Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataCell Reactant:
Cytochrome P450 Reductase (CPR)
Syringe Reactant:
BDBM11941
Meas. Tech.:
Isothermal Titration Calorimetry
Entry Date.:
11/07/06
ΔG°:
-9.8±n/a (kcal/mole)
pH:
7±n/a
Log10Kb:
5.8
Temperature:
298.15±n/a (K)
ΔHobs :
-18±0.1 (kJ/mole)
Corrected for ΔHioniz:
not known
ΔCp :
-0.21±n/a (kJ/mole)
Stoich. Param.:
1
ΔS° :
-0.03±n/a (kJ/mole-K)
Citation
 Hoffman, JM; Grunau, A; Smith, AM; Paine, MJ; Rooney, CS; Ladbury, JE; Fisher, TE; Gutierrez, A; Wai, JS; Thomas, CM; Bamberger, DL; Barnes, JL; Williams, TM; Jones, JH Global effects of the energetics of coenzyme binding: NADPH controls the protein interaction properties of human cytochrome P450 reductase. Biochemistry 45:1421-34 (2006) [PubMed] Article
Hoffman, JM; Grunau, A; Smith, AM; Paine, MJ; Rooney, CS; Ladbury, JE; Fisher, TE; Gutierrez, A; Wai, JS; Thomas, CM; Bamberger, DL; Barnes, JL; Williams, TM; Jones, JH Global effects of the energetics of coenzyme binding: NADPH controls the protein interaction properties of human cytochrome P450 reductase. Biochemistry 45:1421-34 (2006) [PubMed] ArticleCell React
Source:
Human fibroblast CPR (lacking the N-terminal membrane-anchoring region) and the functional FAD-binding domain were expressed in Escherichia coli BL21 (DE3).
Prep. Method:
The recombinant His-tagged proteins were purified to homogeneity by nickel-agarose chromatography. The notable exception is the omission of the 2,5-ADP affinity resin step to avoid the unusual biphasic binding isotherms during ITC experiment.
Name:
NADPH--cytochrome P450 reductase
Synonyms:
CYPOR | Cytochrome P450 Reductase (CPR) | NADPH--cytochrome P450 reductase | NCPR_HUMAN | P450R | POR
Type:
Enzyme
Mol. Mass.:
76675.22
Organism:
Homo sapiens (Human)
Description:
P16435
Residue:
677
Sequence:
MGDSHVDTSSTVSEAVAEEVSLFSMTDMILFSLIVGLLTYWFLFRKKKEEVPEFTKIQTLTSSVRESSFVEKMKKTGRNIIVFYGSQTGTAEEFANRLSKDAHRYGMRGMSADPEEYDLADLSSLPEIDNALVVFCMATYGEGDPTDNAQDFYDWLQETDVDLSGVKFAVFGLGNKTYEHFNAMGKYVDKRLEQLGAQRIFELGLGDDDGNLEEDFITWREQFWPAVCEHFGVEATGEESSIRQYELVVHTDIDAAKVYMGEMGRLKSYENQKPPFDAKNPFLAAVTTNRKLNQGTERHLMHLELDISDSKIRYESGDHVAVYPANDSALVNQLGKILGADLDVVMSLNNLDEESNKKHPFPCPTSYRTALTYYLDITNPPRTNVLYELAQYASEPSEQELLRKMASSSGEGKELYLSWVVEARRHILAILQDCPSLRPPIDHLCELLPRLQARYYSIASSSKVHPNSVHICAVVVEYETKAGRINKGVATNWLRAKEPAGENGGRALVPMFVRKSQFRLPFKATTPVIMVGPGTGVAPFIGFIQERAWLRQQGKEVGETLLYYGCRRSDEDYLYREELAQFHRDGALTQLNVAFSREQSHKVYVQHLLKQDREHLWKLIEGGAHIYVCGDARNMARDVQNTFYDIVAELGAMEHAQAVDYIKKLMTKGRYSLDVWS
Syringe React
Source:
Sigma-Aldrich
Name:
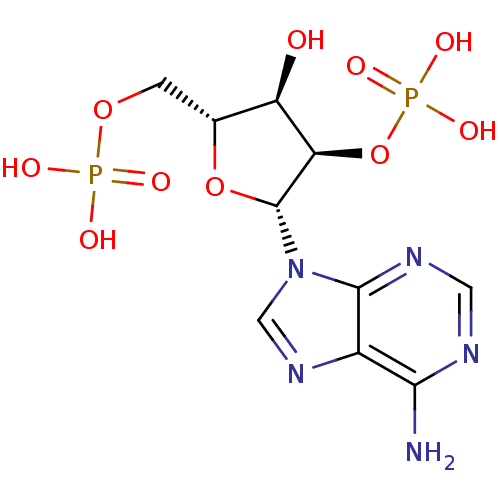
BDBM11941
Synonyms:
2,5-ADP | ADENOSINE-2 -5 -DIPHOSPHATE | {[(2R,3R,4R,5R)-5-(6-amino-9H-purin-9-yl)-3-hydroxy-4-(phosphonooxy)oxolan-2-yl]methoxy}phosphonic acid
Type:
Nucleoside or nucleotide
Emp. Form.:
C10H15N5O10P2
Mol. Mass.:
427.2011
SMILES:
Nc1ncnc2n(cnc12)[C@@H]1O[C@H](COP(O)(O)=O)[C@@H](O)[C@H]1OP(O)(O)=O |r|