Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataCell Reactant:
MAP kinase ERK1
Syringe Reactant:
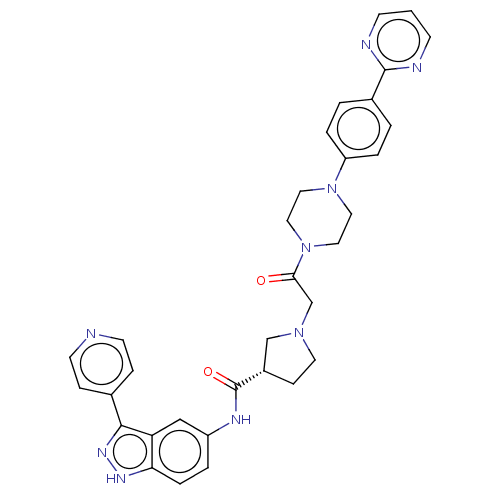
BDBM103302
Meas. Tech.:
Enzyme Inhibition
Entry Date.:
10/22/14
ΔG°:
-36.75±n/a (kJ/mole)
pH:
7.5±0
Temperature:
288.15±0 (K)
ΔH° :
-86.2±0.8 (kJ/mole)
ΔS° :
-0.1716±0 (kJ/mole-K)
Citation
 Chaikuad, A; Tacconi, EM; Zimmer, J; Liang, Y; Gray, NS; Tarsounas, M; Knapp, S A unique inhibitor binding site in ER K1/2 is associated with slow binding kinetics Nat Chem Biol 10:853-860 (2014) [PubMed] Article
Chaikuad, A; Tacconi, EM; Zimmer, J; Liang, Y; Gray, NS; Tarsounas, M; Knapp, S A unique inhibitor binding site in ER K1/2 is associated with slow binding kinetics Nat Chem Biol 10:853-860 (2014) [PubMed] Article More Info.:
Cell React
Name:
Mitogen-activated protein kinase 3
Synonyms:
ERK1 | ERK1/ERK2 | Extracellular signal-regulated kinase 1 | Extracellular signal-regulated kinase 1 (ERK1) | MAP kinase ERK1 | MAPK3 | MAPK3/MAPK7 | MK03_HUMAN | Mitogen-activated protein kinase | Mitogen-activated protein kinase 3 | PRKM3
Type:
Protein
Mol. Mass.:
43136.58
Organism:
Homo sapiens (Human)
Description:
curated by PDB 4QTB
Residue:
379
Sequence:
MAAAAAQGGGGGEPRRTEGVGPGVPGEVEMVKGQPFDVGPRYTQLQYIGEGAYGMVSSAYDHVRKTRVAIKKISPFEHQTYCQRTLREIQILLRFRHENVIGIRDILRASTLEAMRDVYIVQDLMETDLYKLLKSQQLSNDHICYFLYQILRGLKYIHSANVLHRDLKPSNLLINTTCDLKICDFGLARIADPEHDHTGFLTEYVATRWYRAPEIMLNSKGYTKSIDIWSVGCILAEMLSNRPIFPGKHYLDQLNHILGILGSPSQEDLNCIINMKARNYLQSLPSKTKVAWAKLFPKSDSKALDLLDRMLTFNPNKRITVEEALAHPYLEQYYDPTDEPVAEEPFTFAMELDDLPKERLKELIFQETARFQPGVLEAP