Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
UDP-3-O-acyl-N-acetylglucosamine deacetylase
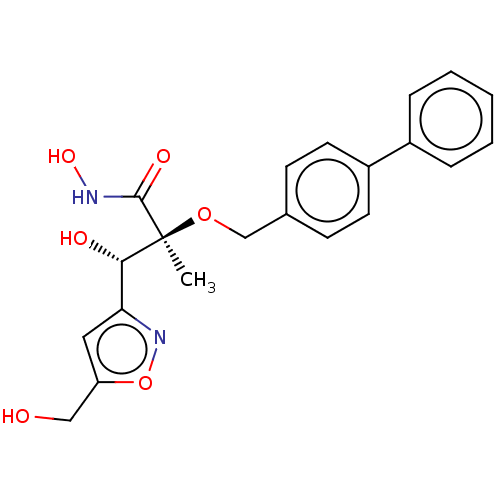
Ligand
BDBM50484896
Substrate
n/a
Meas. Tech.
ChEMBL_810002 (CHEMBL2015111)
IC50
0.600000±n/a nM
Citation
 Warmus, JS; Quinn, CL; Taylor, C; Murphy, ST; Johnson, TA; Limberakis, C; Ortwine, D; Bronstein, J; Pagano, P; Knafels, JD; Lightle, S; Mochalkin, I; Brideau, R; Podoll, T Structure based design of an in vivo active hydroxamic acid inhibitor of P. aeruginosa LpxC. Bioorg Med Chem Lett 22:2536-43 (2012) [PubMed] Article
Warmus, JS; Quinn, CL; Taylor, C; Murphy, ST; Johnson, TA; Limberakis, C; Ortwine, D; Bronstein, J; Pagano, P; Knafels, JD; Lightle, S; Mochalkin, I; Brideau, R; Podoll, T Structure based design of an in vivo active hydroxamic acid inhibitor of P. aeruginosa LpxC. Bioorg Med Chem Lett 22:2536-43 (2012) [PubMed] Article More Info.:
Target
Name:
UDP-3-O-acyl-N-acetylglucosamine deacetylase
Synonyms:
LPXC_PSEAE | Protein envA | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase (LpxC | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase (LpxC) | UDP-3-O-acyl-GlcNAc deacetylase | UDP-3-O-acyl-GlcNAc deacetylase (LpxC) | envA | lpxC
Type:
Enzyme
Mol. Mass.:
33428.15
Organism:
Pseudomonas aeruginosa
Description:
P47205
Residue:
303
Sequence:
MIKQRTLKNIIRATGVGLHSGEKVYLTLKPAPVDTGIVFCRTDLDPVVEIPARAENVGETTMSTTLVKGDVKVDTVEHLLSAMAGLGIDNAYVELSASEVPIMDGSAGPFVFLIQSAGLQEQEAAKKFIRIKREVSVEEGDKRAVFVPFDGFKVSFEIDFDHPVFRGRTQQASVDFSSTSFVKEVSRARTFGFMRDIEYLRSQNLALGGSVENAIVVDENRVLNEDGLRYEDEFVKHKILDAIGDLYLLGNSLIGEFRGFKSGHALNNQLLRTLIADKDAWEVVTFEDARTAPISYMRPAAAV