Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Lysine-specific demethylase 5C
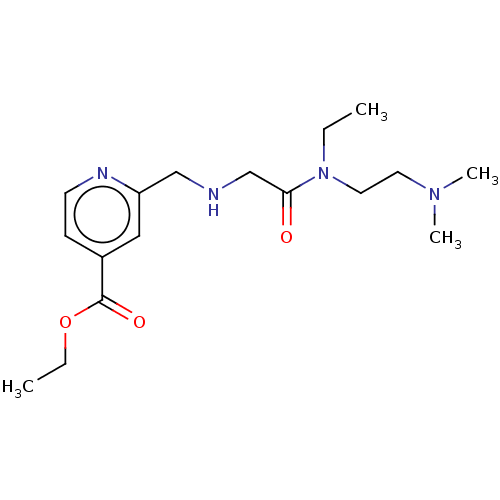
Ligand
BDBM191599
Substrate
n/a
Meas. Tech.
AlphaLISA Assay
pH
7.2±n/a
Temperature
298.15±n/a K
IC50
5.8e+2± 2e+1 nM
Comments
extracted
Citation
 Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol 23:769-81 (2016) [PubMed] Article
Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol 23:769-81 (2016) [PubMed] Article More Info.:
Target
Name:
Lysine-specific demethylase 5C
Synonyms:
DXS1272E | Histone demethylase JARID1C | JARID1C | Jumonji/ARID domain-containing protein 1C | KDM5C | KDM5C_HUMAN | Lysine-specific demethylase 5C (KDM5C) | Lysine-specific demethylase 5C (KDM5Flag-5C-FL) | Protein SmcX | Protein Xe169 | SMCX | XE169
Type:
PROTEIN
Mol. Mass.:
175691.17
Organism:
Homo sapiens (Human)
Description:
P41229
Residue:
1560
Sequence:
MEPGSDDFLPPPECPVFEPSWAEFRDPLGYIAKIRPIAEKSGICKIRPPADWQPPFAVEVDNFRFTPRIQRLNELEAQTRVKLNYLDQIAKFWEIQGSSLKIPNVERRILDLYSLSKIVVEEGGYEAICKDRRWARVAQRLNYPPGKNIGSLLRSHYERIVYPYEMYQSGANLVQCNTRPFDNEEKDKEYKPHSIPLRQSVQPSKFNSYGRRAKRLQPDPEPTEEDIEKNPELKKLQIYGAGPKMMGLGLMAKDKTLRKKDKEGPECPPTVVVKEELGGDVKVESTSPKTFLESKEELSHSPEPCTKMTMRLRRNHSNAQFIESYVCRMCSRGDEDDKLLLCDGCDDNYHIFCLLPPLPEIPKGVWRCPKCVMAECKRPPEAFGFEQATREYTLQSFGEMADSFKADYFNMPVHMVPTELVEKEFWRLVNSIEEDVTVEYGADIHSKEFGSGFPVSDSKRHLTPEEEEYATSGWNLNVMPVLEQSVLCHINADISGMKVPWLYVGMVFSAFCWHIEDHWSYSINYLHWGEPKTWYGVPSLAAEHLEEVMKKLTPELFDSQPDLLHQLVTLMNPNTLMSHGVPVVRTNQCAGEFVITFPRAYHSGFNQGYNFAEAVNFCTADWLPAGRQCIEHYRRLRRYCVFSHEELICKMAACPEKLDLNLAAAVHKEMFIMVQEERRLRKALLEKGITEAEREAFELLPDDERQCIKCKTTCFLSALACYDCPDGLVCLSHINDLCKCSSSRQYLRYRYTLDELPAMLHKLKVRAESFDTWANKVRVALEVEDGRKRSLEELRALESEARERRFPNSELLQQLKNCLSEAEACVSRALGLVSGQEAGPHRVAGLQMTLTELRAFLDQMNNLPCAMHQIGDVKGVLEQVEAYQAEAREALASLPSSPGLLQSLLERGRQLGVEVPEAQQLQRQVEQARWLDEVKRTLAPSARRGTLAVMRGLLVAGASVAPSPAVDKAQAELQELLTIAERWEEKAHLCLEARQKHPPATLEAIIREAENIPVHLPNIQALKEALAKARAWIADVDEIQNGDHYPCLDDLEGLVAVGRDLPVGLEELRQLELQVLTAHSWREKASKTFLKKNSCYTLLEVLCPCADAGSDSTKRSRWMEKELGLYKSDTELLGLSAQDLRDPGSVIVAFKEGEQKEKEGILQLRRTNSAKPSPLASSSTASSTTSICVCGQVLAGAGALQCDLCQDWFHGRCVSVPRLLSSPRPNPTSSPLLAWWEWDTKFLCPLCMRSRRPRLETILALLVALQRLPVRLPEGEALQCLTERAISWQGRARQALASEDVTALLGRLAELRQRLQAEPRPEEPPNYPAAPASDPLREGSGKDMPKVQGLLENGDSVTSPEKVAPEEGSGKRDLELLSSLLPQLTGPVLELPEATRAPLEELMMEGDLLEVTLDENHSIWQLLQAGQPPDLERIRTLLELEKAERHGSRARGRALERRRRRKVDRGGEGDDPAREELEPKRVRSSGPEAEEVQEEEELEEETGGEGPPAPIPTTGSPSTQENQNGLEPAEGTTSGPSAPFSTLTPRLHLPCPQQPPQQQL