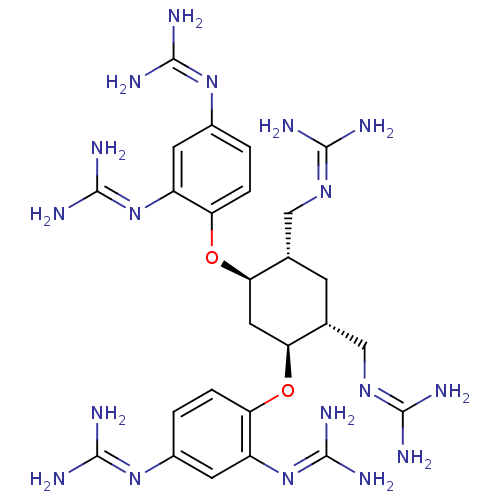
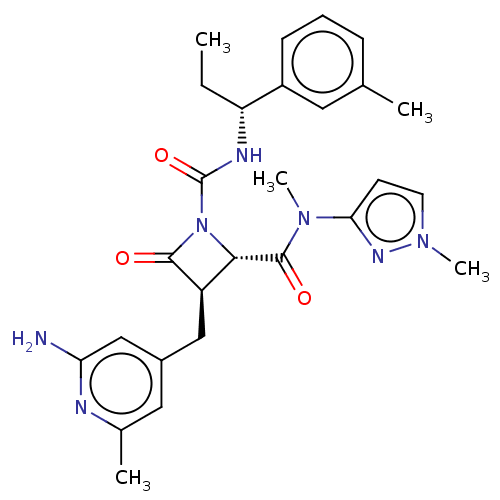
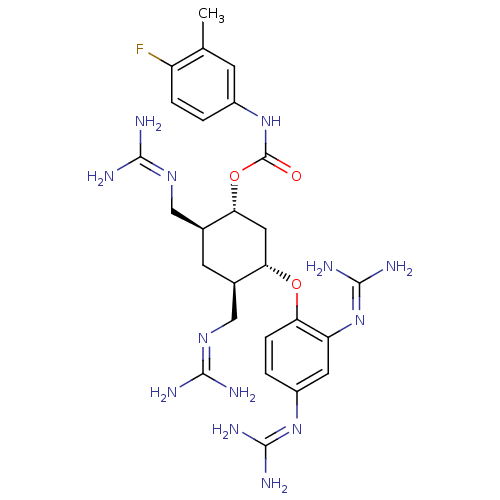
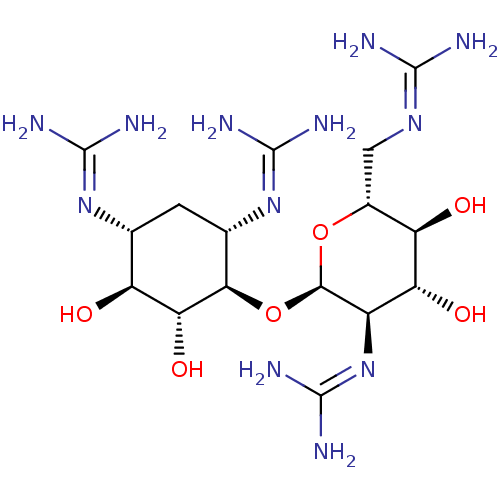
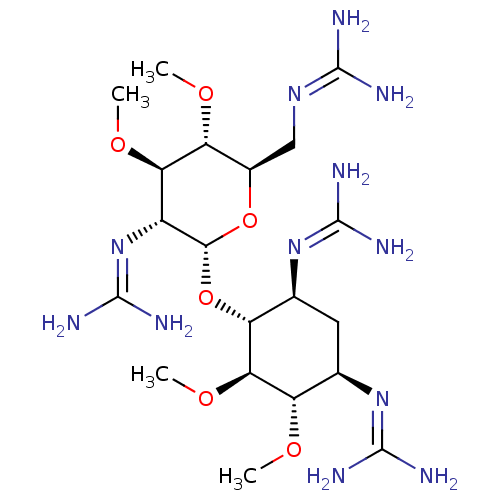
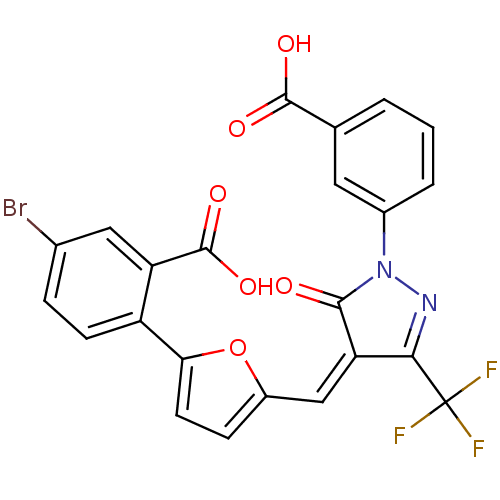
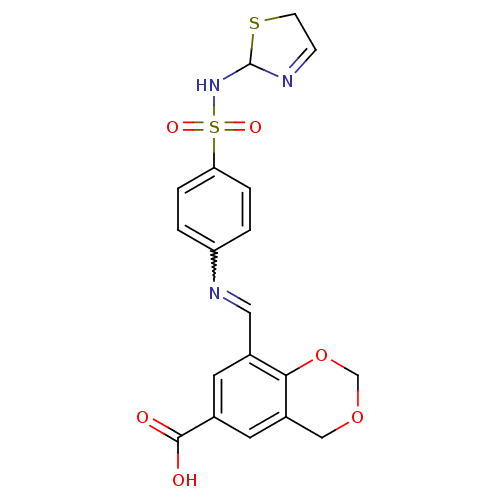
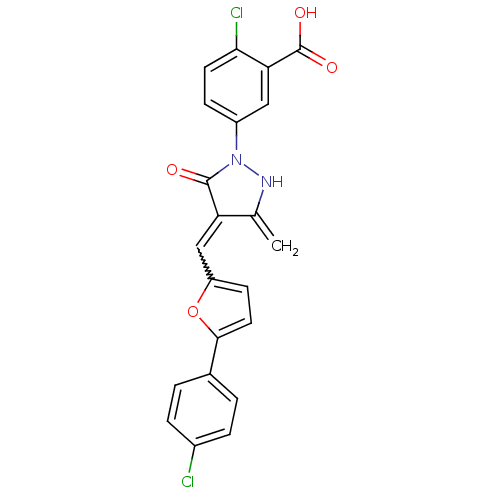
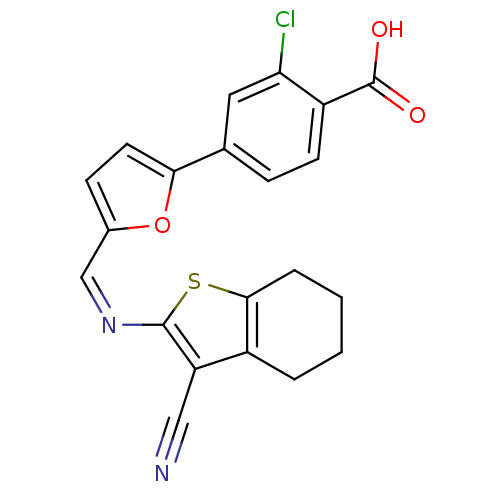
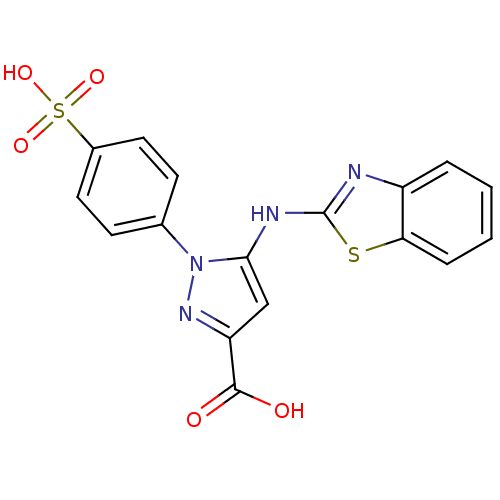
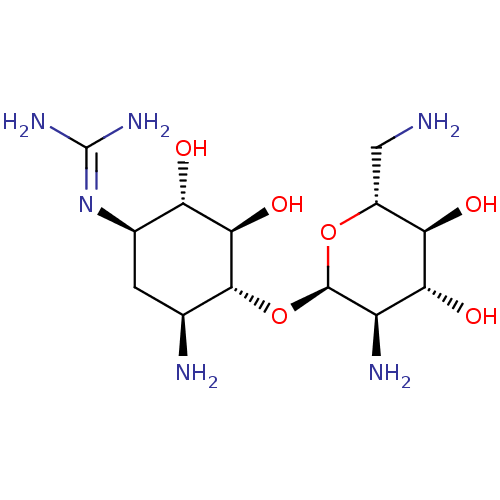
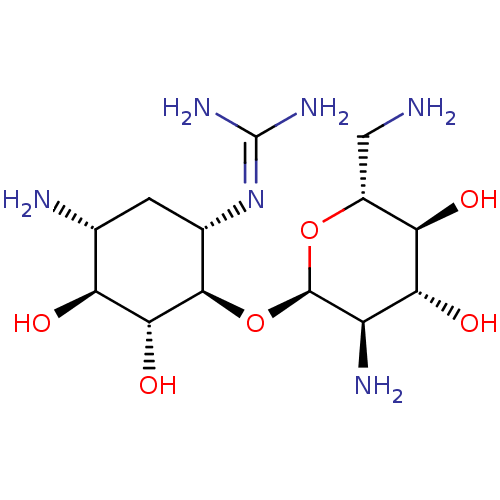
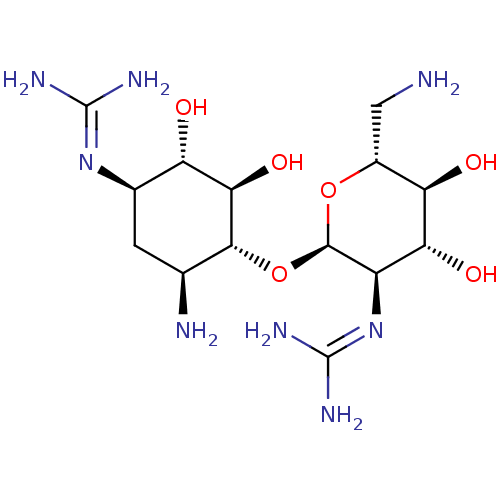
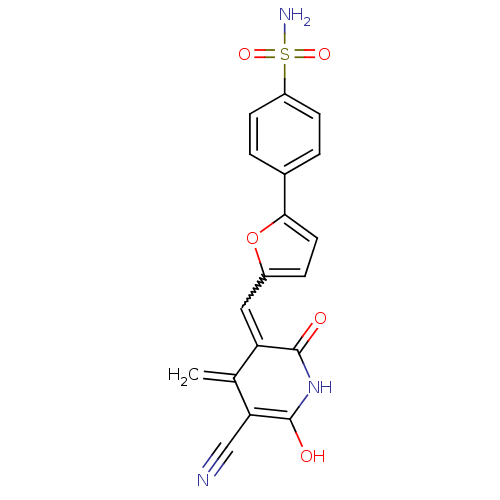
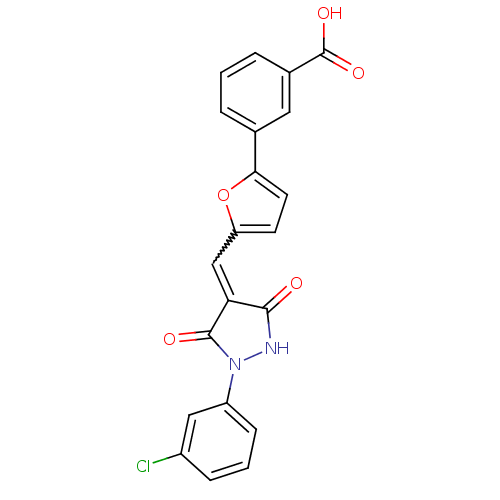
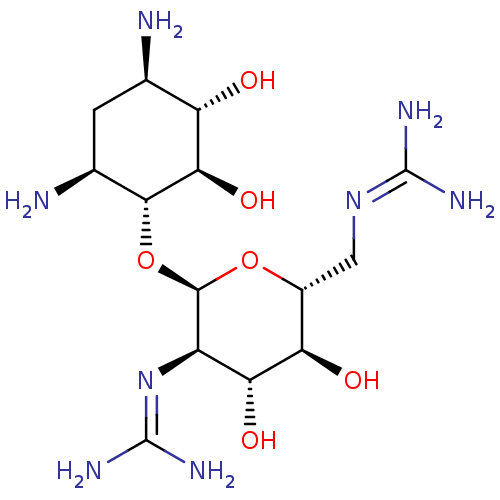
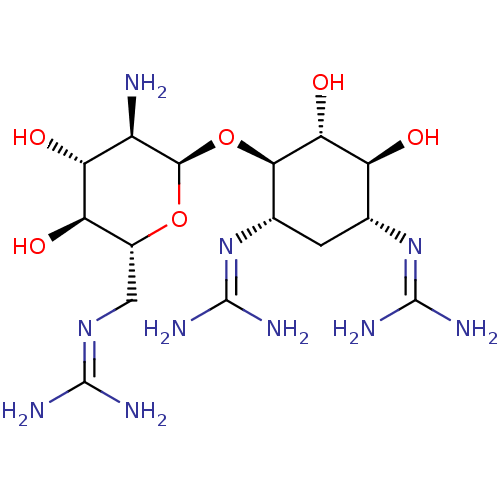
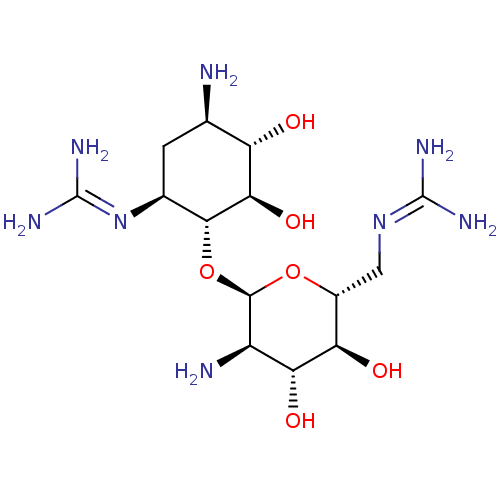
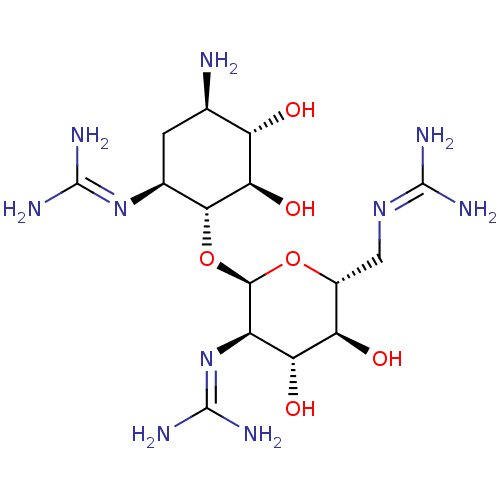
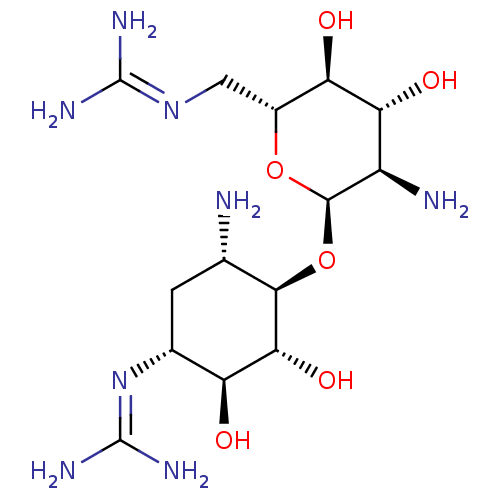
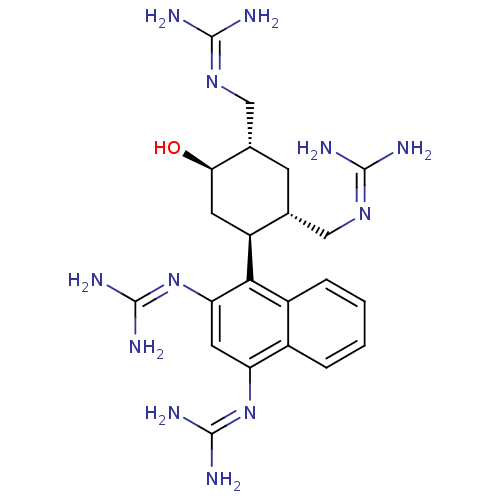
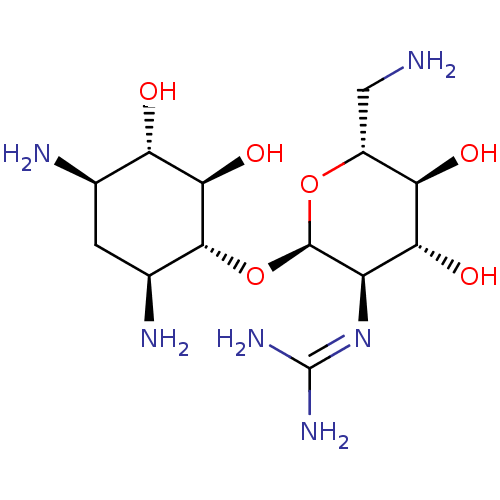
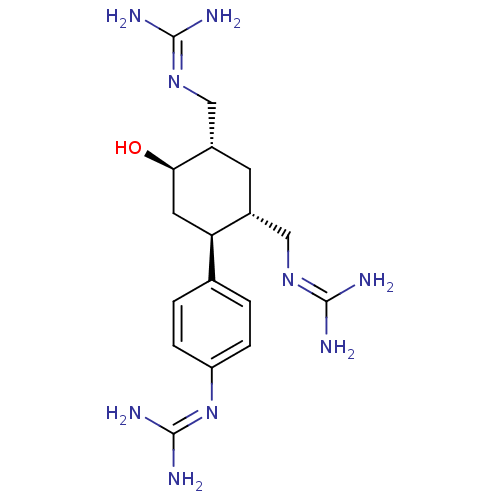
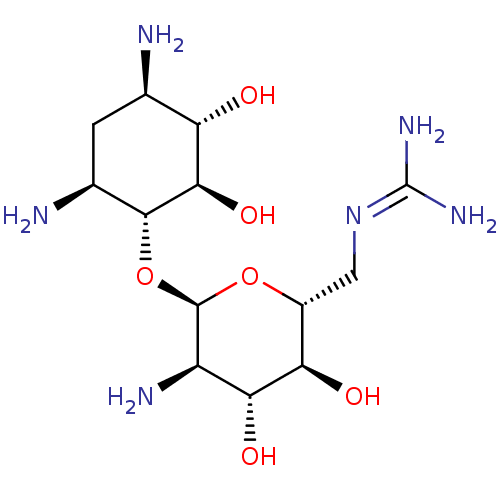
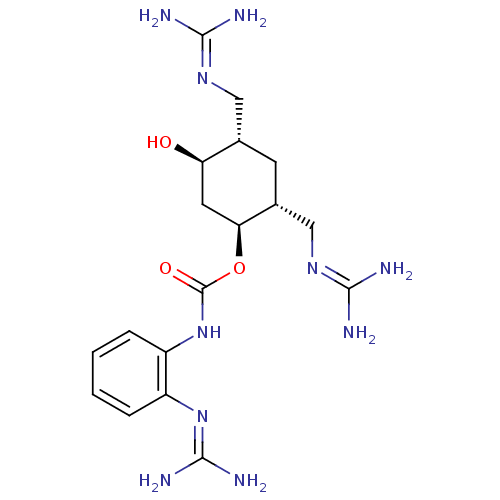
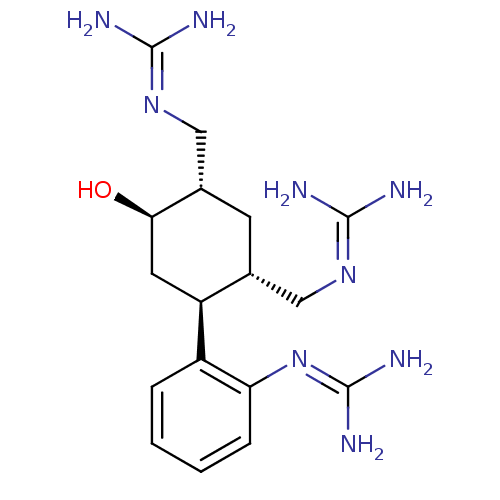
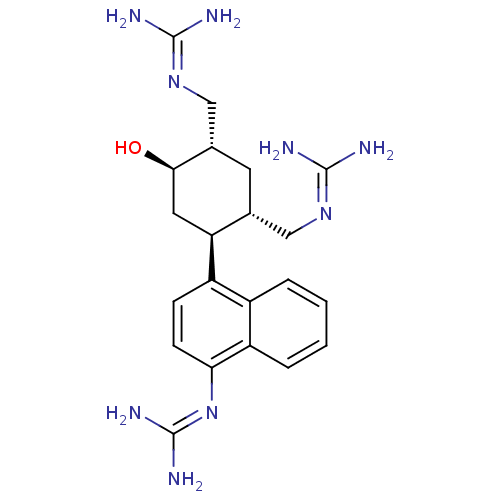
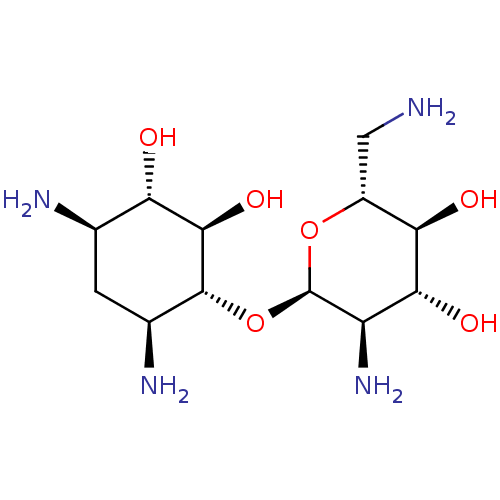
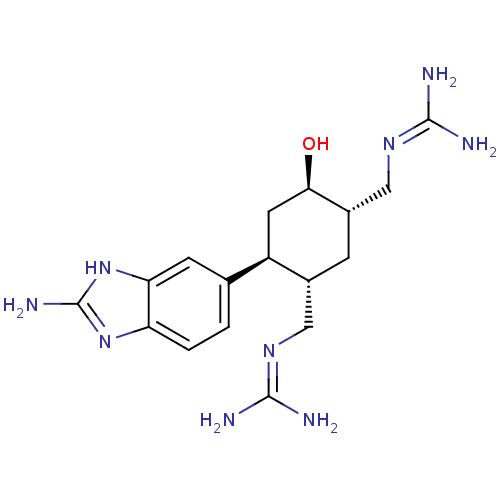
Affinity DataKi: 65nM ΔG°: -40.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
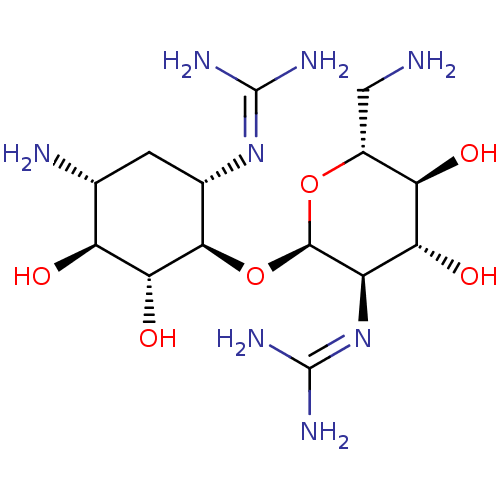
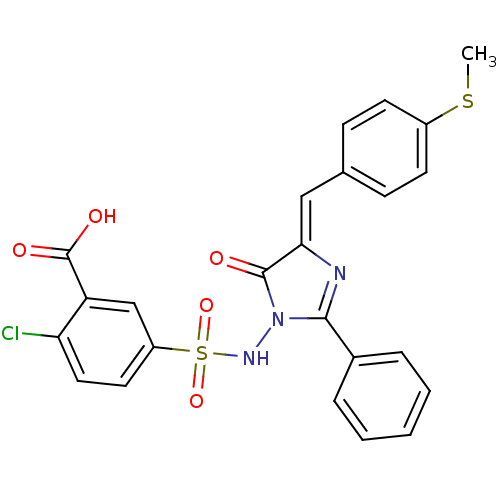
Affinity DataKi: 500nM ΔG°: -35.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
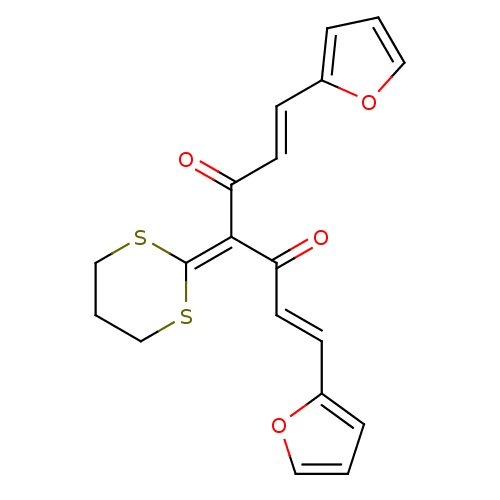
Affinity DataKi: 500nM ΔG°: -35.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
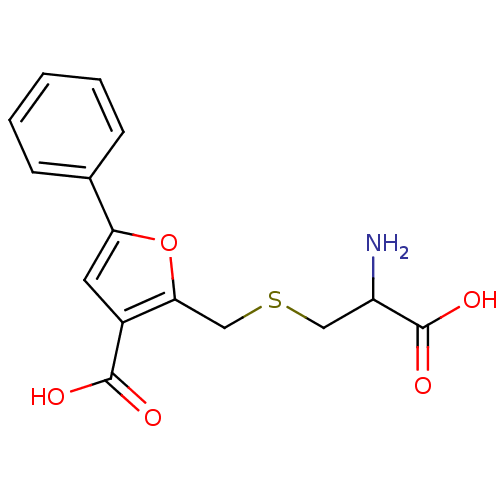
Affinity DataKi: 500nM ΔG°: -35.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.2kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -35.0kJ/mole IC50: 1.10E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 4.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 7.60E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 8.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nM ΔG°: -34.2kJ/mole IC50: 3.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nM ΔG°: -33.5kJ/mole IC50: 3.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nM ΔG°: -32.9kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -33.3kJ/mole IC50: 1.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -33.3kJ/mole IC50: 2.90E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nM ΔG°: -33.0kJ/mole IC50: 8.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+3nM ΔG°: -32.6kJ/mole IC50: 1.00E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.3kJ/mole IC50: 2.90E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.3kJ/mole IC50: 5.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nM ΔG°: -32.2kJ/mole IC50: 3.30E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nM ΔG°: -32.0kJ/mole IC50: 3.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.90E+3nM ΔG°: -31.8kJ/mole IC50: 4.10E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 3.10E+3nM ΔG°: -31.7kJ/mole IC50: 4.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 3.10E+3nM ΔG°: -31.1kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 3.30E+3nM ΔG°: -31.5kJ/mole IC50: 4.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+3nM ΔG°: -30.4kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nM ΔG°: -30.9kJ/mole IC50: 9.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nM ΔG°: -30.0kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+3nM ΔG°: -29.9kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 5.30E+3nM ΔG°: -29.8kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 5.40E+3nM ΔG°: -30.3kJ/mole IC50: 1.08E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 5.60E+3nM ΔG°: -30.2kJ/mole IC50: 6.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 5.60E+3nM ΔG°: -29.7kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 6.60E+3nM ΔG°: -29.3kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 7.20E+3nM ΔG°: -29.5kJ/mole IC50: 1.39E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 7.70E+3nM ΔG°: -28.9kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 8.70E+3nM ΔG°: -28.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 8.80E+3nM ΔG°: -28.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 9.20E+3nM ΔG°: -28.5kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.02E+4nM ΔG°: -28.2kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.07E+4nM ΔG°: -28.1kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.09E+4nM ΔG°: -28.0kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.49E+4nM ΔG°: -27.3kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 2.43E+4nM ΔG°: -26.1kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 2.48E+4nM ΔG°: -26.0kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 3.06E+4nM ΔG°: -25.5kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 3.14E+4nM ΔG°: -25.4kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 4.29E+4nM ΔG°: -24.7kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair
Affinity DataKi: 1.54E+5nM ΔG°: -21.5kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic reaction started by the addition fluorogenic peptide substrate, MAPKKide to the buffer containing LF and inhibitor compound. Cleavage o...More data for this Ligand-Target Pair