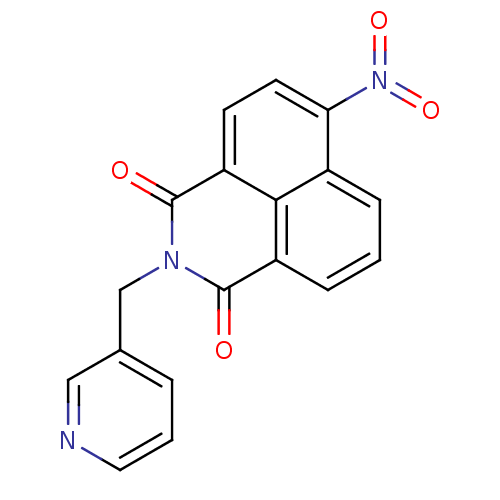
Affinity DataIC50: 28nM Kd: 5.30E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nM Kd: 9.40E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nM Kd: 9.80E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+3nM Kd: 160nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nM Kd: 4.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nM Kd: 9.30E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nM Kd: 6.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nM Kd: 4.90E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nM Kd: 1.60E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nM Kd: 1.20E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+4nM Kd: 1.07E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nM Kd: 1.00E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: 1.41E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >2.00E+6nMAssay Description:Binding affinity to Mycobacterium tuberculosis UGM assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: 100nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >1.00E+6nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 4.01E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 6.53E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 9.48E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 3.58E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 6.30E+4nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 1.97E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 3.50E+4nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by fluorescence polarization a...More data for this Ligand-Target Pair
Affinity DataKd: 1.05E+5nMAssay Description:Inhibition of Mycobacterium tuberculosis UGM expressed in Escherichia coli BL21(DE3) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.00E+6nMAssay Description:Binding affinity to Mycobacterium tuberculosis UGM assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >1.12E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: 8.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair






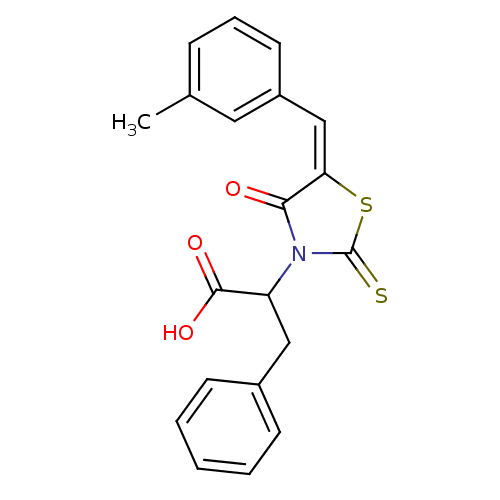
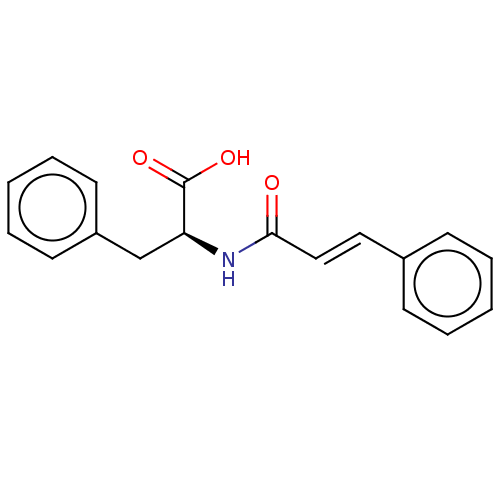

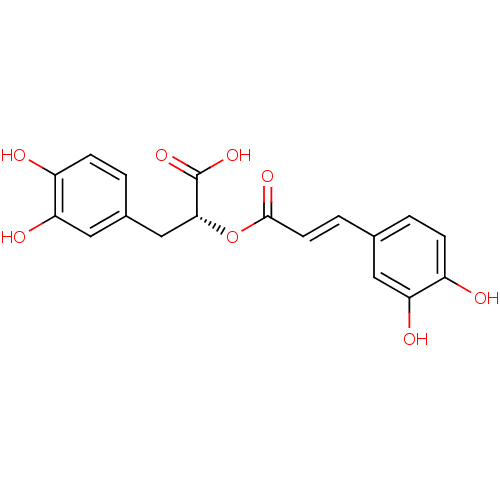


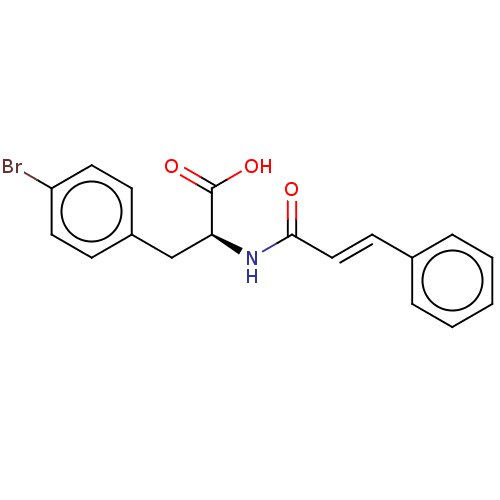
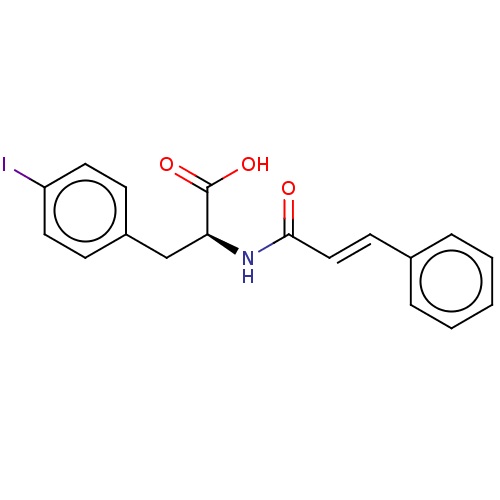
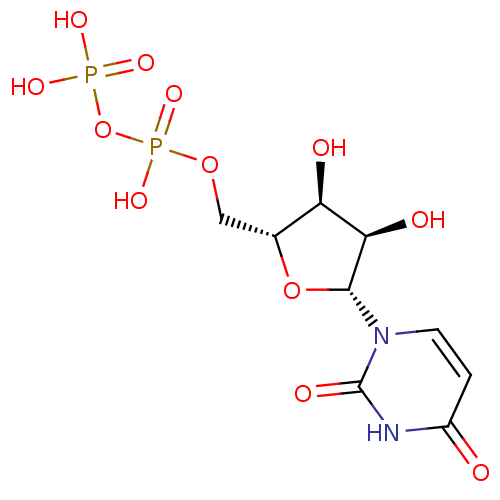
 3D Structure (crystal)
3D Structure (crystal)