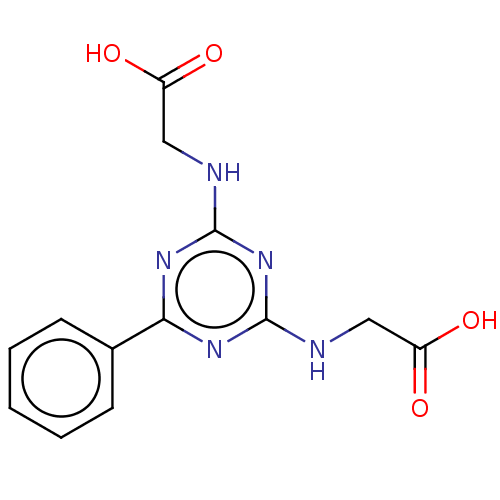
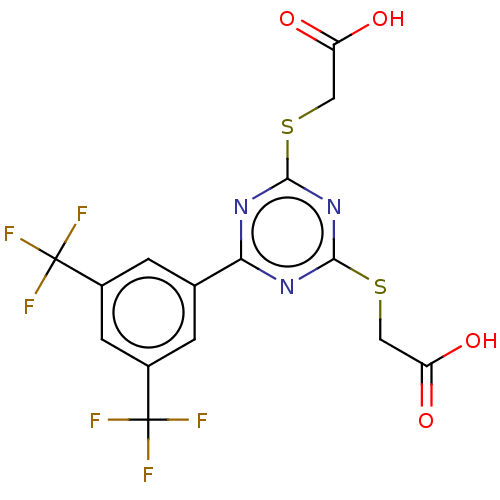
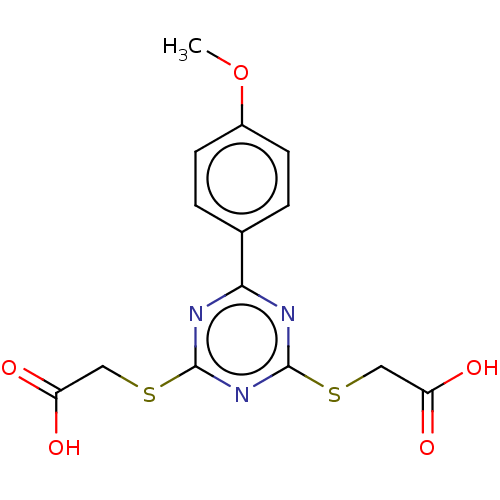
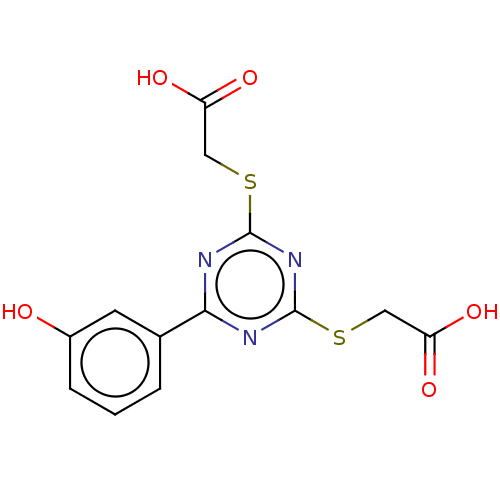
Affinity DataKd: 7.70E+4nMAssay Description:Binding affinity to human recombinant CXCL12 by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.88E+4nMAssay Description:Binding affinity to human CXCL12 assessed as dissociation constant incubated for 3 mins by surface plasmon resonance analysisMore data for this Ligand-Target Pair
Affinity DataKd: 2.40E+4nMAssay Description:Binding affinity to human recombinant CXCL12 by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.20E+6nMAssay Description:Binding affinity to His6-SUMO fusion tagged wild type human [U-15N]CXCL12 expressed in Escherichia coli by 2D 1H-15N HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.70E+6nMAssay Description:Binding affinity to His6-SUMO fusion tagged wild type human [U-15N]CXCL12 expressed in Escherichia coli by 2D 1H-15N HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.10E+6nMAssay Description:Binding affinity to His6-SUMO fusion tagged wild type human [U-15N]CXCL12 expressed in Escherichia coli by 2D 1H-15N HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 4.10E+6nMAssay Description:Binding affinity to His6-SUMO fusion tagged wild type human [U-15N]CXCL12 expressed in Escherichia coli by 2D 1H-15N HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 4.10E+6nMAssay Description:Binding affinity to His6-SUMO fusion tagged wild type human [U-15N]CXCL12 expressed in Escherichia coli by 2D 1H-15N HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 7.40E+4nMAssay Description:Binding affinity to human [U-15N]CXCL12 L36C/A65C dimer mutant expressed in Escherichia coli by 2D 1H-15N SOFAST HMQC NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+5nMAssay Description:Binding affinity to human [U-15N]CXCL12 L36C/A65C dimer mutant expressed in Escherichia coli assessed as induction of chemical shift perturbations at...More data for this Ligand-Target Pair
Affinity DataKd: 780nMAssay Description:Binding affinity to CXCL12 (unknown origin) assessed as changes in tryptophan fluorescence intensityMore data for this Ligand-Target Pair
Affinity DataKd: 1.69E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.20E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 9.00E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.58E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 3.55E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.58E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.56E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 3.83E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 6.46E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 3.57E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 3.06E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.80E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.33E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.34E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.64E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 8.00E+4nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 2.51E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.36E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.70E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 1.03E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 2.34E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 2.33E+5nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 3.02E+6nMAssay Description:Binding affinity to wild type CXCL12 (unknown origin) by NMR spectroscopyMore data for this Ligand-Target Pair
Affinity DataKd: 2.50E+8nMAssay Description:Binding affinity to human recombinant CXCL12 by surface plasmon resonance assayMore data for this Ligand-Target Pair