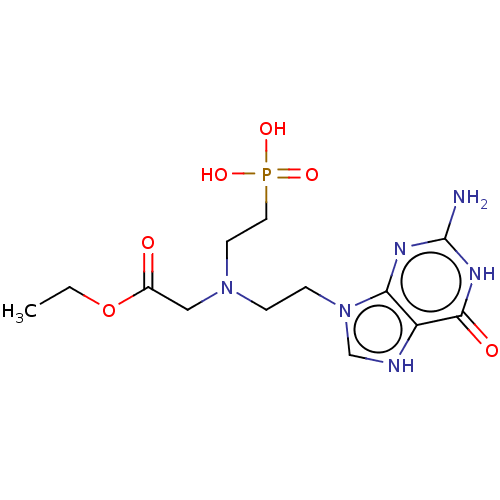
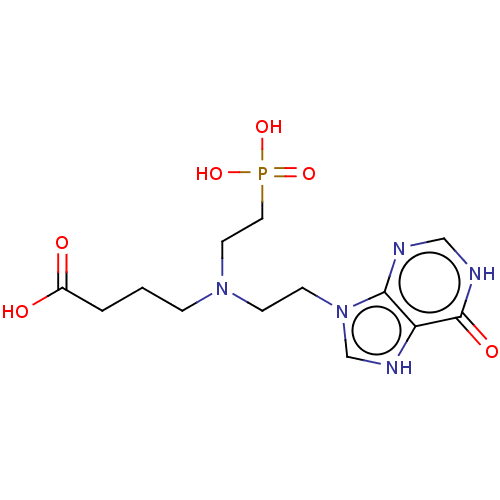
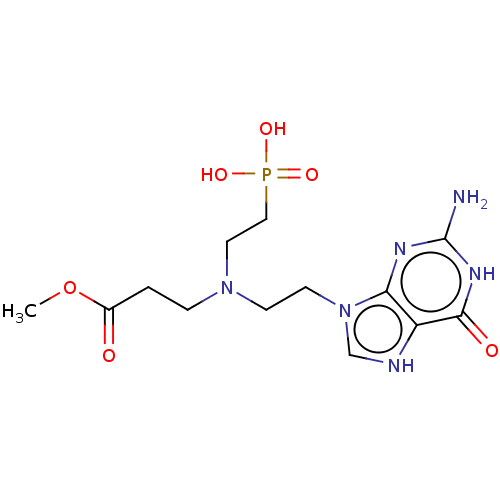
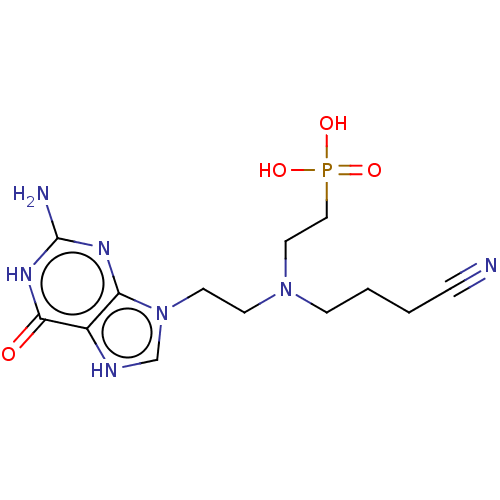
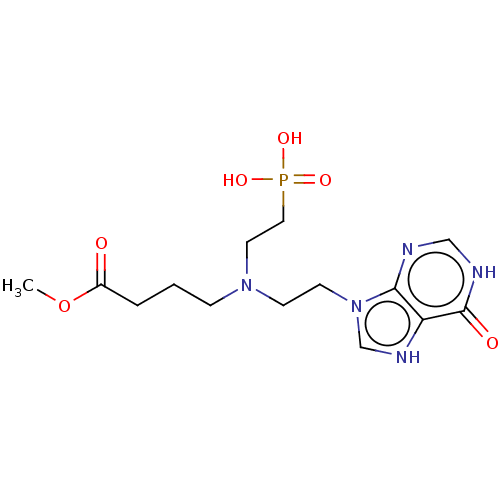
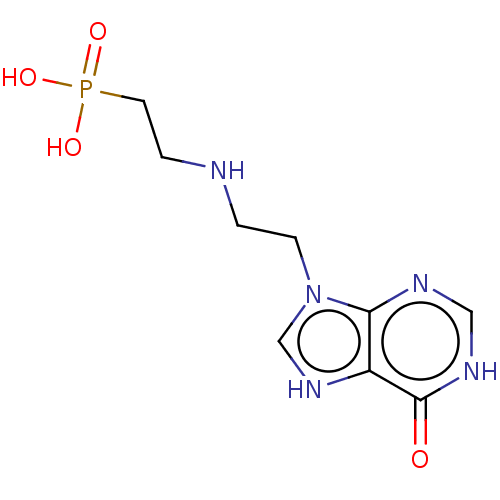
Affinity DataKi: 50nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 300nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -36.5kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 900nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -34.4kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nM ΔG°: -29.4kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+4nM ΔG°: -29.0kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nM ΔG°: -27.8kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nM ΔG°: -27.8kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+4nM ΔG°: -26.0kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 4.70E+4nM ΔG°: -25.7kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: 1.78E+5nM ΔG°: -22.3kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+5nMpH: 7.4Assay Description:The Ki values were determined using a spectrophotometric assay at 25� C., 0.1 M Tris-HCl, 10 mM MgCl2, pH 7.4 (Keough, D. T.; Ng, A. L.; Winzor, D. J...More data for this Ligand-Target Pair
Affinity DataKi: >2.30E+5nM ΔG°: >-21.6kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+5nM ΔG°: >-21.4kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+5nM ΔG°: >-21.4kJ/moleT: 2°CAssay Description:The [3H]-hypoxanthine growth inhibition assay (Desjardins et al., 1979
Antimicrobial Agents Chemother 16: 710-718) was used to evaluate the in
vitr...More data for this Ligand-Target Pair