Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Macrophage-stimulating protein receptor
Ligand
BDBM50306682
Substrate
n/a
Meas. Tech.
ChEMBL_531097 (CHEMBL976905)
IC50
300±n/a nM
Citation
 McDermott, U; Sharma, SV; Dowell, L; Greninger, P; Montagut, C; Lamb, J; Archibald, H; Raudales, R; Tam, A; Lee, D; Rothenberg, SM; Supko, JG; Sordella, R; Ulkus, LE; Iafrate, AJ; Maheswaran, S; Njauw, CN; Tsao, H; Drew, L; Hanke, JH; Ma, XJ; Erlander, MG; Gray, NS; Haber, DA; Settleman, J Identification of genotype-correlated sensitivity to selective kinase inhibitors by using high-throughput tumor cell line profiling. Proc Natl Acad Sci U S A 104:19936-41 (2007) [PubMed] Article
McDermott, U; Sharma, SV; Dowell, L; Greninger, P; Montagut, C; Lamb, J; Archibald, H; Raudales, R; Tam, A; Lee, D; Rothenberg, SM; Supko, JG; Sordella, R; Ulkus, LE; Iafrate, AJ; Maheswaran, S; Njauw, CN; Tsao, H; Drew, L; Hanke, JH; Ma, XJ; Erlander, MG; Gray, NS; Haber, DA; Settleman, J Identification of genotype-correlated sensitivity to selective kinase inhibitors by using high-throughput tumor cell line profiling. Proc Natl Acad Sci U S A 104:19936-41 (2007) [PubMed] Article More Info.:
Target
Name:
Macrophage-stimulating protein receptor
Synonyms:
2.7.10.1 | CD_antigen=CD136 | CDw136 | MSP receptor | MST1R | Macrophage-stimulating protein receptor (MST1R) | Macrophage-stimulating protein receptor alpha chain | Macrophage-stimulating protein receptor beta chain | PTK8 | Protein-tyrosine kinase 8 | RON | RON_HUMAN | Tyrosine kinase receptor ron | p185-Ron
Type:
Protein
Mol. Mass.:
152270.76
Organism:
Homo sapiens (Human)
Description:
Q04912
Residue:
1400
Sequence:
MELLPPLPQSFLLLLLLPAKPAAGEDWQCPRTPYAASRDFDVKYVVPSFSAGGLVQAMVTYEGDRNESAVFVAIRNRLHVLGPDLKSVQSLATGPAGDPGCQTCAACGPGPHGPPGDTDTKVLVLDPALPALVSCGSSLQGRCFLHDLEPQGTAVHLAAPACLFSAHHNRPDDCPDCVASPLGTRVTVVEQGQASYFYVASSLDAAVAASFSPRSVSIRRLKADASGFAPGFVALSVLPKHLVSYSIEYVHSFHTGAFVYFLTVQPASVTDDPSALHTRLARLSATEPELGDYRELVLDCRFAPKRRRRGAPEGGQPYPVLRVAHSAPVGAQLATELSIAEGQEVLFGVFVTGKDGGPGVGPNSVVCAFPIDLLDTLIDEGVERCCESPVHPGLRRGLDFFQSPSFCPNPPGLEALSPNTSCRHFPLLVSSSFSRVDLFNGLLGPVQVTALYVTRLDNVTVAHMGTMDGRILQVELVRSLNYLLYVSNFSLGDSGQPVQRDVSRLGDHLLFASGDQVFQVPIQGPGCRHFLTCGRCLRAWHFMGCGWCGNMCGQQKECPGSWQQDHCPPKLTEFHPHSGPLRGSTRLTLCGSNFYLHPSGLVPEGTHQVTVGQSPCRPLPKDSSKLRPVPRKDFVEEFECELEPLGTQAVGPTNVSLTVTNMPPGKHFRVDGTSVLRGFSFMEPVLIAVQPLFGPRAGGTCLTLEGQSLSVGTSRAVLVNGTECLLARVSEGQLLCATPPGATVASVPLSLQVGGAQVPGSWTFQYREDPVVLSISPNCGYINSHITICGQHLTSAWHLVLSFHDGLRAVESRCERQLPEQQLCRLPEYVVRDPQGWVAGNLSARGDGAAGFTLPGFRFLPPPHPPSANLVPLKPEEHAIKFEYIGLGAVADCVGINVTVGGESCQHEFRGDMVVCPLPPSLQLGQDGAPLQVCVDGECHILGRVVRPGPDGVPQSTLLGILLPLLLLVAALATALVFSYWWRRKQLVLPPNLNDLASLDQTAGATPLPILYSGSDYRSGLALPAIDGLDSTTCVHGASFSDSEDESCVPLLRKESIQLRDLDSALLAEVKDVLIPHERVVTHSDRVIGKGHFGVVYHGEYIDQAQNRIQCAIKSLSRITEMQQVEAFLREGLLMRGLNHPNVLALIGIMLPPEGLPHVLLPYMCHGDLLQFIRSPQRNPTVKDLISFGLQVARGMEYLAEQKFVHRDLAARNCMLDESFTVKVADFGLARDILDREYYSVQQHRHARLPVKWMALESLQTYRFTTKSDVWSFGVLLWELLTRGAPPYRHIDPFDLTHFLAQGRRLPQPEYCPDSLYQVMQQCWEADPAVRPTFRVLVGEVEQIVSALLGDHYVQLPATYMNLGPSTSHEMNVRPEQPQFSPMPGNVRRPRPLSEPPRPT
Inhibitor
Name:
BDBM50306682
Synonyms:
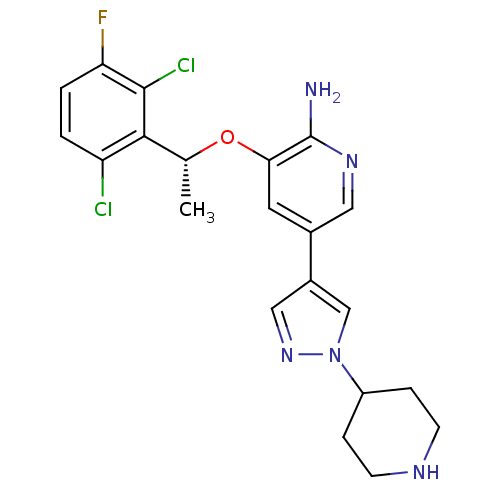
(R)-3-(1-(2,6-dichloro-3-fluorophenyl)ethoxy)-5-(1-(piperidin-4-yl)-1H-pyrazol-4-yl)pyridin-2-amine | 3-(2,6-dichloro-3-fluorobenzyloxy)-5-(1-(piperidin-4-yl)-1H-pyrazol-4-yl)pyridin-2-amine | 3-[(1R)-1-(2,6-dichloro-3-fluorophenyl)ethoxy]-5-(1-piperidin-4-yl-1H-pyrazol-4-yl)pyridin-2-amine | CHEMBL601719 | CRIZOTINIB | PF-2341066 | US10370379, Crizotinib | US10543199, Compound Crizotinib | US10780082, Compound Crizotinib | US11059827, Compound Crizotinib | US11517561, Compound Crizotinib | US9126941, PF-2341066 | US9199944, Crizotinib | US9226923, Crizotinib
Type:
Small organic molecule
Emp. Form.:
C21H22Cl2FN5O
Mol. Mass.:
450.337
SMILES:
C[C@@H](Oc1cc(cnc1N)-c1cnn(c1)C1CCNCC1)c1c(Cl)ccc(F)c1Cl |r|