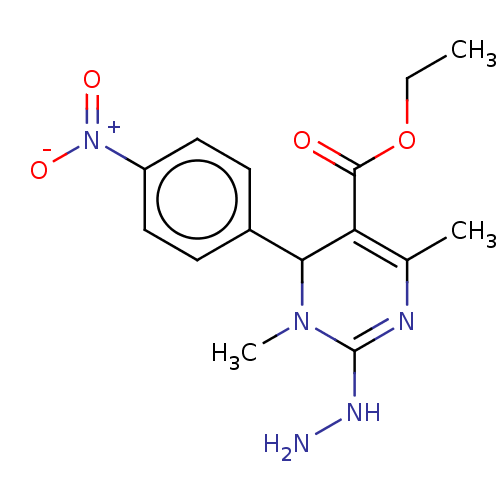
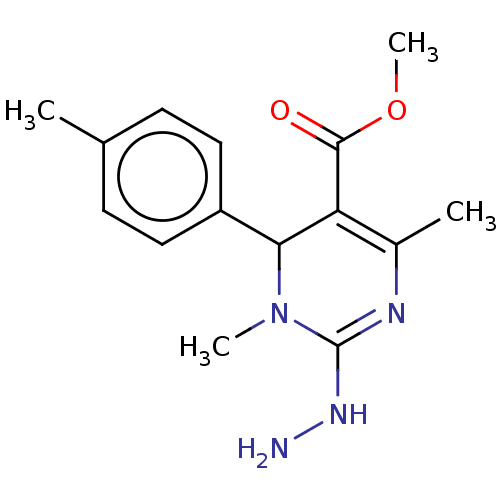
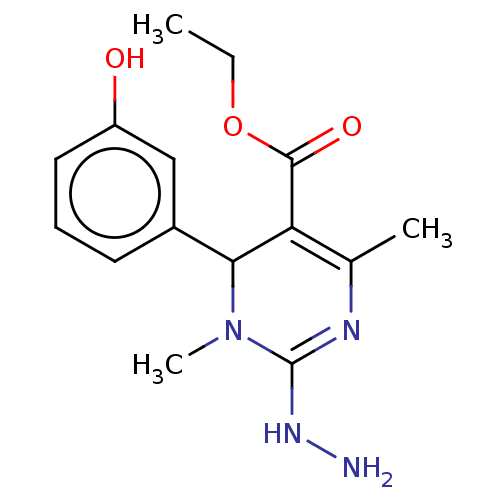
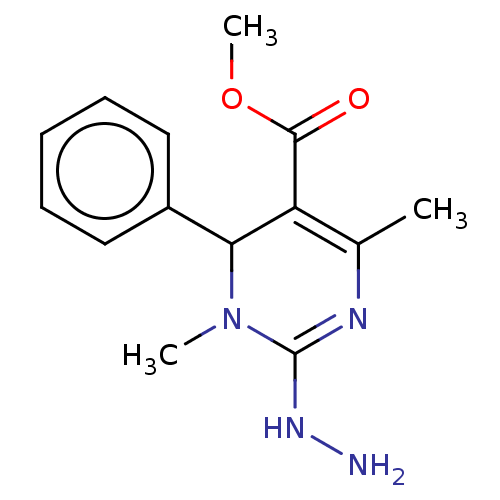
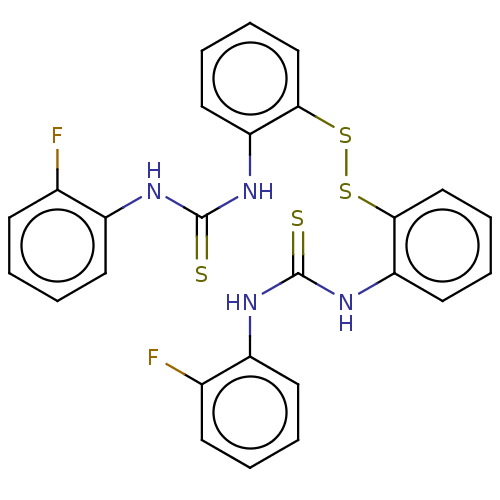
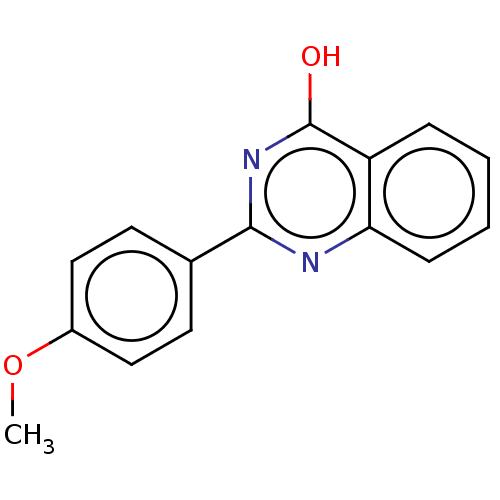
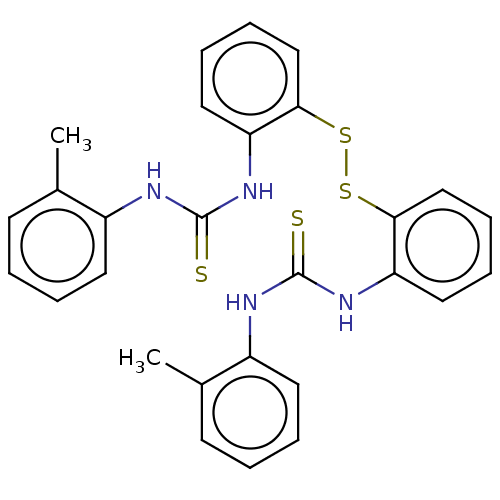
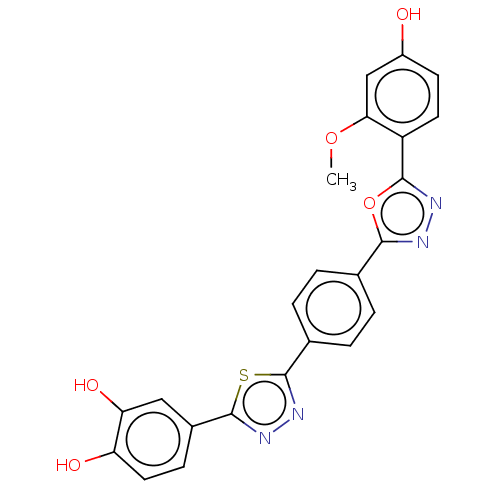
Affinity DataKi: 1.46E+4nM IC50: 1.50E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.51E+4nM IC50: 1.78E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.53E+4nM IC50: 1.74E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.58E+4nM IC50: 3.56E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+4nM IC50: 4.29E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.76E+4nM IC50: 2.08E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.94E+4nM IC50: 4.05E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 1.99E+4nM IC50: 2.36E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM IC50: 2.10E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+4nM IC50: 4.13E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 2.28E+4nM IC50: 3.60E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 2.57E+4nM IC50: 3.47E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataKi: 2.94E+4nM IC50: 2.60E+4nMAssay Description:Reaction mixture consisting of 25 �L of Jack bean (Canavalia ensiformis) urease (1 unit/well), 55 �L of 100 mM urea dissolved in phosphate buffer (4 ...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.7Assay Description:Total volume of the reaction mixture was 100 �L containing 60 �L, Na2HPO4 buffer, 50 mM and pH 7.7. Ten �L test compound 0.5 mM well-1 was added foll...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:β-Glucuronidase activity was determined by measuring the absorbance at 405 nm of p-nitrophenol formed from the substrate by the spectrophotometr...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:β-Glucuronidase activity was determined by measuring the absorbance at 405 nm of p-nitrophenol formed from the substrate by the spectrophotometr...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm), Puncak Alam Campus
Universiti Teknologi Mara (Uitm), Puncak Alam Campus
Affinity DataIC50: 165nMAssay Description:The 135 �L of 50 mM phosphate saline buffer pH (6.8), solution of enzyme (20 �L), and 20 �L of test sample with 70% DMSO was added to 96-well plate. ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:β-Glucuronidase activity was determined by measuring the absorbance at 405 nm of p-nitrophenol formed from the substrate by the spectrophotometr...More data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:β-Glucuronidase activity was determined by measuring the absorbance at 405 nm of p-nitrophenol formed from the substrate by the spectrophotometr...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Compound was tested for TXA2 receptor antagonism by measuring its ability to inhibit contraction of rat aorta induced by the stable TXA2 agonist U-46...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 850nMpH: 7.7Assay Description:Total volume of the reaction mixture was 100 �L. It contained 60 �L Na2HPO4 buffer with concentration of 50 mM and pH 7.7. 10 �L test compound (0.5 m...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Inhibition of human beta-glucuronidase pre-incubated for 30 mins before p-nitrophenyl-beta-D-glucuronide substrate addition by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Compound was tested for TXA2 receptor antagonism by measuring its ability to inhibit contraction of rat aorta induced by the stable TXA2 agonist U-46...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of human beta-glucuronidase pre-incubated for 30 mins before p-nitrophenyl-beta-D-glucuronide substrate addition by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMpH: 6.8 T: 2°CAssay Description:To understand the binding modes of the compounds given in Table 1, all ligands were docked into the binding site of Urease enzyme. The top ranked con...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Antagonistic activity for inhibition of histamine H2 receptor from anesthetized ratsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMpH: 6.8 T: 2°CAssay Description:To understand the binding modes of the compounds given in Table 1, all ligands were docked into the binding site of Urease enzyme. The top ranked con...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of jack bean urease using urea as substrate assessed as ammonia production incubated for 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+3nMpH: 7.0Assay Description:This assay was modified from Berthelot assay and was employed for the determination of urease activity. The assay is based on the hydrolysis of urea ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.86E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate after 30 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.04E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of bovine liver beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMpH: 6.8 T: 2°CAssay Description:To understand the binding modes of the compounds given in Table 1, all ligands were docked into the binding site of Urease enzyme. The top ranked con...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of bovine liver beta glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrate preincubated for 30 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+3nMAssay Description:Inhibition of bovine beta-glucuronidase assessed as p-nitrophenol formation preincubated for 30 mins followed by addition of p-nitrophenyl-beta-D-glu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:10 �L of test samples (5 mg/mL DMSO solution) were reconstituted in 100 �L of 100 mM-phosphate buffer (pH6.8) in 96-well microplate and incubated wit...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:β-glucuronidase activity was determined by measuring absorbance at 405 nm of p-nitrophenol formed substrate by spectrophotometric method. 250 �L...More data for this Ligand-Target Pair