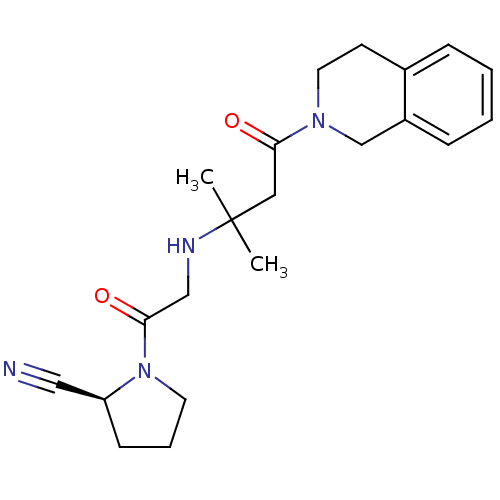
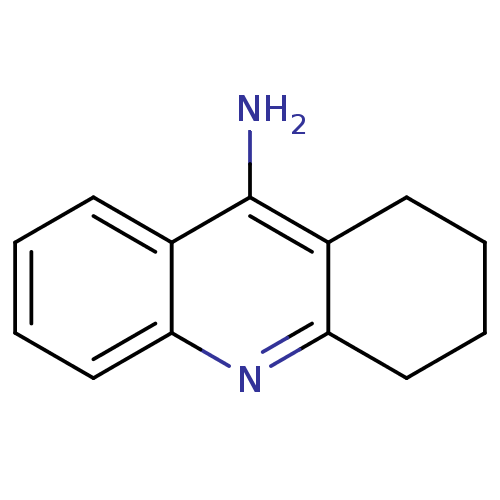
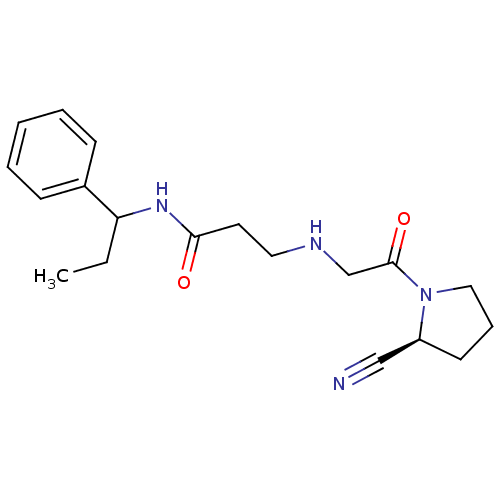
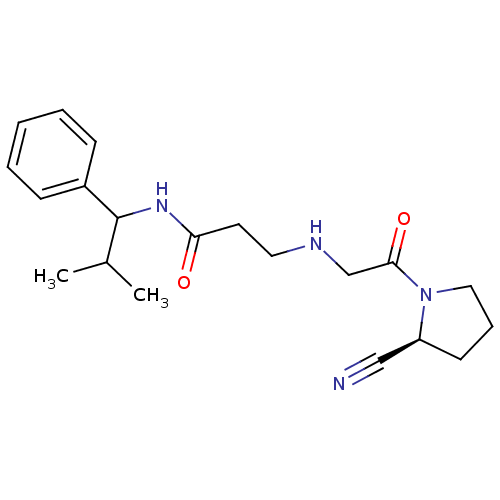
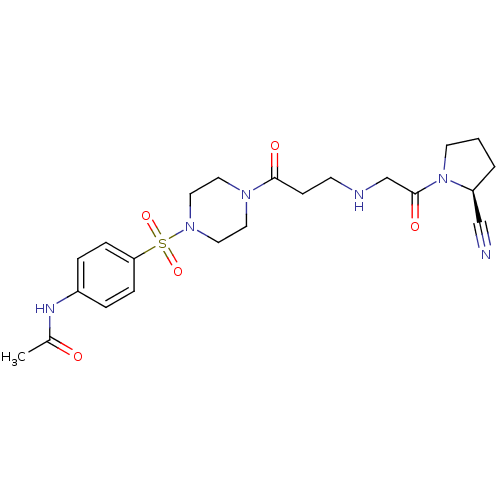
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 8.60nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant human MMP8 (Phe21 to Gly467 residues) expressed in mouse myeloma cells using Mca-Pro-LeuGly-Leu-Dpa-Ala-Arg-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant human MMP9 (Ala20 to Asp707 residues) expressed in CHO cells using Mca-Pro-LeuGly-Leu-Dpa-Ala-Arg-NH2 as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
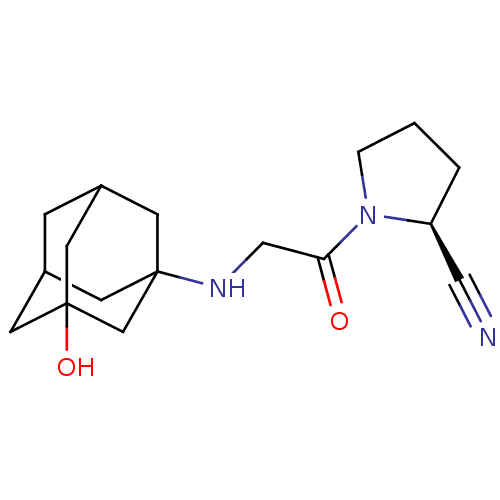
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
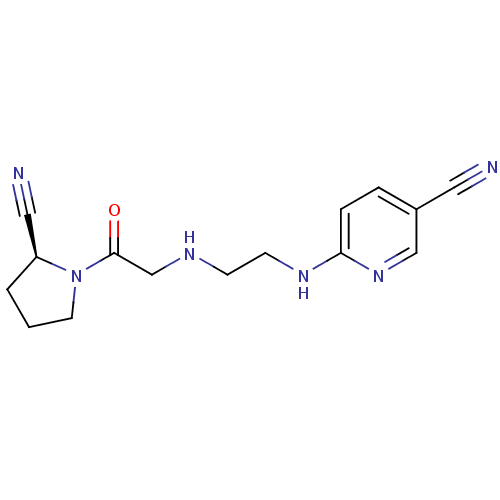
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
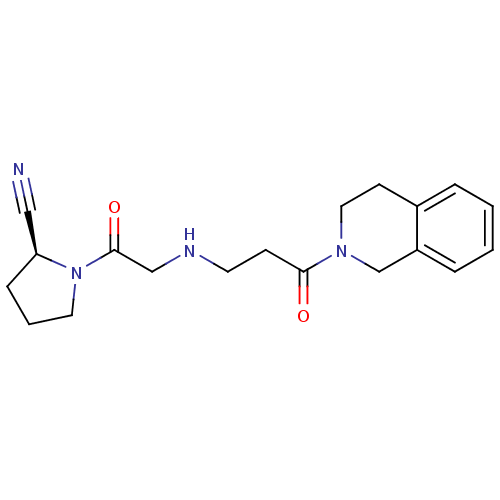
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant human MMP3 preincubated for 30 mins followed by McaPro-Leu-Gly-Leu-Dpa-Ala-Arg-NH2 substrate addition measured by fluoresce...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant human MMP3 (Tyr18 to Cys477 residues) expressed in mouse myeloma cells using Mca-Pro-LeuGly-Leu-Dpa-Ala-Arg-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of recombinant human MMP9 preincubated for 30 mins followed by McaPro-Leu-Gly-Leu-Dpa-Ala-Arg-NH2 substrate addition measured by fluoresce...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of recombinant human MMP2 preincubated for 30 mins followed by McaPro-Leu-Gly-Leu-Dpa-Ala-Arg-NH2 substrate addition measured by fluoresce...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 116nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 119nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 132nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of human AChE by Ellman's assayMore data for this Ligand-Target Pair
Affinity DataIC50: 202nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 298nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 317nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 369nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Inhibition of recombinant human MMP3 (Tyr18 to Cys477 residues) expressed in mouse myeloma cells using Mca-Pro-LeuGly-Leu-Dpa-Ala-Arg-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 428nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 447nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 452nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 527nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 564nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 629nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 651nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 676nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 784nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 811nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 855nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of recombinant human MMP3 (Tyr18 to Cys477 residues) expressed in mouse myeloma cells using Mca-Pro-LeuGly-Leu-Dpa-Ala-Arg-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+3nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 1.36E+3nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+3nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+3nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+3nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
TargetProteasome subunit beta type-5(Homo sapiens (Human))
Guangxi Normal University
Curated by ChEMBL
Guangxi Normal University
Curated by ChEMBL
Affinity DataIC50: 1.97E+3nMAssay Description:Inhibition of chymotrypsin-like activity of human 20S proteasome using Suc-LLVY-AMC as substrate by fluorometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 2.12E+3nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair


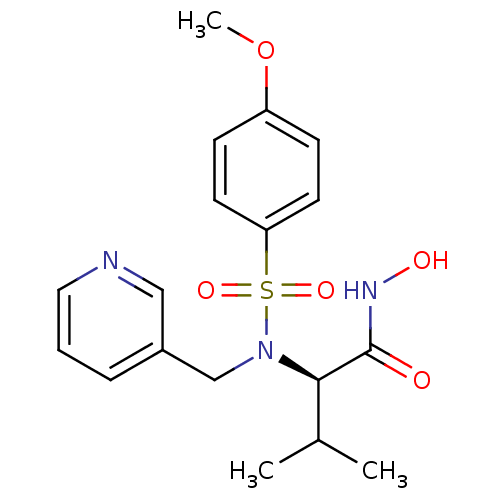




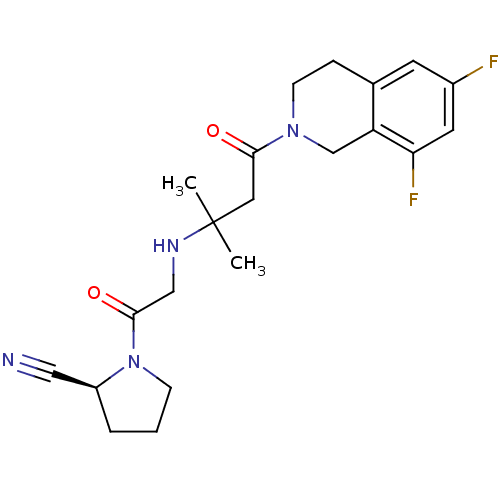
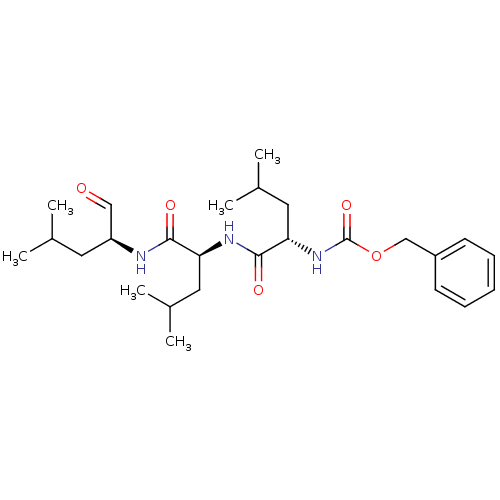
 3D Structure (crystal)
3D Structure (crystal)