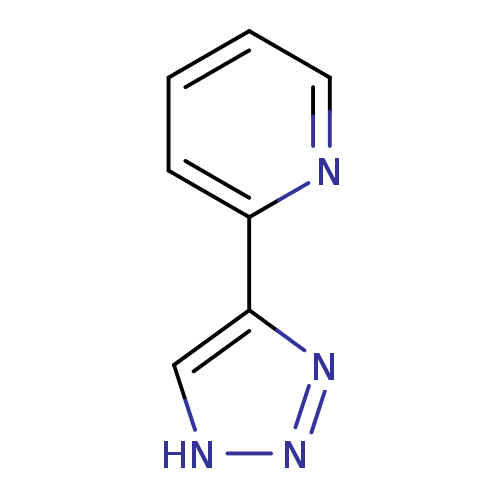
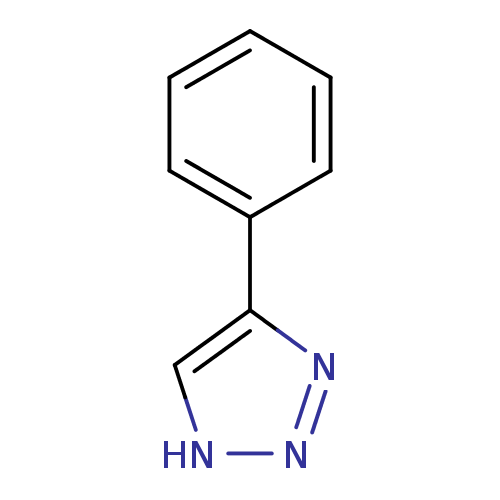
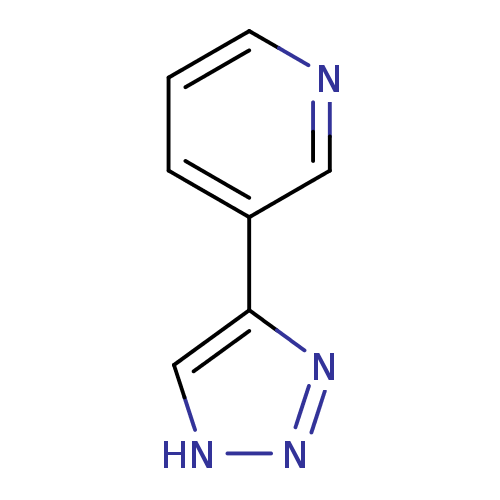
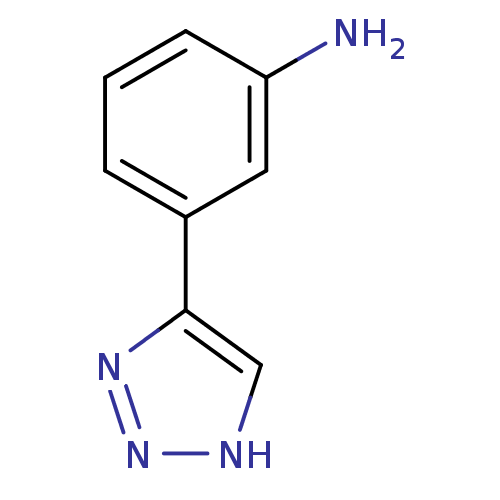
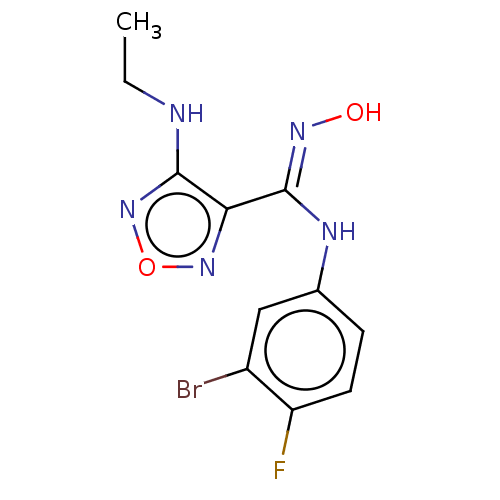
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 15nM ΔG°: -44.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -44.0kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 18nM ΔG°: -43.8kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 29nM ΔG°: -42.6kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -42.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 41nM ΔG°: -41.7kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 52nM ΔG°: -41.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 52nM ΔG°: -41.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 70nM ΔG°: -40.4kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 112nM ΔG°: -39.3kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 150nM ΔG°: -38.6kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 260nM ΔG°: -37.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 280nM ΔG°: -37.0kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 280nM ΔG°: -37.0kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 300nM ΔG°: -36.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 337nM ΔG°: -36.6kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-33.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-33.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-33.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nM ΔG°: >-33.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition of N-terminal his-tagged human indoleamine 2,3-dioxygenase expressed in Escherichia coli assessed as N'-formylkynurenine formation by spec...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+4nMAssay Description:Competitive inhibition of IDO1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 7.40nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in IFN-gamma-stimulated human HeLa cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in IFN-gamma-stimulated human HeLa cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in IFN-gamma-stimulated human HeLa cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in IFN-gamma-stimulated human HeLa cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Inhibition of IDO1 in IFN-gamma-stimulated human HeLa cells assessed as decrease in kynurenine levels after 48 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in mouse B16 cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of human N-terminal His-tagged IDO1 expressed in Escherichia coli using D-Trp as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:Inhibition of indoleamine 2,3-dioxygenase in IFN-gamma-stimulated human HeLa cells assessed as kynurenine formation by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of N-terminal his-tagged human indoleamine 2,3-dioxygenase expressed in Escherichia coli assessed as N'-formylkynurenine formation by spec...More data for this Ligand-Target Pair
Affinity DataIC50: 67nMAssay Description:Inhibition of N-terminal his-tagged human indoleamine 2,3-dioxygenase expressed in Escherichia coli assessed as N'-formylkynurenine formation by spec...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:Inhibition of N-terminal his-tagged human indoleamine 2,3-dioxygenase expressed in Escherichia coli assessed as N'-formylkynurenine formation by spec...More data for this Ligand-Target Pair


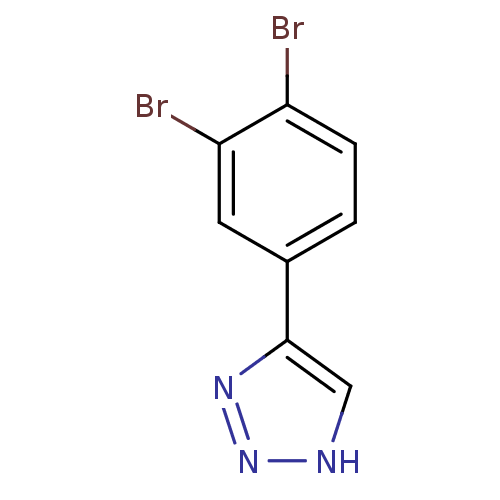


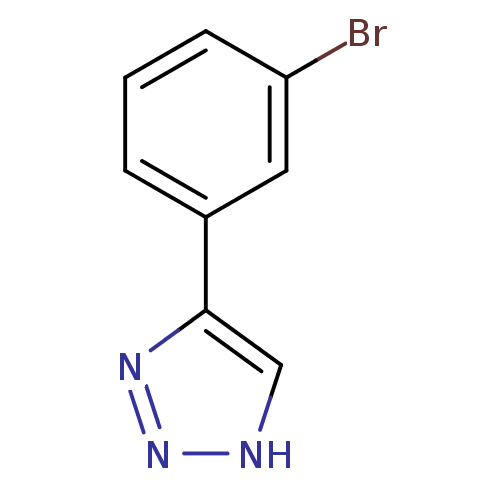
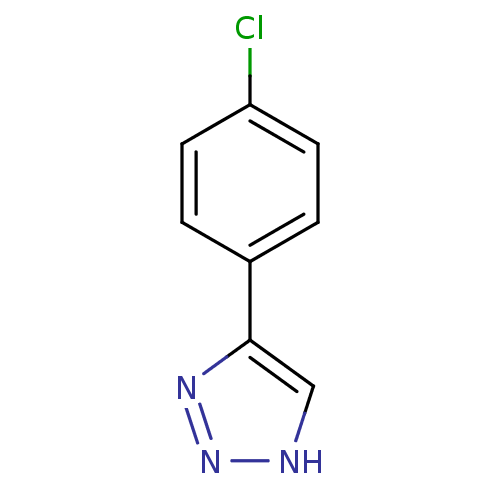




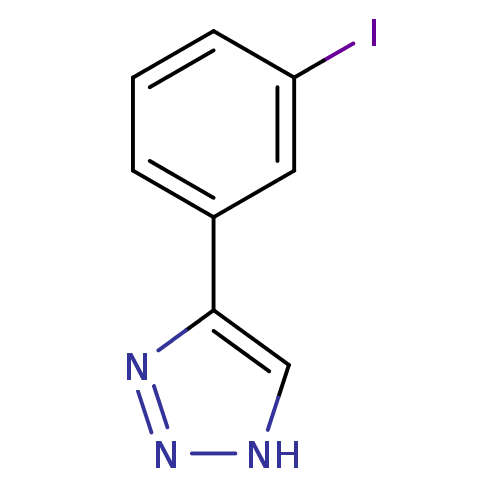
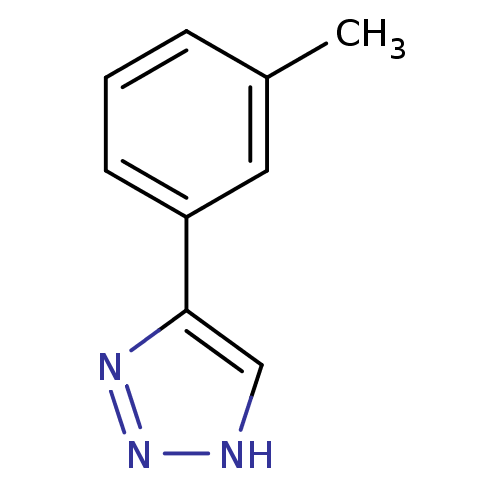
 3D Structure (crystal)
3D Structure (crystal)