Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 14nM ΔG°: -45.6kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -44.7kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 28nM ΔG°: -43.8kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 32nM ΔG°: -43.5kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 150nM ΔG°: -39.6kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 168nM ΔG°: -39.3kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 190nM ΔG°: -39.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 190nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.9kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 240nM ΔG°: -38.4kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 340nM ΔG°: -37.5kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 380nM ΔG°: -37.3kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 570nM ΔG°: -36.2kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -36.1kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.7kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.7kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 820nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.63E+3nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 2.04E+3nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 2.42E+3nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 4.41E+3nMAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+3nM ΔG°: >-30.8kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+3nM ΔG°: >-30.8kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.0kJ/molepH: 7.4 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 200nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 19nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 50nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 3.65E+3nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 3.70E+3nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 60nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: >1.00E+4nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 2.64E+4nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 40nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 20nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 610nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 2.40E+3nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 47nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Homo sapiens (Human))
Universita Del Piemonte Orientale
Universita Del Piemonte Orientale
Affinity DataEC50: 23nMAssay Description:The effect of the substances on Ca2+ influx was determined by using HEK-293 cells stably overexpressing recombinant human TRPV1 cDNA. The cells were ...More data for this Ligand-Target Pair


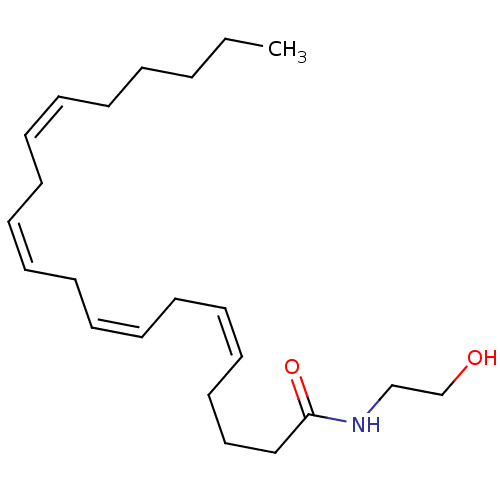
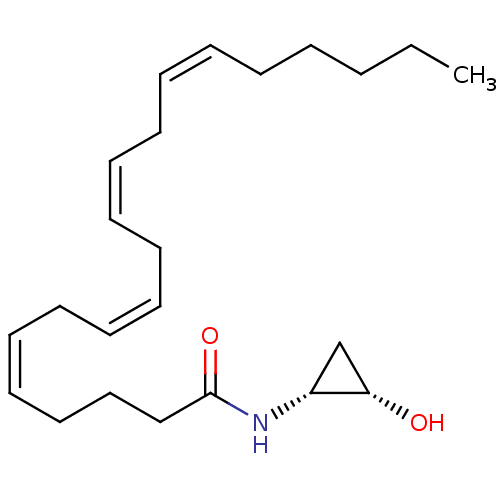
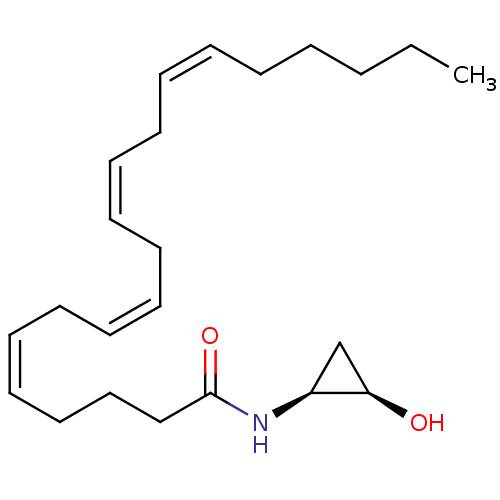
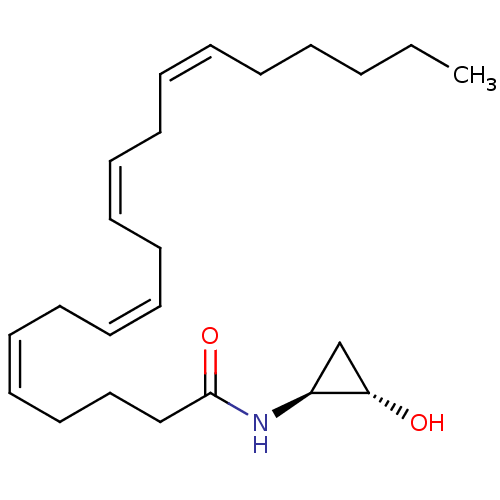
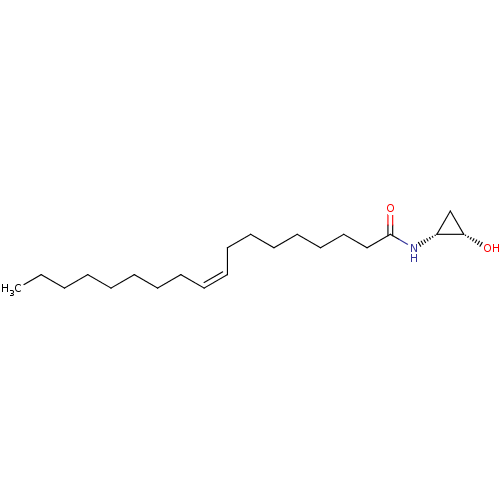
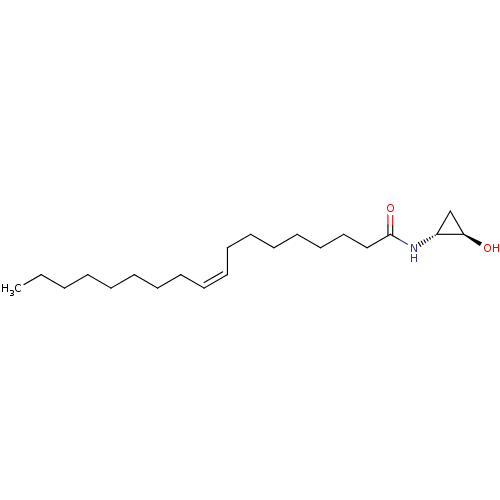
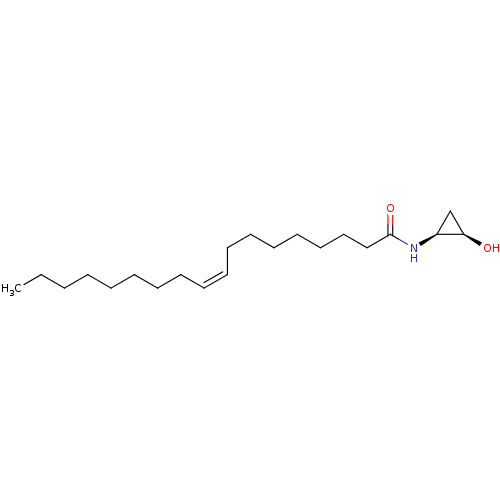
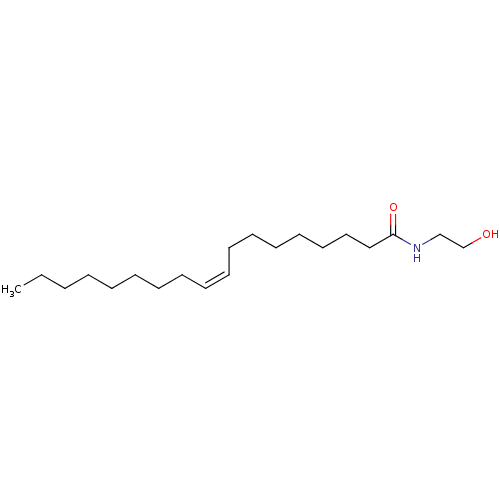
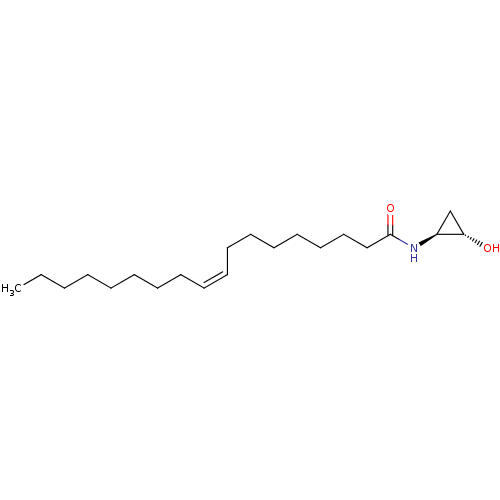
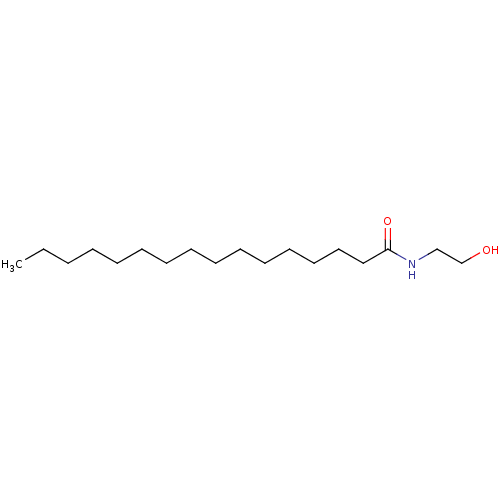
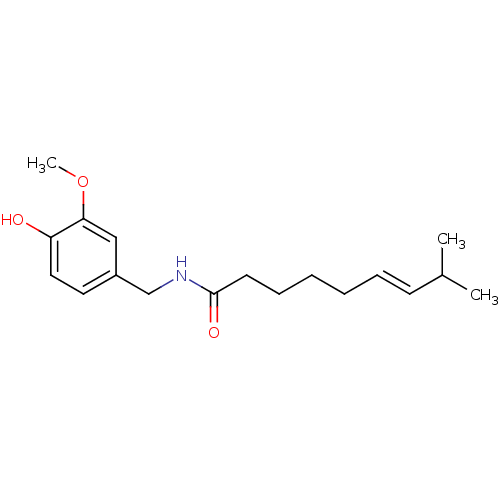
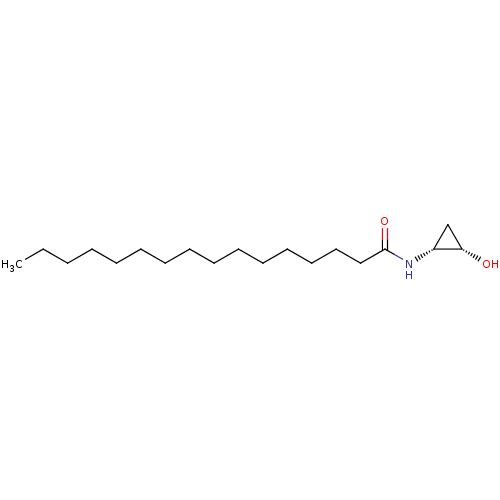

 3D Structure (crystal)
3D Structure (crystal)