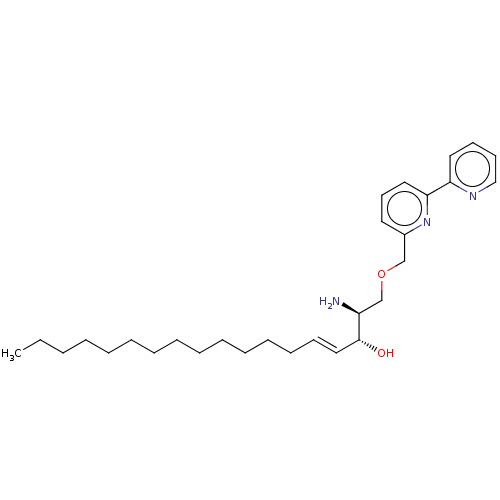
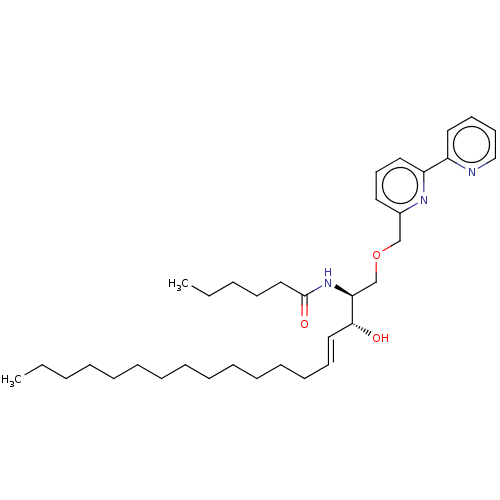
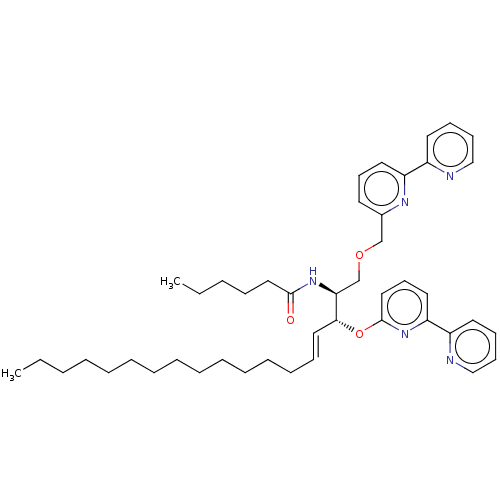
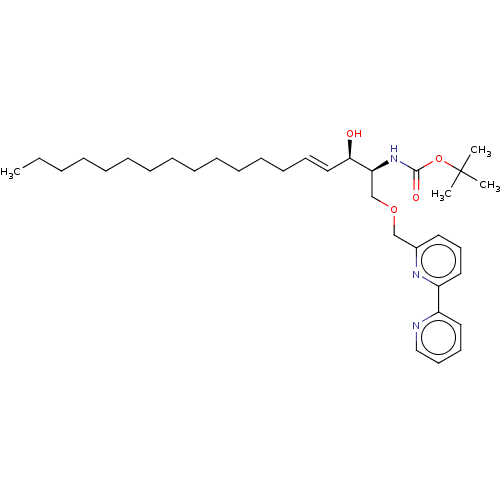
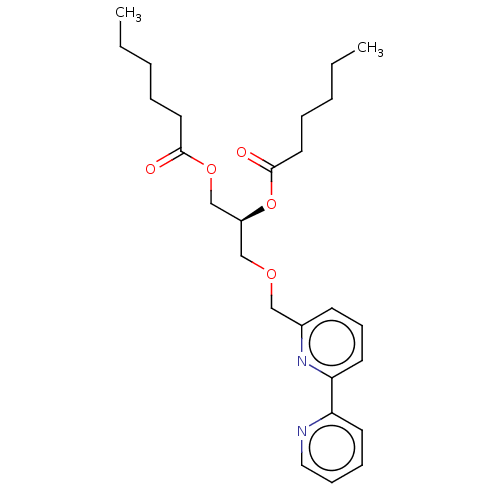
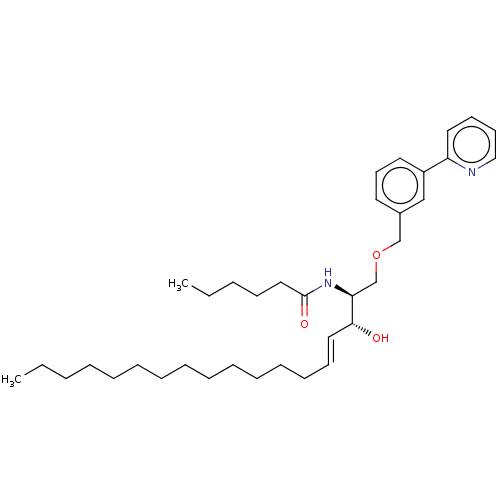
Affinity DataIC50: 800nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nM ΔG°: -34.9kJ/mole IC50: 800nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+3nM ΔG°: -33.0kJ/mole IC50: 900nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+3nM ΔG°: -31.4kJ/mole IC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nMAssay Description:Competitive inhibition of Bacillus cereus SMaseC assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+3nMAssay Description:Competitive inhibition of Bacillus cereus SMaseC assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+3nMAssay Description:Competitive inhibition of Bacillus cereus SMaseC assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 1.83E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+4nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+4nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+5nMAssay Description:Inhibition of Bacillus cereus SMaseCMore data for this Ligand-Target Pair