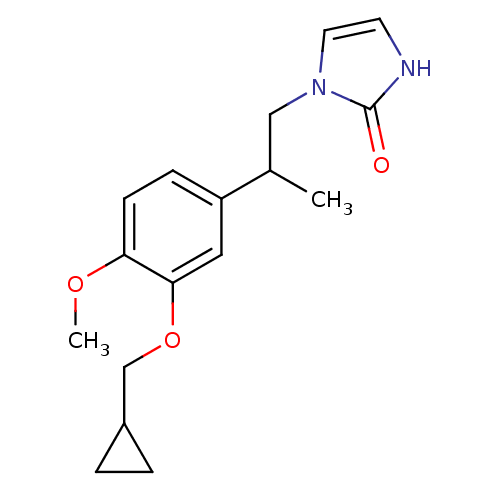
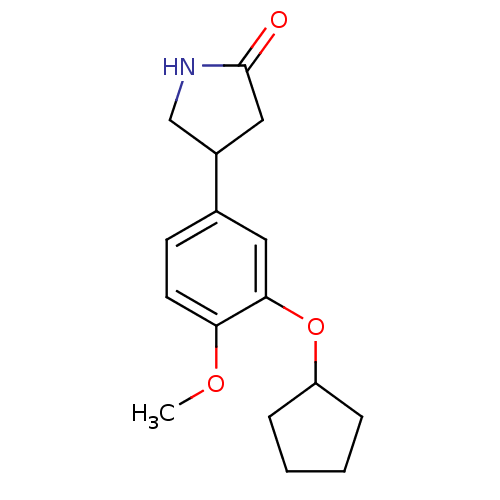
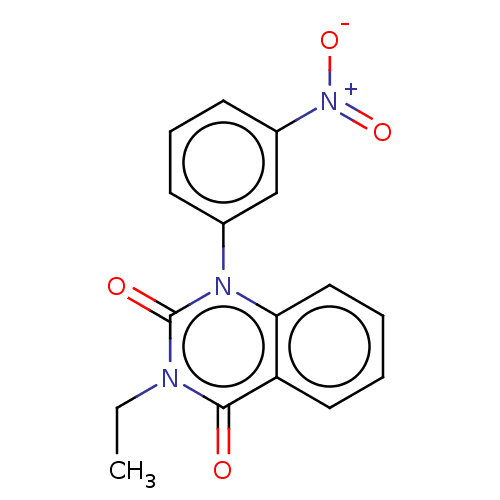
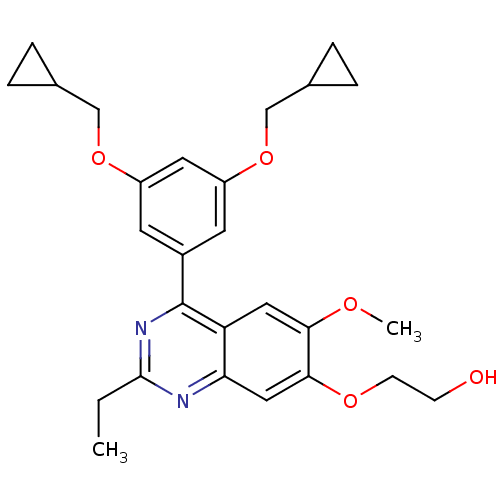
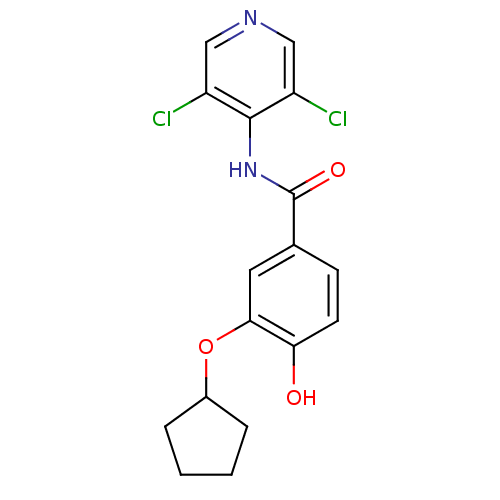
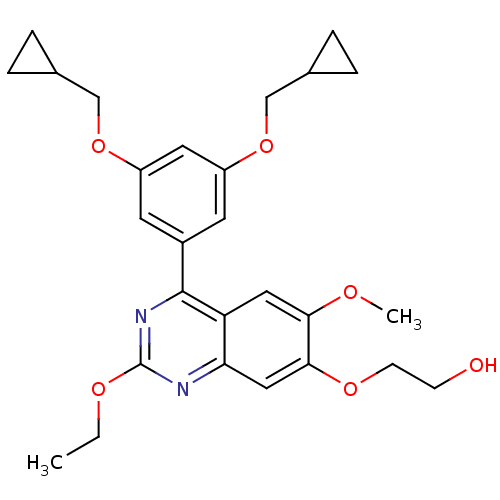
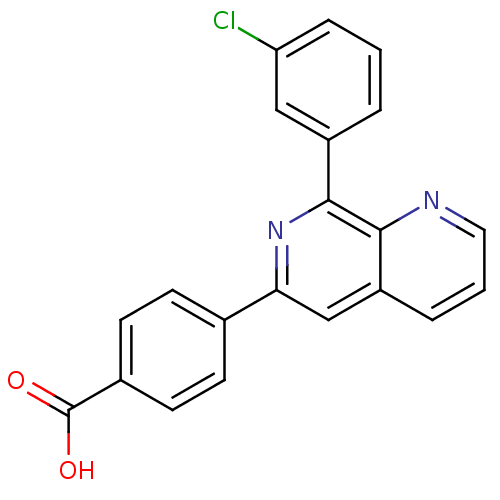
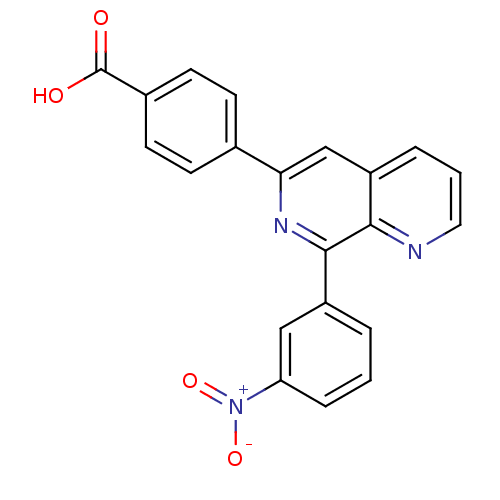
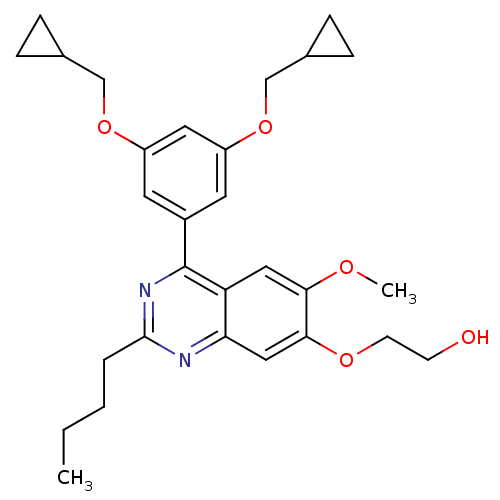
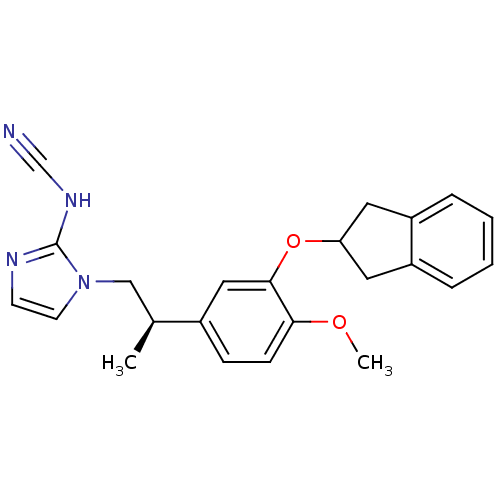
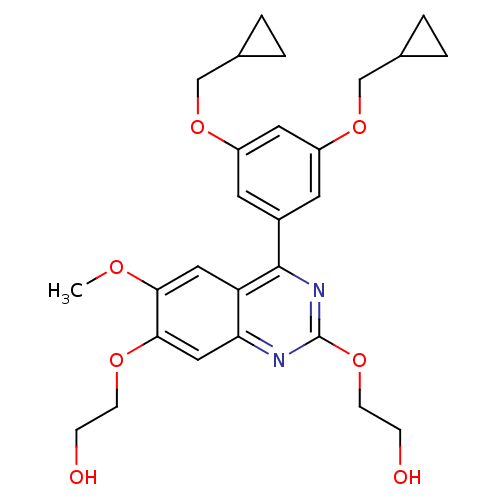
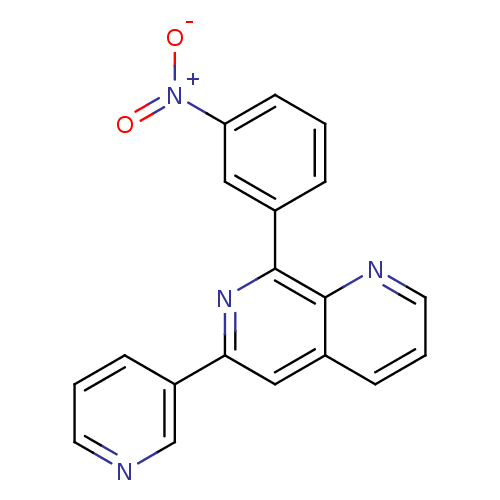
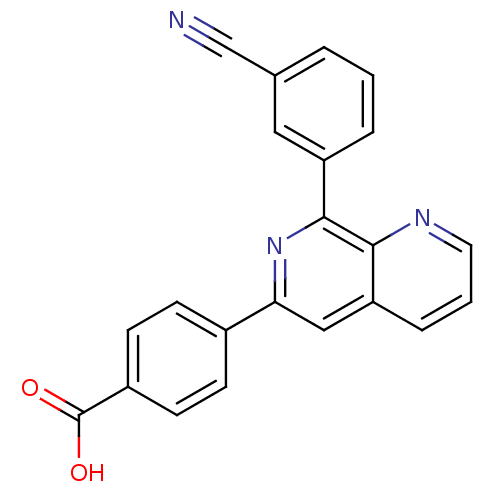
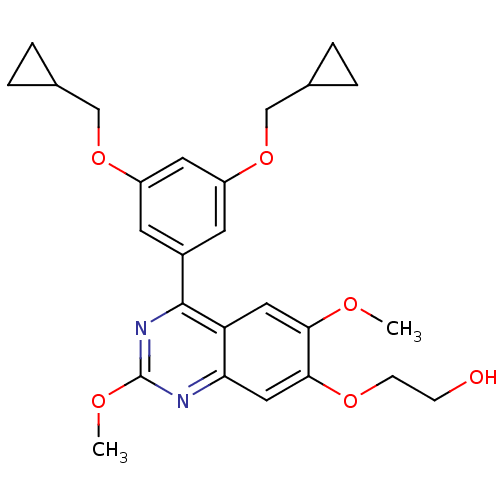
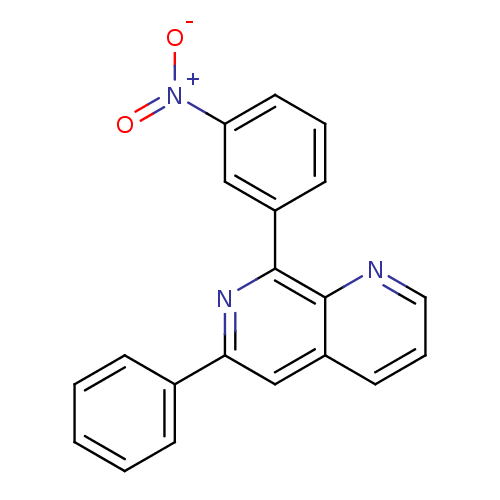
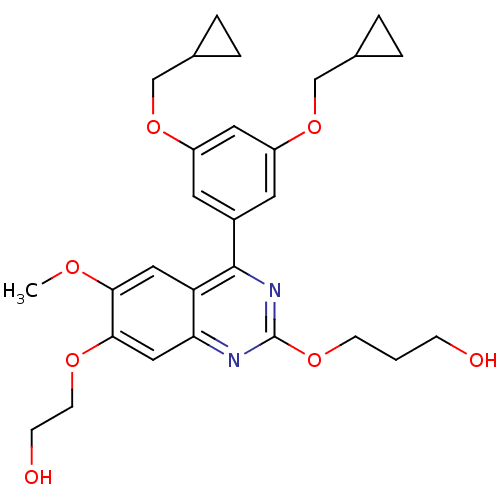
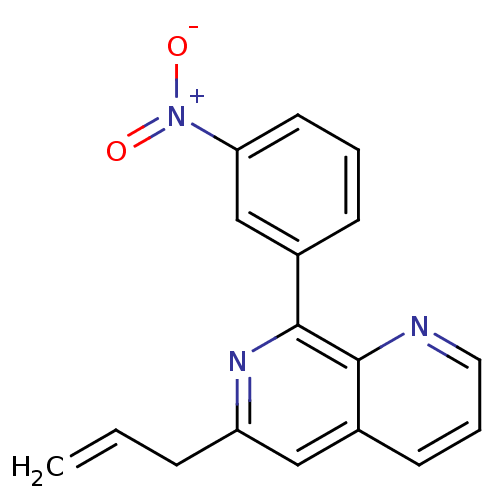
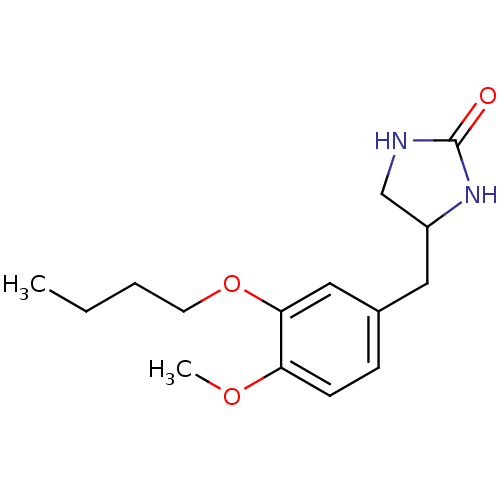
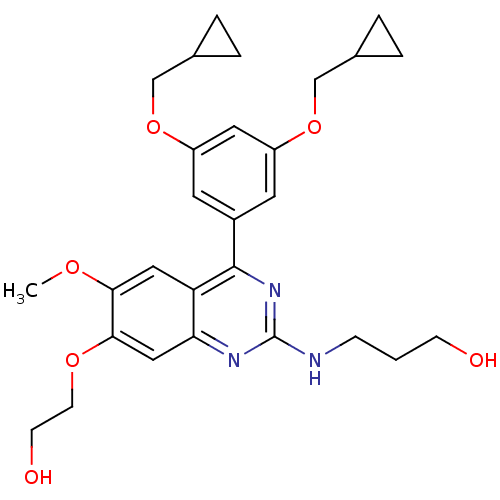
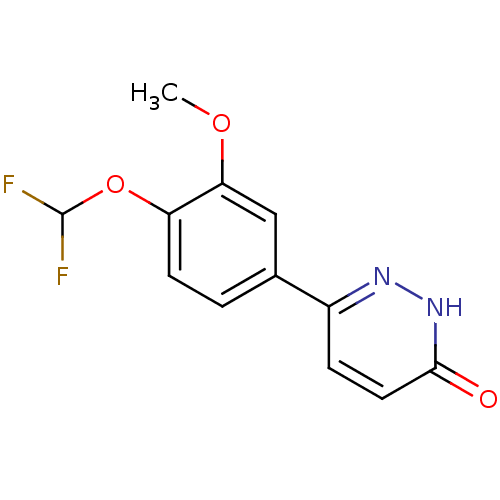
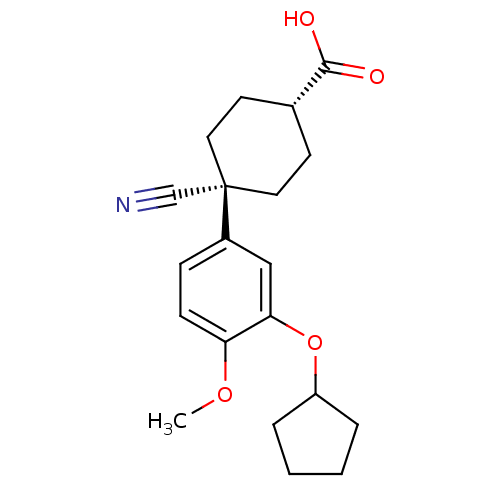
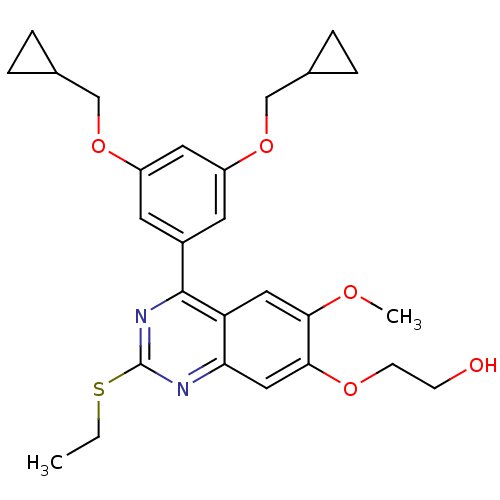
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of rat PDE4BMore data for this Ligand-Target Pair
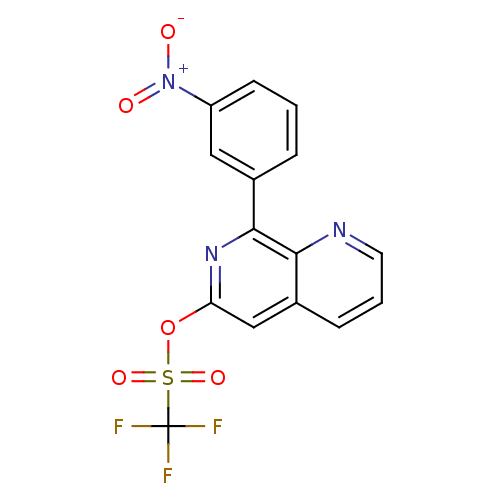
Affinity DataIC50: 0.0251nMAssay Description:Inhibition of rat PDE4BMore data for this Ligand-Target Pair
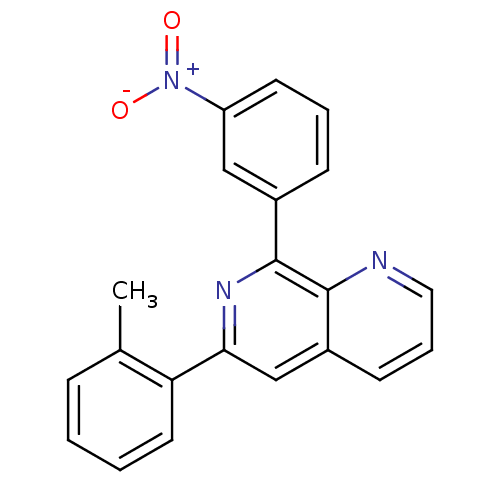
Affinity DataKi: 3.5nMAssay Description:Displacement of [3H]rolipram from rat forebrain membraneMore data for this Ligand-Target Pair
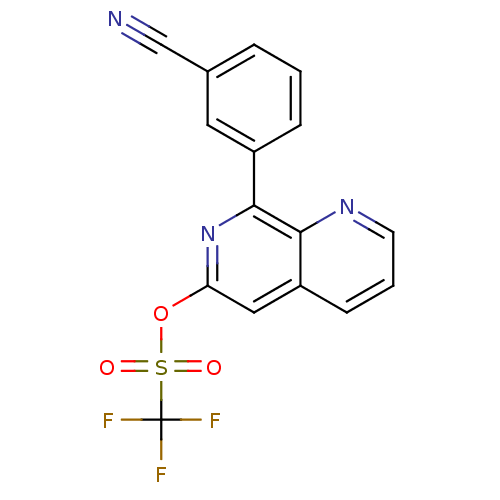
Affinity DataIC50: 5.70nMAssay Description:Inhibition of phosphodiesterase (PDE) 4BMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Displacement of [3H]rolipram from high affinity binding site (HARBS) in rat brain membrane.More data for this Ligand-Target Pair
Affinity DataIC50: 7.70nMAssay Description:Inhibition of phosphodiesterase (PDE) 4BMore data for this Ligand-Target Pair
Affinity DataIC50: 8.30nMAssay Description:Inhibition of phosphodiesterase (PDE) 4BMore data for this Ligand-Target Pair
Affinity DataIC50: 8.60nMAssay Description:Inhibition of phosphodiesterase (PDE) 4BMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]rolipram from high affinity binding site (HARBS) in rat brain membrane.More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 of rat forebrain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Evaluated for the inhibition of [3H]rolipram binding to membrane-bound PDE4.More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 of rat forebrain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Evaluated for the inhibition of [3H]rolipram binding to membrane-bound PDE4.More data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of [3H]-rolipram binding to membrane-bound PDE4More data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Displacement of [3H]rolipram from rat forebrain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 81nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 85nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 145nMAssay Description:Evaluated for the inhibition of [3H]rolipram binding to membrane-bound PDE4.More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Displacement of [3H]rolipram from rat forebrain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Evaluated for the inhibition of [3H]rolipram binding to membrane-bound PDE4.More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Inhibitory concentration against phosphodiesterase 4 (PDE4) from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Evaluated for the inhibition of [3H]rolipram binding to membrane-bound PDE4.More data for this Ligand-Target Pair
Affinity DataKi: 220nMAssay Description:Displacement of [3H]rolipram from rat forebrain membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 420nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:Evaluated for its ability to inhibit PDE4B.More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Inhibitory activity against Phosphodiesterase 4B (PDE4B) from rat source expressed in Saccharomyces cerevisiaeMore data for this Ligand-Target Pair

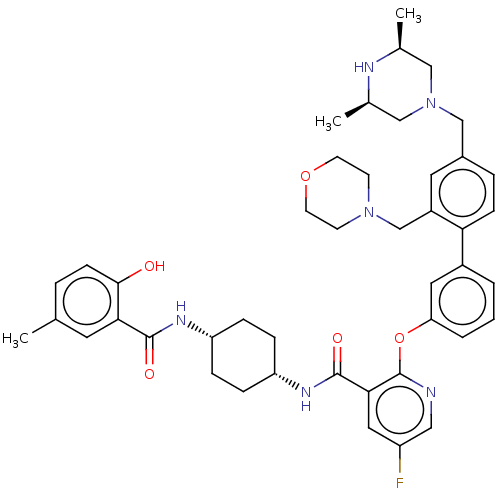
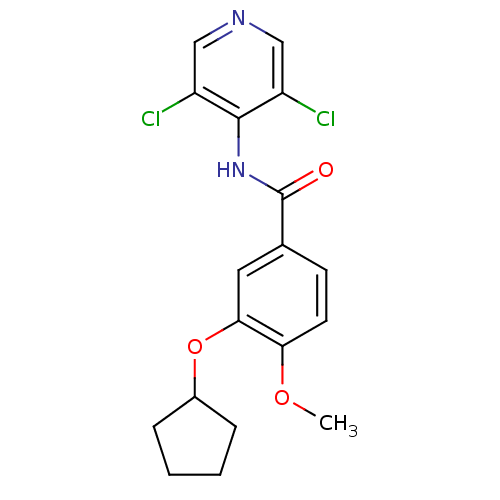
 3D Structure (crystal)
3D Structure (crystal)