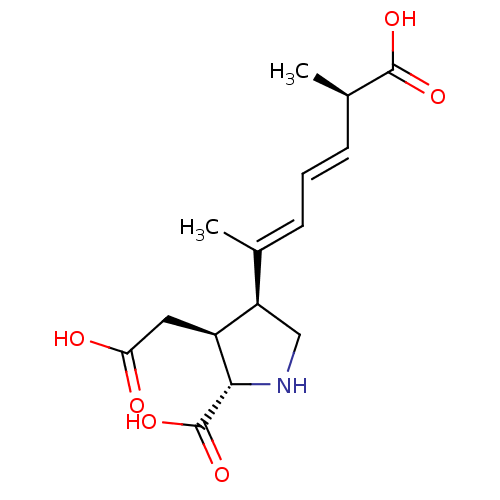
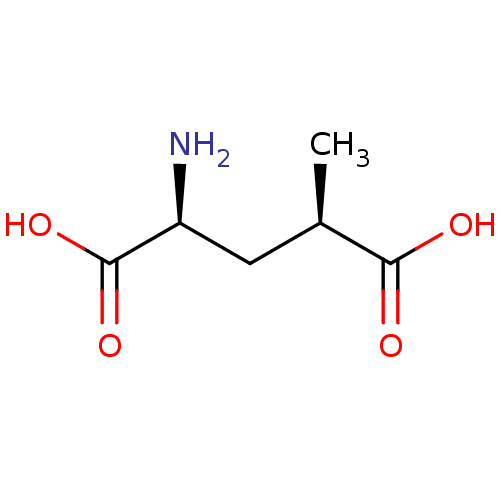
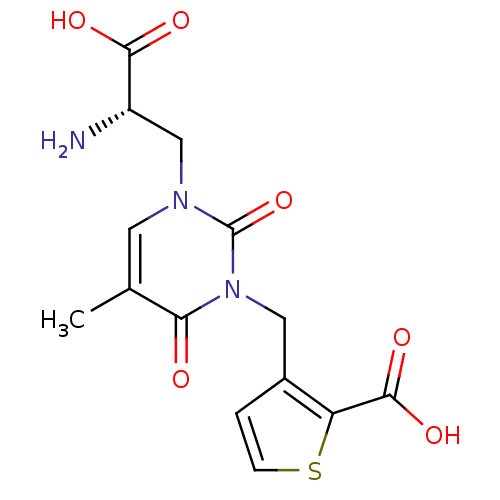
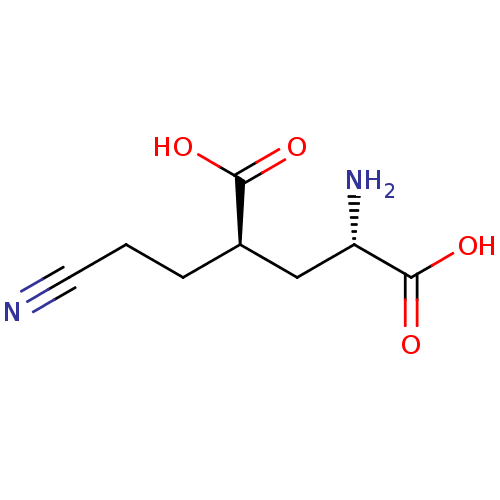
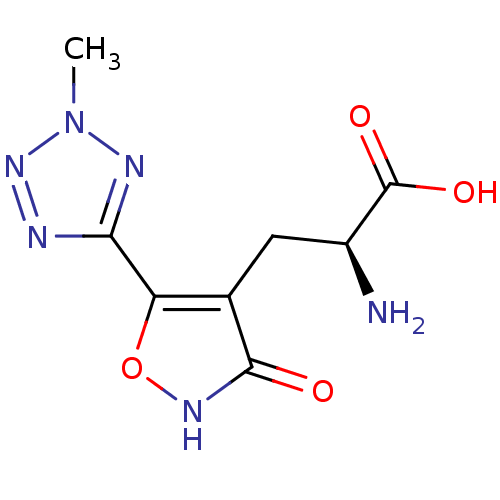
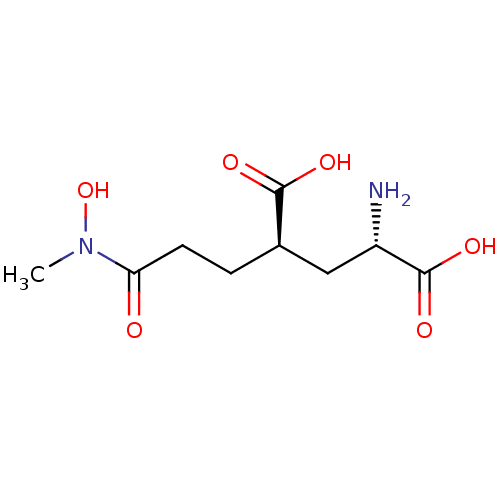
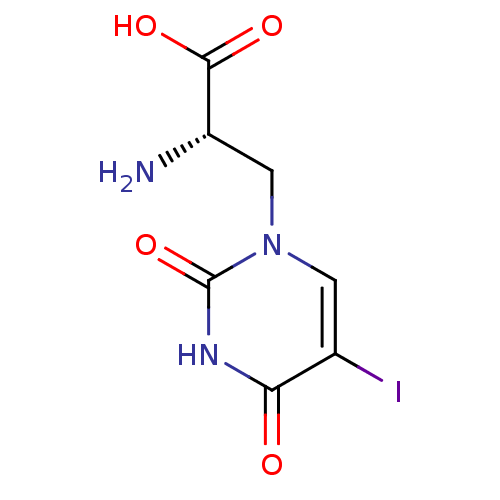
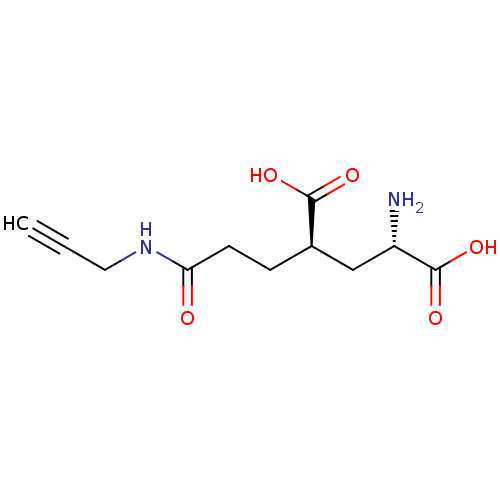
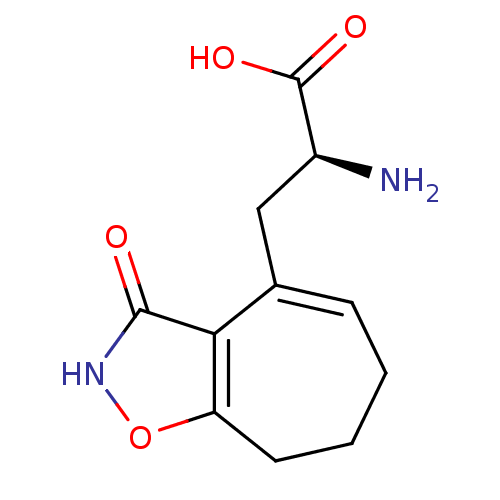
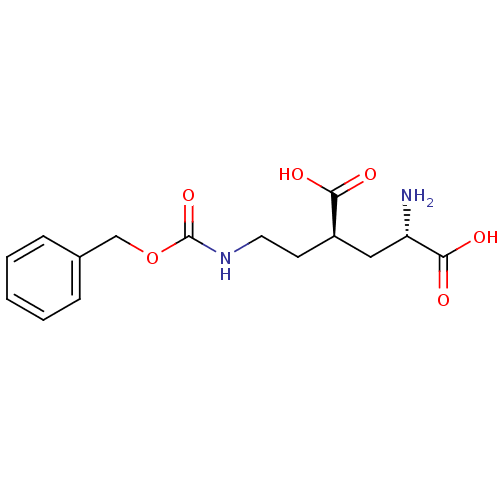
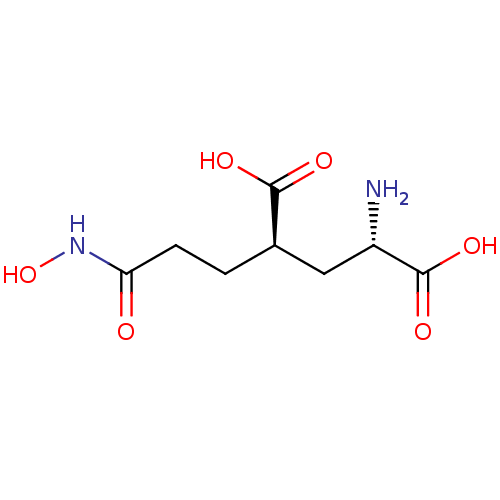
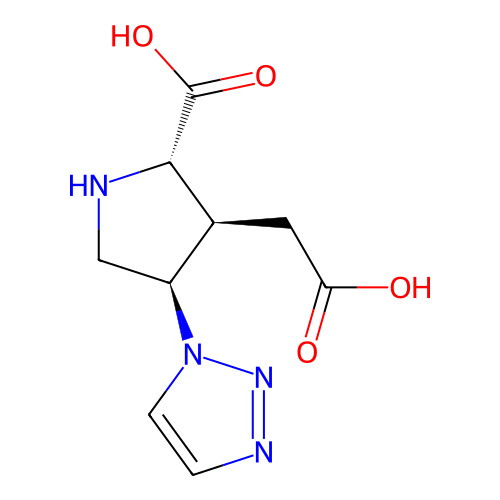
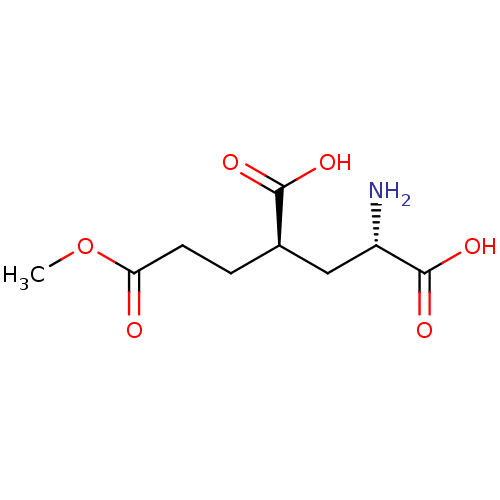
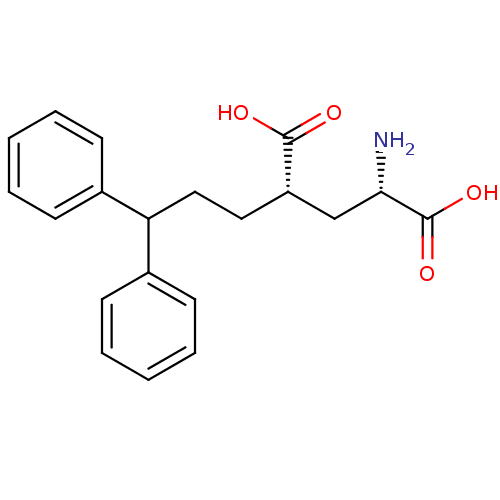
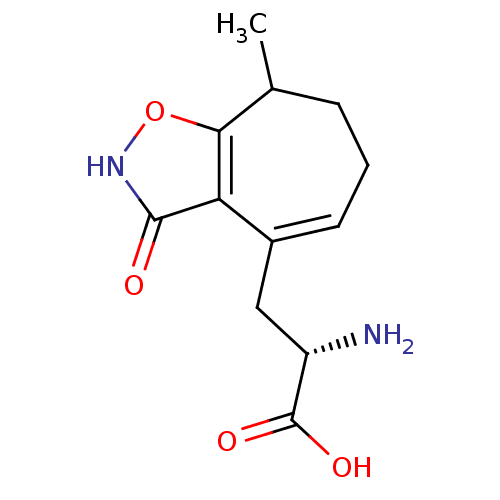
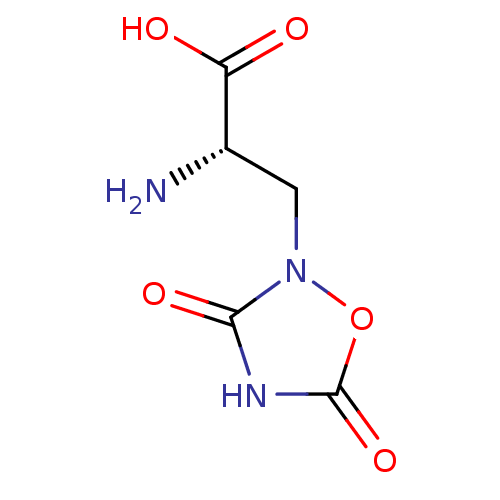
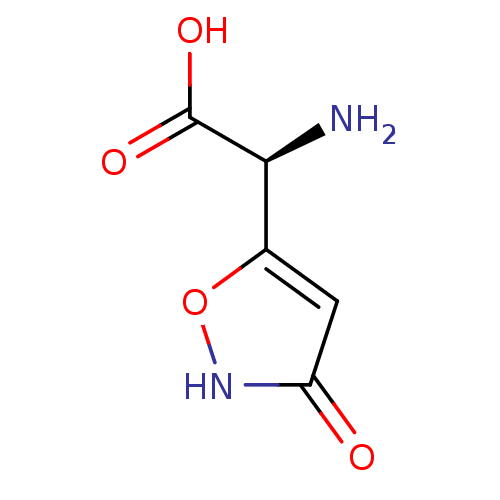
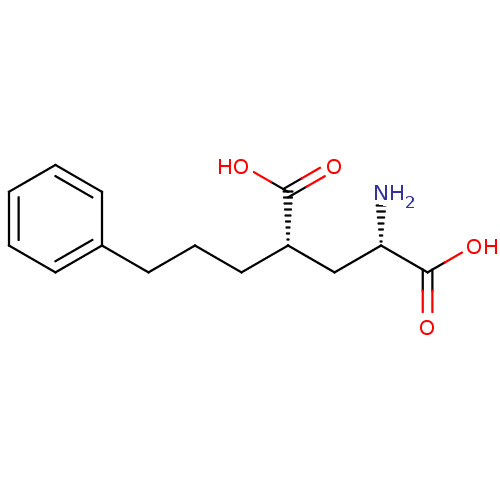
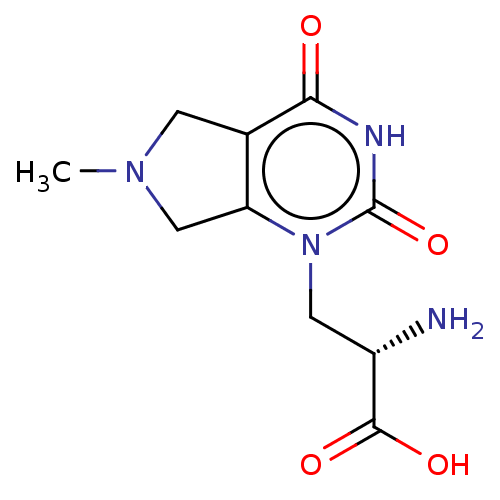
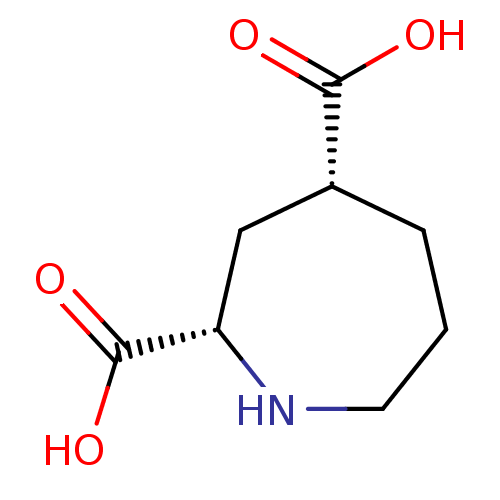
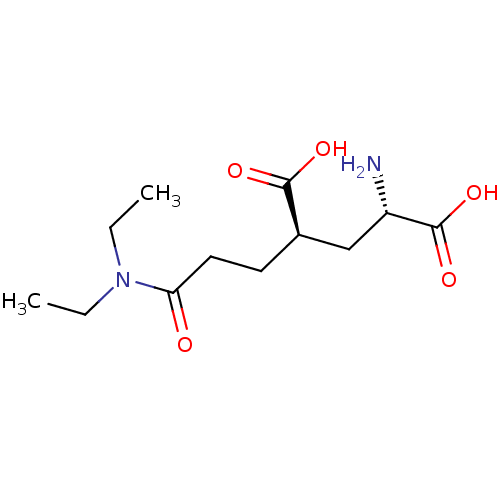
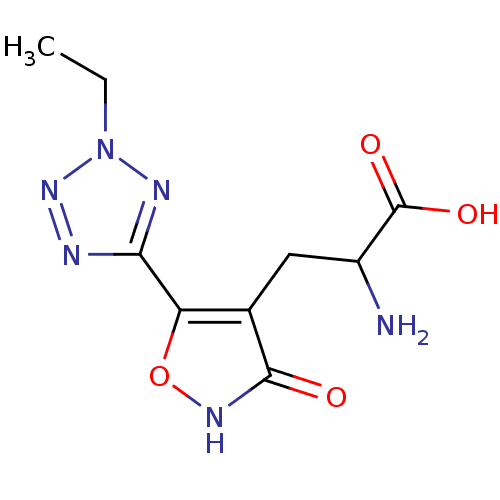
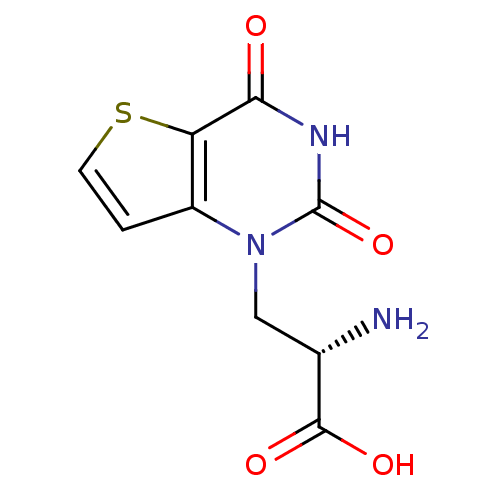
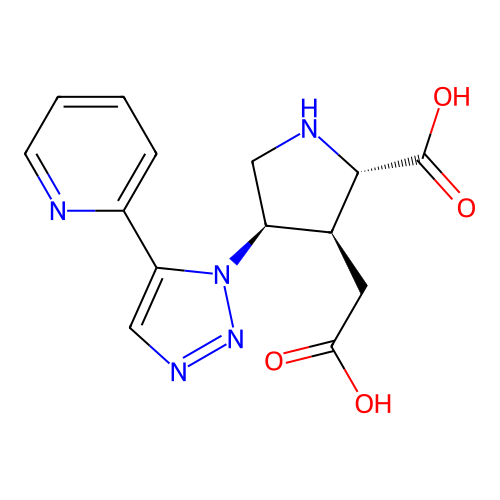
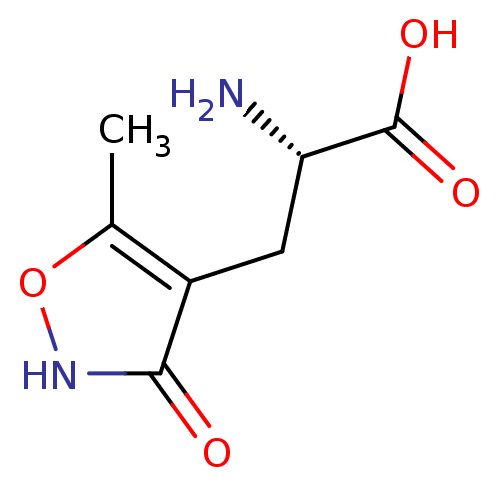
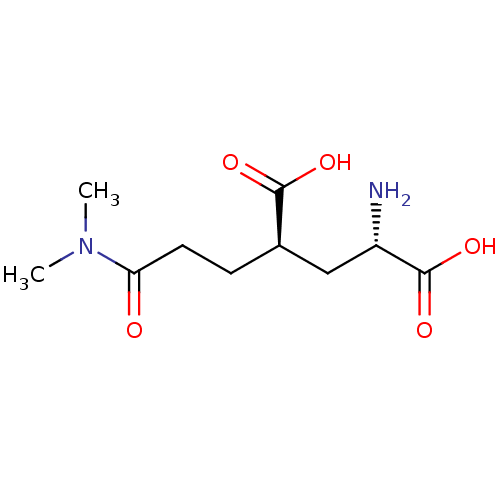
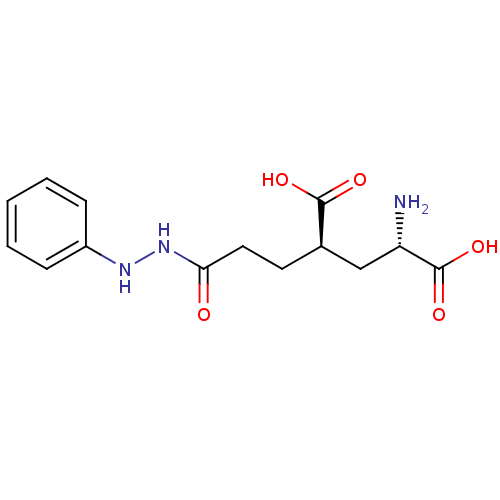
Affinity DataKi: 3.84nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 5.69nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 32.8nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 57.3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataEC50: 110nMAssay Description:In vitro activity at Ionotropic glutamate receptor using cortical wedge assays in rat brain synaptosomaMore data for this Ligand-Target Pair
Affinity DataKi: 123nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataEC50: 140nMAssay Description:Potency on Ionotropic glutamate receptor kainate in dorsal root ganglion cellsMore data for this Ligand-Target Pair
Affinity DataKi: 185nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 199nMAssay Description:Displacement of [3H]SYM from rat recombinant GluR7A expressed in baculovirus infected Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 217nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 239nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 280nMAssay Description:Displacement of [3H]-KA from rat GluK3More data for this Ligand-Target Pair
Affinity DataEC50: 330nMAssay Description:In vitro activity at Ionotropic glutamate receptor using cortical wedge assays in rat brain synaptosomaMore data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 376nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 376nMAssay Description:Displacement of [3H]SYM2081 from rat cloned iGluR7More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Binding affinity against Ionotropic glutamate receptor using ACPD sensitive [3H]- glutamate displacement from rat forebrain preparationsMore data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:Displacement of [3H]SYM2081 from rat iGluR7 receptor expressed in Sf9 cells baculovirus system after 1 to 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 494nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 494nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Compound was tested in vitro for binding affinity against Ionotropic glutamate receptor kainate using [3H]kainate as the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.01E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 1.02E+3nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+3nMAssay Description:Binding affinity against Ionotropic glutamate receptor using ACPD sensitive [3H]- glutamate displacement from rat forebrain preparationsMore data for this Ligand-Target Pair
Affinity DataKi: 1.38E+3nMAssay Description:Displacement of [3H]SYM from rat recombinant GluR7A expressed in baculovirus infected Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.67E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 1.74E+3nMAssay Description:Displacement of [3H[SYM2081 from rat recombinant GluK3a expressed in sS9 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.84E+3nMAssay Description:Binding affinity against Ionotropic glutamate receptor using ACPD sensitive [3H]- glutamate displacement from rat forebrain preparationsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.84E+3nMAssay Description:Binding affinity against Ionotropic glutamate receptor using ACPD sensitive [3H]- glutamate displacement from rat forebrain preparationsMore data for this Ligand-Target Pair
Affinity DataKi: 2.15E+3nMAssay Description:Displacement of [3H]SYM2081 from recombinant homomeric rat GluK3 receptor expressed in baculovirus infected Sf9 insect cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 2.53E+3nMAssay Description:Displacement of [3H]SYM from rat recombinant GluR7A expressed in baculovirus infected Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.81E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 2.82E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 3.24E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR7More data for this Ligand-Target Pair
Affinity DataKi: 3.25E+3nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 3.29E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataEC50: 3.50E+3nMAssay Description:In vitro activity at Ionotropic glutamate receptor using cortical wedge assays in rat brain synaptosomaMore data for this Ligand-Target Pair
Affinity DataKi: 3.65E+3nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 3.75E+3nMAssay Description:Displacement of [3H]-KA from rat GluK3More data for this Ligand-Target Pair
Affinity DataEC50: 3.80E+3nMAssay Description:In vitro activity at Ionotropic glutamate receptor using cortical wedge assays in rat brain synaptosomaMore data for this Ligand-Target Pair
Affinity DataKi: 4.66E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 4.70E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 4.75E+3nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.09E+3nMAssay Description:Displacement of [3H]SYM2081 from recombinant homomeric rat GluK3 receptor expressed in baculovirus infected Sf9 insect cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5.09E+3nMAssay Description:Displacement of [3H]SYM2081 from rat GluK3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.39E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 5.66E+3nMAssay Description:Displacement of [3H]-(2S,4R)-4-methylglutamic acid from full length recombinant rat GluK3 receptor expressed in sf9 cells by liquid scintillation cou...More data for this Ligand-Target Pair