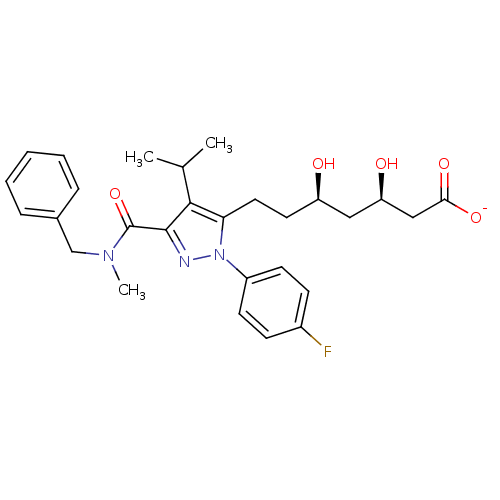
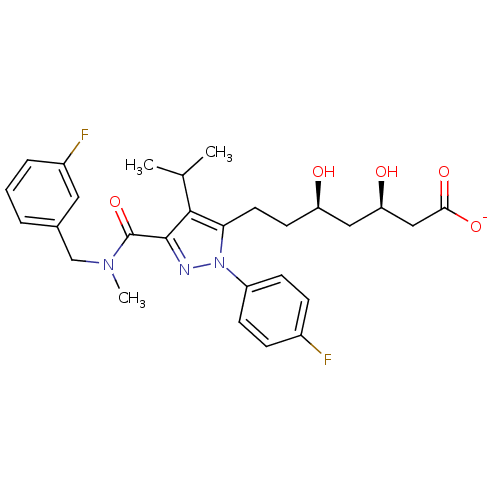
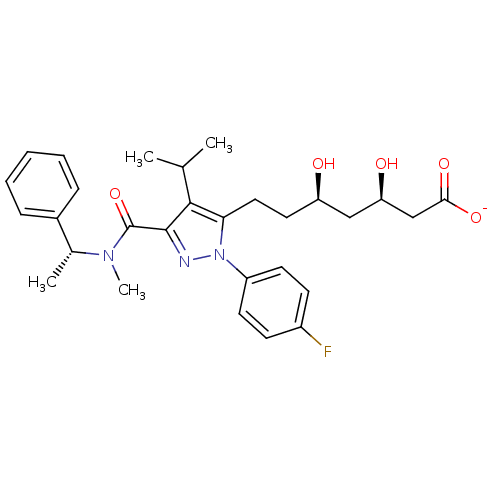
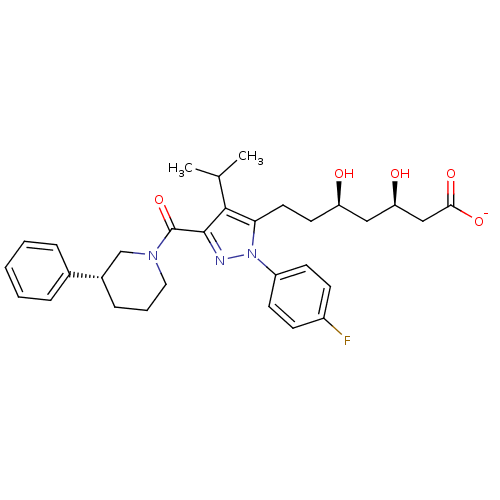
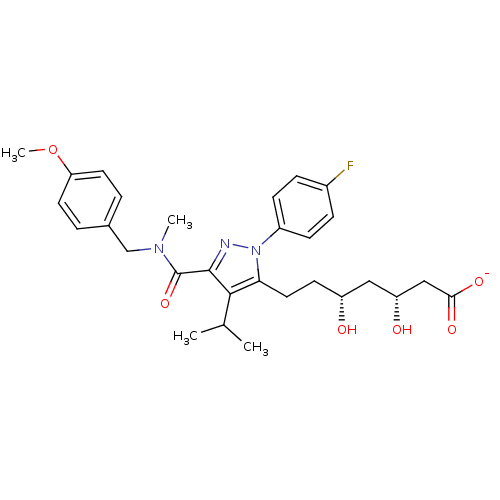
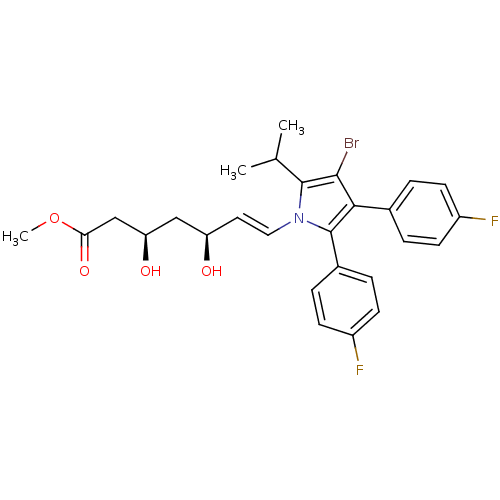
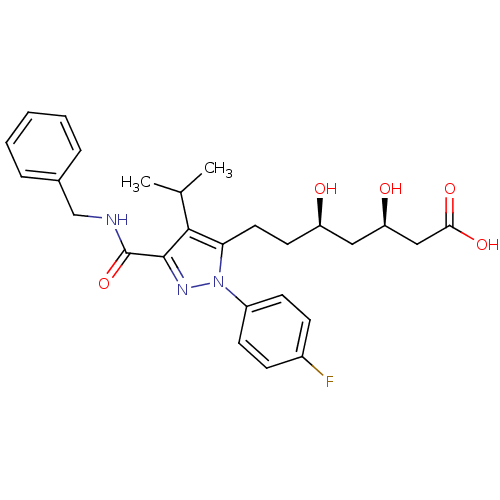
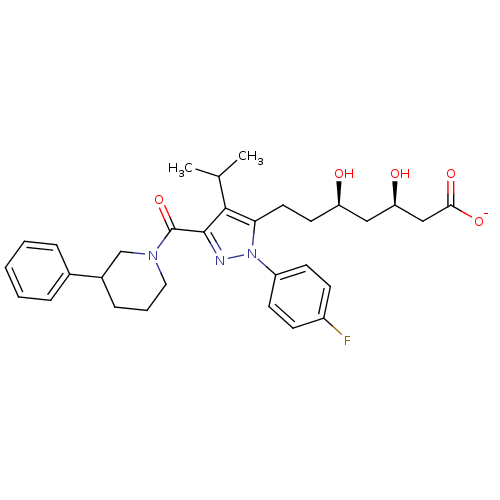
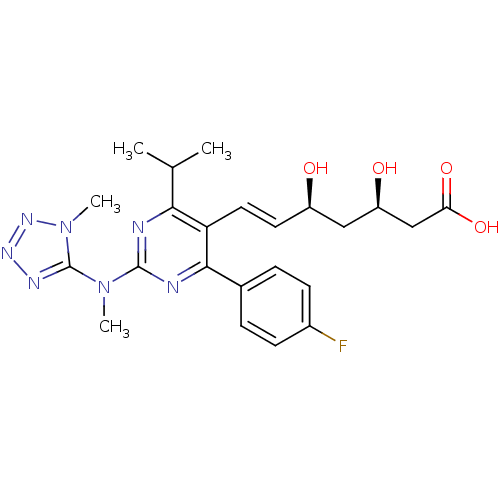
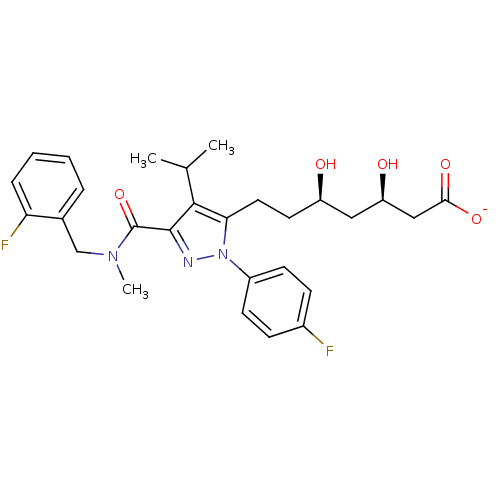
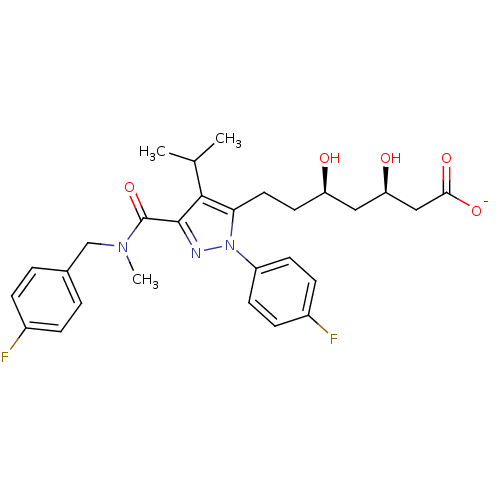
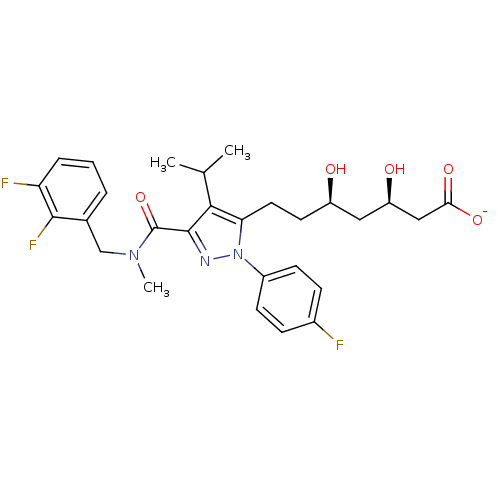
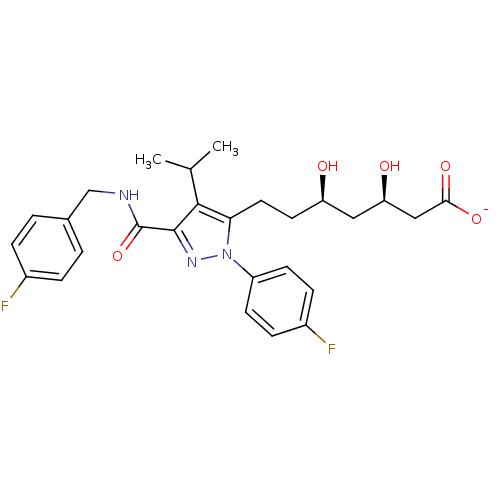
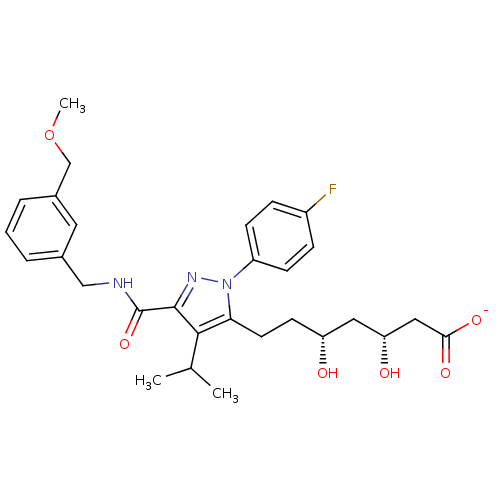
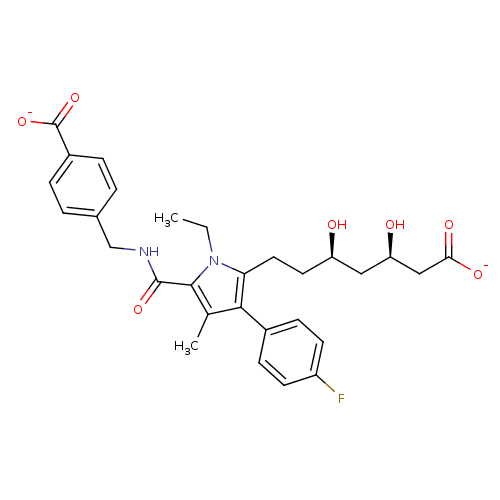
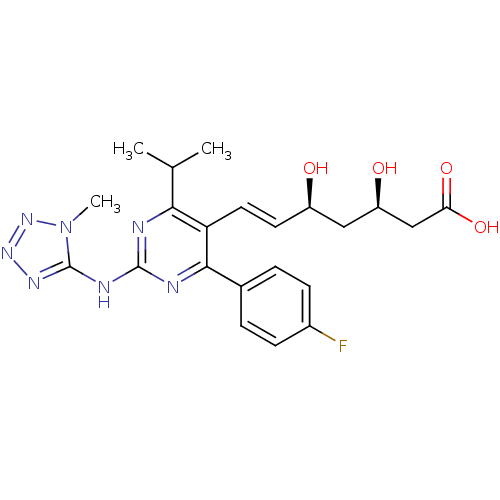
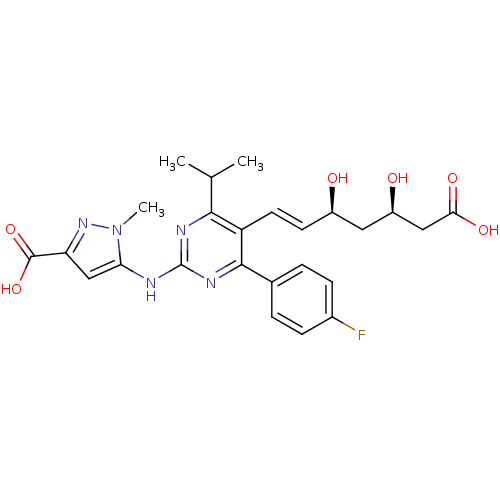
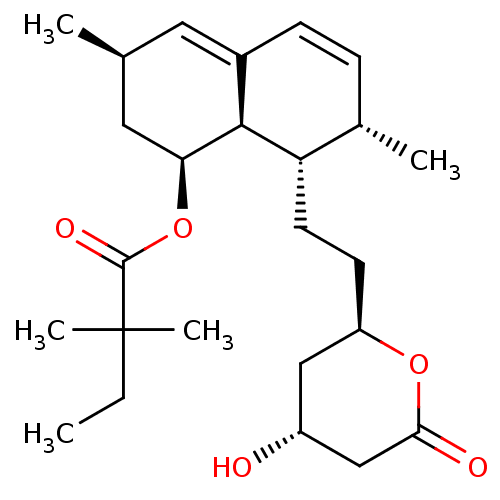
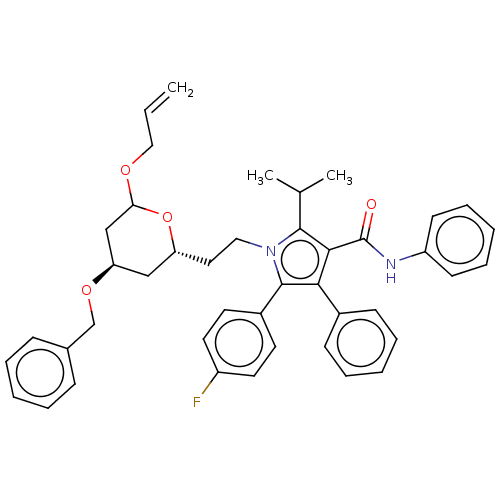
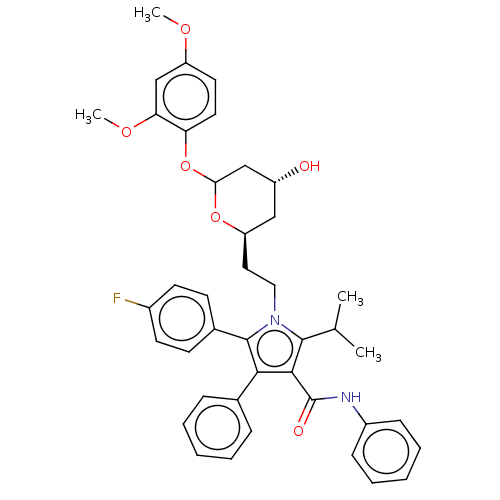
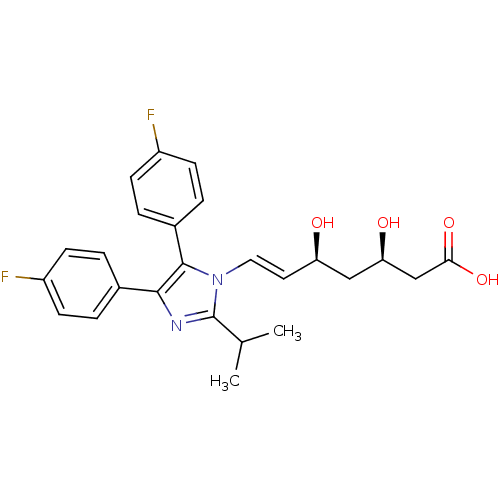
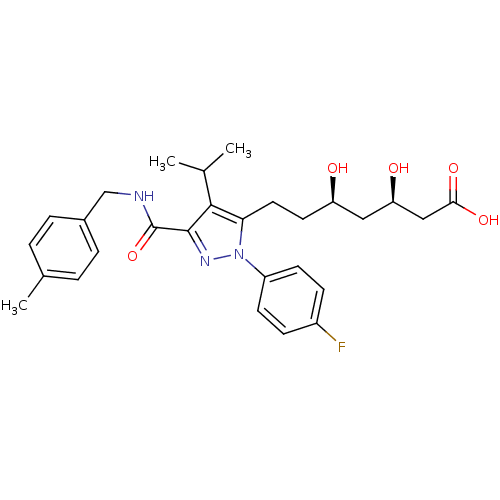
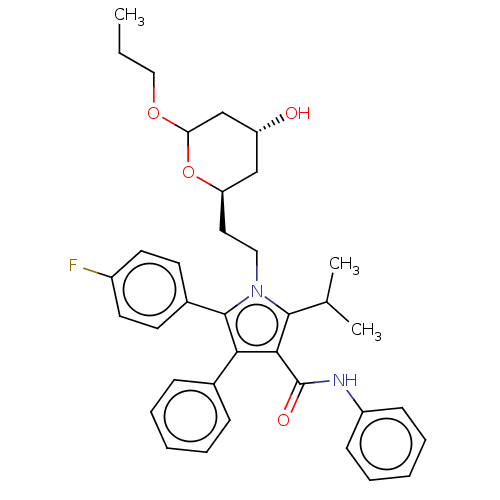
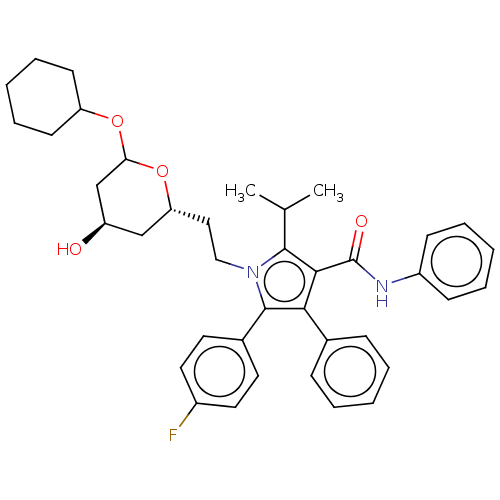
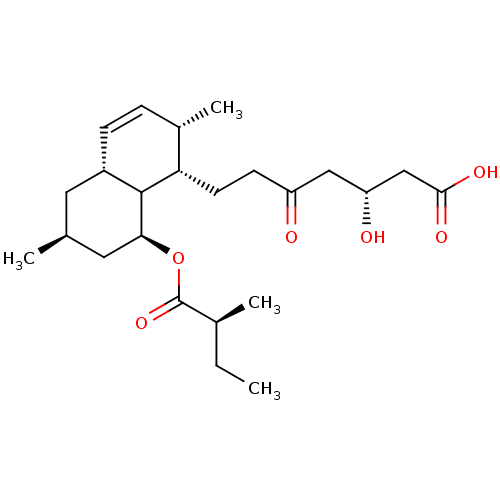
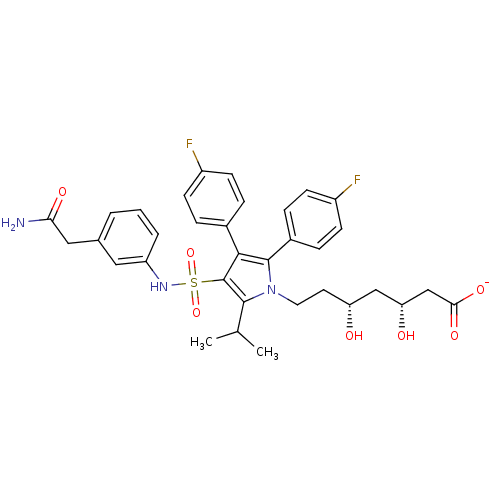
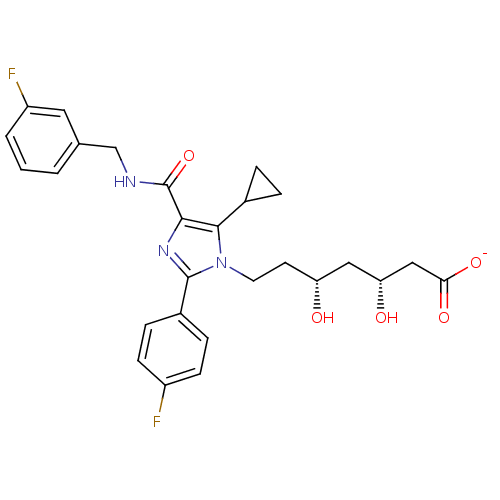
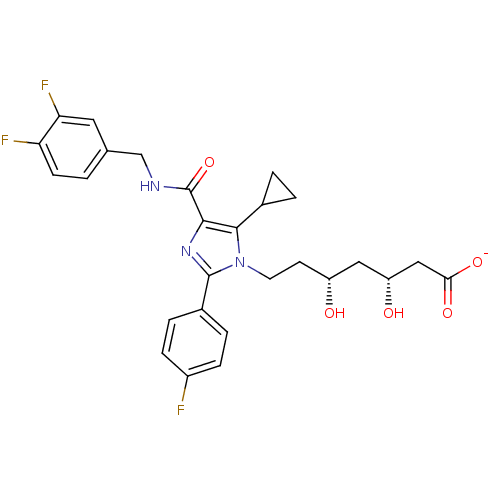
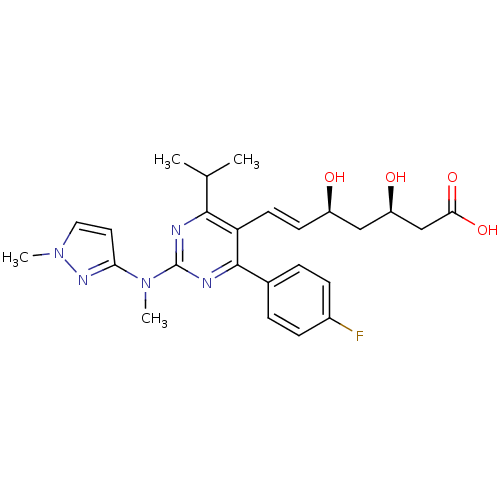
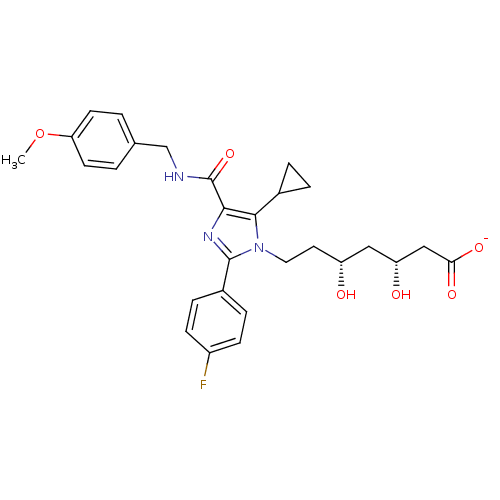
Affinity DataIC50: 0.106nMAssay Description:In vitro inhibition of HMG-CoA reductase of rat liverMore data for this Ligand-Target Pair
Affinity DataIC50: 5.90nM EC50: 0.200nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90nM EC50: 0.200nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.5nM EC50: 0.300nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2nM EC50: 0.300nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 12nM EC50: 0.300nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of rat liver microsomal HMG-CoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10nM EC50: 0.300nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nM EC50: 0.300nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.5nM EC50: 0.400nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nM EC50: 0.400nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nM EC50: 1.40nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nM EC50: 0.5nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60nM EC50: 0.5nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of rat liver microsomal HMG-CoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9nM EC50: 0.600nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10nM EC50: 0.600nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nM EC50: 0.600nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nM EC50: 0.700nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 7nM EC50: 0.700nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.850nMpH: 7.2 T: 2°CAssay Description:Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme source...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90nM EC50: 0.900nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nM EC50: 1.5nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nM EC50: 13.7nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Inhibition of rat liver HMG-CoA reductase pre incubated for 5 mins followed by substrate addition using NADPH as substrate by scintillation counter a...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Inhibition of solubilized, purified rat liver HMG-CoA reductase.More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory activity against washed rat liver microsomal HMG-CoA reductase (HMGR)More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of HMG-CoA reductase in Sprague-Dawley rat liver microsomes using [14C]HMG-CoA as substrate preincubated for 0.5 hrs before substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20nM EC50: 1nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Ability of compound to inhibit the activity of 3-hydroxy-3-methylglutarylcoenzyme A(HMGR) reductase in rat liver microsomesMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory activity against washed rat liver microsomal HMG-CoA reductase (HMGR)More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:In vitro inhibition of rat liver HMG-CoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory activity against washed rat liver microsomal HMG-CoA reductase (HMGR)More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:In vitro inhibition of rat liver HMG-CoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of rat microsomal HMGCoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:In vitro inhibition of solubilized HMG-CoA reductase in rat liver.More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:In vitro inhibition of rat liver HMG-CoA reductaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2Assay Description:All assays were carried out in a reaction buffer containing 100 nM KxPO4 at pH 7.2, 1 mM EDTA, 500 mM KCl and 1 mg/ml BSA. The concentrations of NADP...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10nM EC50: 1.10nMpH: 7.2 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of HMG-CoA reductase in Sprague-Dawley rat liver microsomes using [14C]HMG-CoA as substrate preincubated for 0.5 hrs before substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of HMG-CoA reductase in Sprague-Dawley rat liver microsomes using [14C]HMG-CoA as substrate preincubated for 0.5 hrs before substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.5nM EC50: 1.20nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of HMG-CoA reductase in Sprague-Dawley rat liver microsomes using [14C]HMG-CoA as substrate preincubated for 0.5 hrs before substrate addi...More data for this Ligand-Target Pair