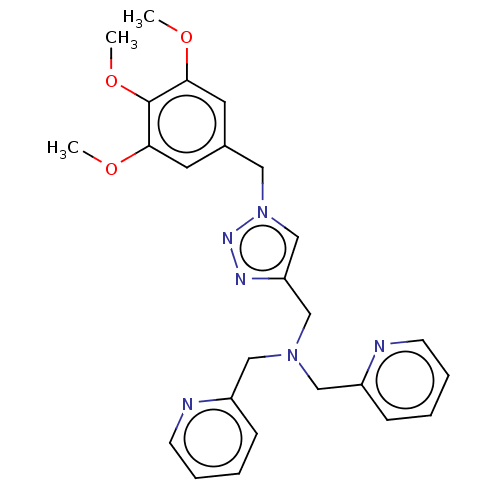
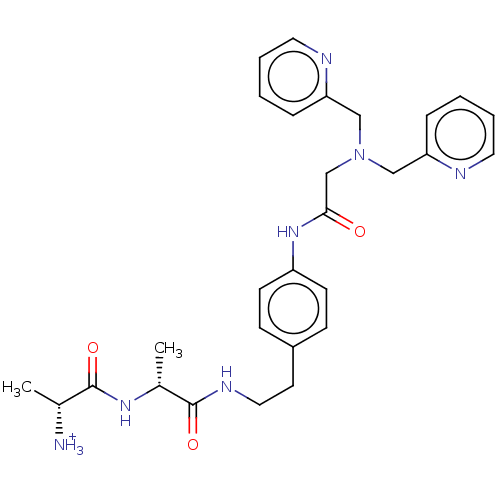
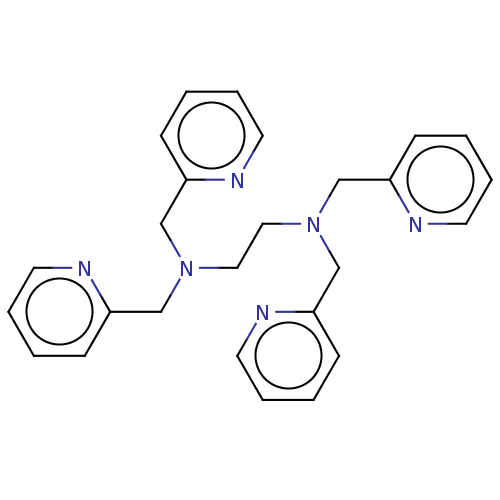
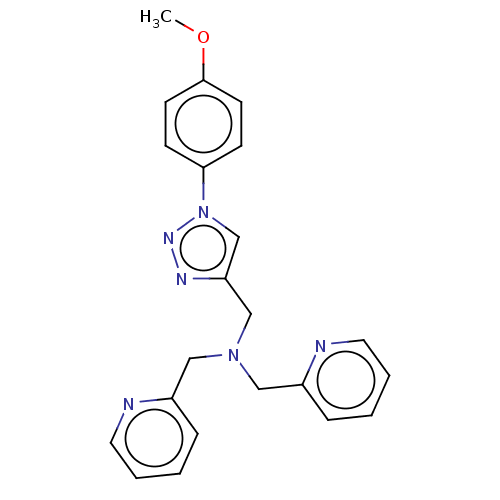
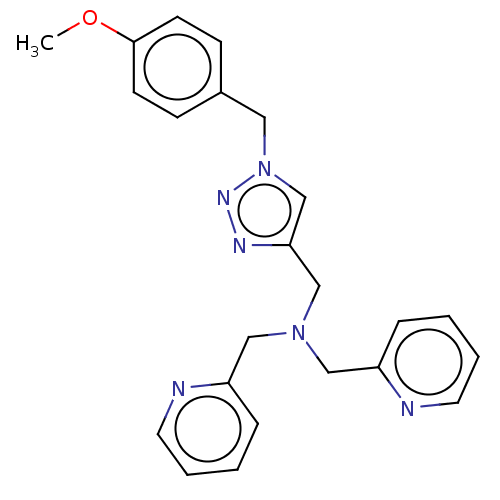
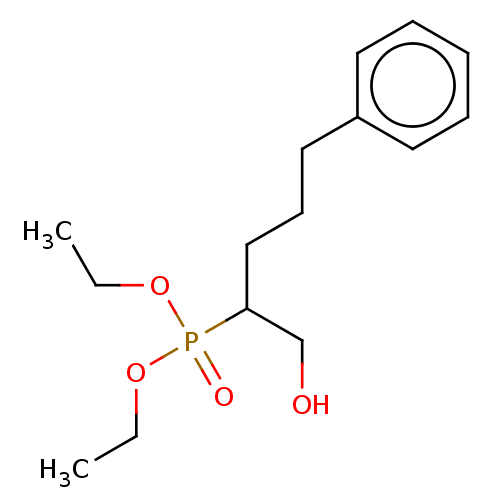
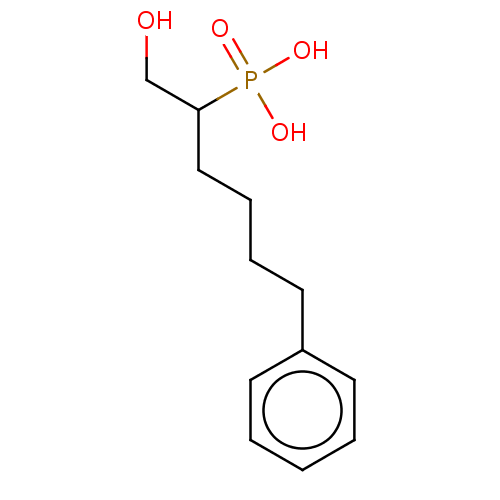
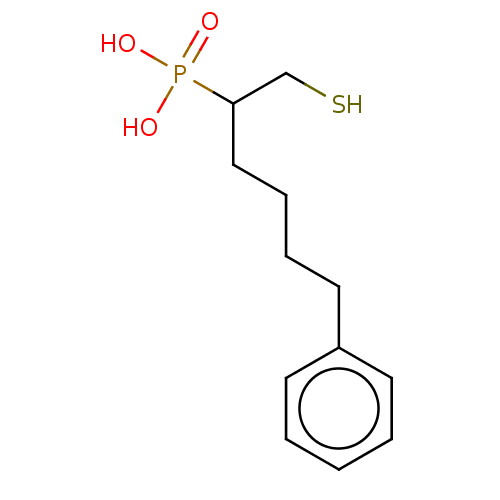
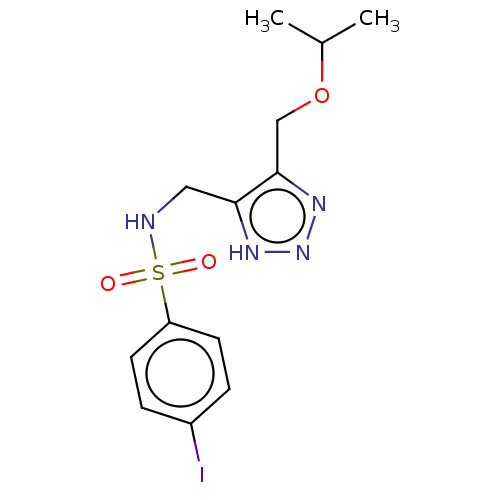
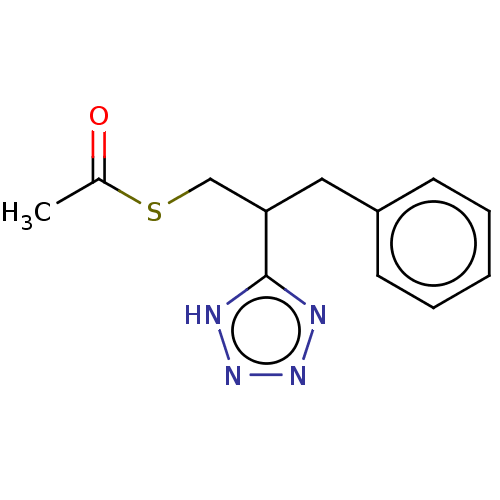
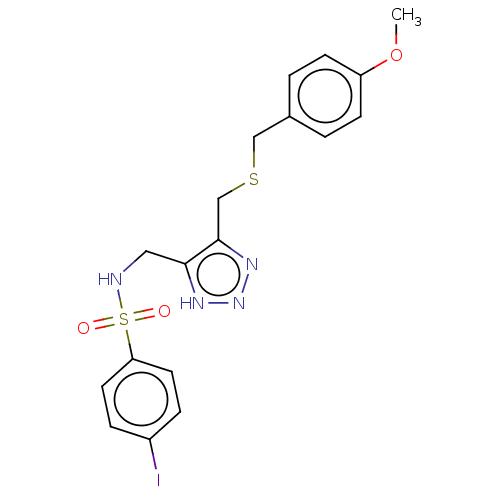
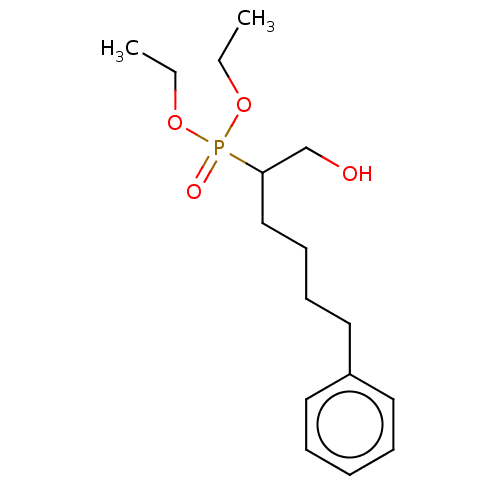
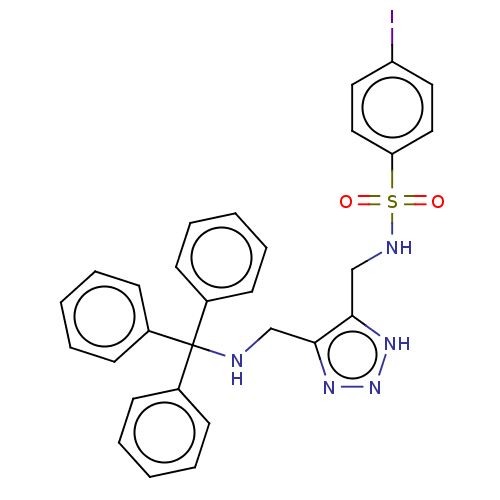
Affinity DataIC50: 140nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 610nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 710nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 860nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+3nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 1.43E+3nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 2.26E+3nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 3.15E+3nMAssay Description:a. Transfer 50 μl LB with 0.8 mM IPTG (0.4 mM final concentration in assay) to each wellb. Add 1 μl inhibitor/positive control (500/1000 &#...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90E+4nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+4nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+5nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.69E+5nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 1.93E+5nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+5nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.27E+5nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+5nMAssay Description:Inhibition of Pseudomonas aeruginosa 6His-tagged GIM-1 (Q19 to D250 residues) expressed in Escherichia coli BL21 Star(DE3) pLysS using nitrocefin as ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.53E+5nMAssay Description:Inhibition of beta lactamase GIM-1 in Pseudomonas aeruginosa in presence of nitrocefin as reporter substrate measured after 5 mins by microplate spec...More data for this Ligand-Target Pair