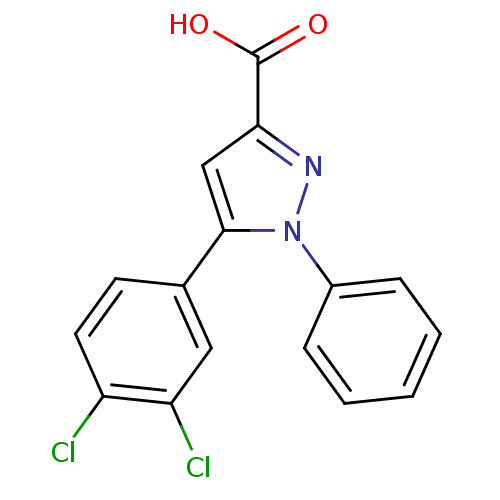
BDBM64868 5-(3,4-dichlorophenyl)-1-phenyl-1H-pyrazole-3-carboxylic acid::5-(3,4-dichlorophenyl)-1-phenyl-3-pyrazolecarboxylic acid::5-(3,4-dichlorophenyl)-1-phenyl-pyrazole-3-carboxylic acid::5-(3,4-dichlorophenyl)-1-phenylpyrazole-3-carboxylic acid::MLS001166361::SMR000549909::cid_1477263
SMILES OC(=O)c1cc(-c2ccc(Cl)c(Cl)c2)n(n1)-c1ccccc1
InChI Key InChIKey=CGAOXSZEALULKE-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 64868
Found 2 hits for monomerid = 64868
TargetMu-type opioid receptor(Human)
The Scripps Research Institute Molecular Screening Center
Curated by PubChem BioAssay
The Scripps Research Institute Molecular Screening Center
Curated by PubChem BioAssay
Affinity DataIC50: 5.82E+3nMAssay Description:Source (MLPCN Center Name): The Scripps Research Institute Molecular Screening Center (SRIMSC) Center Affiliation: The Scripps Research Institute (TS...More data for this Ligand-Target Pair
TargetTrans-activator protein BZLF1(HHV-4)
The Scripps Research Institute Molecular Screening Center
Curated by PubChem BioAssay
The Scripps Research Institute Molecular Screening Center
Curated by PubChem BioAssay
Affinity DataIC50: 6.70E+4nMAssay Description:Source (MLPCN Center Name): The Scripps Research Institute Molecular Screening Center (SRIMSC) Center Affiliation: The Scripps Research Institute (TS...More data for this Ligand-Target Pair