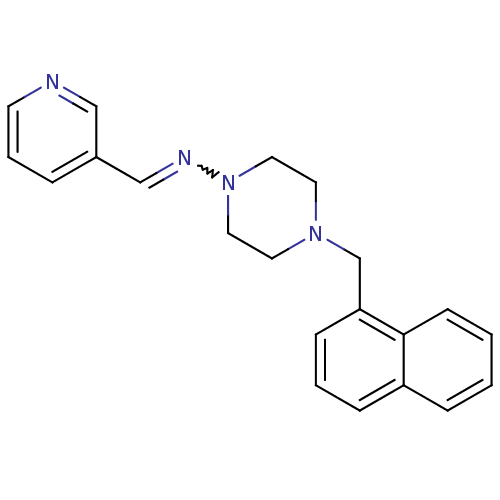
BDBM72133 4-(1-naphthylmethyl)-N-(3-pyridinylmethylene)-1-piperazinamine::MLS000570883::N-[4-(1-naphthalenylmethyl)-1-piperazinyl]-1-(3-pyridinyl)methanimine::N-[4-(naphthalen-1-ylmethyl)piperazin-1-yl]-1-pyridin-3-yl-methanimine::N-[4-(naphthalen-1-ylmethyl)piperazin-1-yl]-1-pyridin-3-ylmethanimine::SMR000193305::[4-(1-naphthylmethyl)piperazino]-(3-pyridylmethylene)amine::cid_873924
SMILES C(N1CCN(CC1)N=Cc1cccnc1)c1cccc2ccccc12
InChI Key InChIKey=NYLAUQJTBURUOI-UHFFFAOYSA-N
Data 1 EC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 72133
Found 2 hits for monomerid = 72133
TargetXaa-Pro dipeptidase(Human)
Memorial Sloan-Kettering Cancer Center; Memorial Hospital for Cancer and Allied Diseases; Sloan-Kettering Institute for Cancer Research
US Patent
Memorial Sloan-Kettering Cancer Center; Memorial Hospital for Cancer and Allied Diseases; Sloan-Kettering Institute for Cancer Research
US Patent
Affinity DataIC50: 530nMAssay Description:Source (MLPCN Center Name): The Scripps Research Institute Molecular Screening Center (SRISMC) Center Affiliation: The Scripps Research Institute (TS...More data for this Ligand-Target Pair
TargetXaa-Pro aminopeptidase 1(Human)
Memorial Sloan-Kettering Cancer Center; Memorial Hospital for Cancer and Allied Diseases; Sloan-Kettering Institute for Cancer Research
US Patent
Memorial Sloan-Kettering Cancer Center; Memorial Hospital for Cancer and Allied Diseases; Sloan-Kettering Institute for Cancer Research
US Patent
Affinity DataIC50: 9.20E+4nMAssay Description:Source (MLPCN Center Name): The Scripps Research Institute Molecular Screening Center (SRISMC) Center Affiliation: The Scripps Research Institute (TS...More data for this Ligand-Target Pair