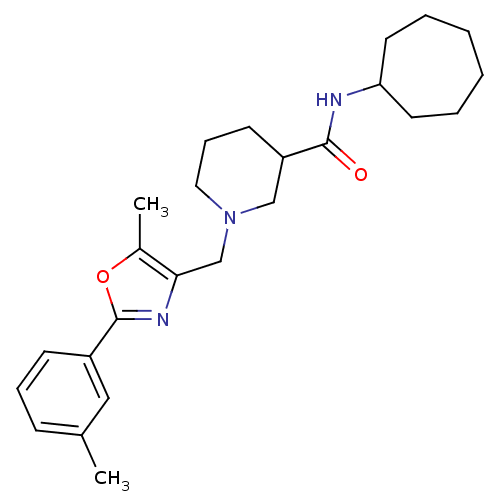
BDBM76927 MLS000089349::N-cycloheptyl-1-[[5-methyl-2-(3-methylphenyl)-1,3-oxazol-4-yl]methyl]piperidine-3-carboxamide::N-cycloheptyl-1-[[5-methyl-2-(3-methylphenyl)-4-oxazolyl]methyl]-3-piperidinecarboxamide::N-cycloheptyl-1-[[5-methyl-2-(m-tolyl)oxazol-4-yl]methyl]nipecotamide::N-cycloheptyl-1-{[5-methyl-2-(3-methylphenyl)-1,3-oxazol-4-yl]methyl}piperidine-3-carboxamide::SMR000027724::cid_3238767
SMILES Cc1oc(nc1CN1CCCC(C1)C(=O)NC1CCCCCC1)-c1cccc(C)c1
InChI Key InChIKey=OZPGNDYAYQWBDW-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 76927
Found 2 hits for monomerid = 76927
TargetVoltage-dependent T-type calcium channel subunit alpha-1H(Human)
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Vanderbilt Screening Center For Gpcrs, Ion Channels and Transporters
Curated by PubChem BioAssay
Affinity DataEC50: 5.56E+3nMAssay Description:Assay Provider: Xinmin Xie Assay Provider Affiliation: Bioscience Division, SRI International, Menlo Park, CA Grant Title: HTS Assay for Cav3 T-Type ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily KQT member 2(Rat)
Johns Hopkins Ion Channel Center
Curated by PubChem BioAssay
Johns Hopkins Ion Channel Center
Curated by PubChem BioAssay
Affinity DataIC50: 7.19E+3nMAssay Description:Source (MLPCN Center Name): Johns Hopkins Ion Channel Center (JHICC_KCNQ2_Inh_HFCP_IWS_CRC) Center Affiliation: Johns Hopkins University, School of M...More data for this Ligand-Target Pair