Report error Found 332 Enz. Inhib. hit(s) with Target = 'ATP-binding cassette sub-family C member 9'
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
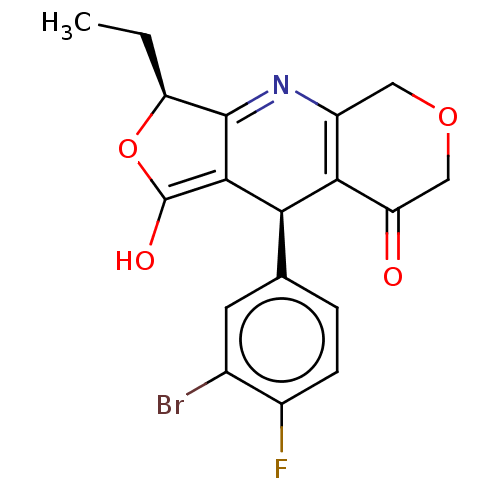
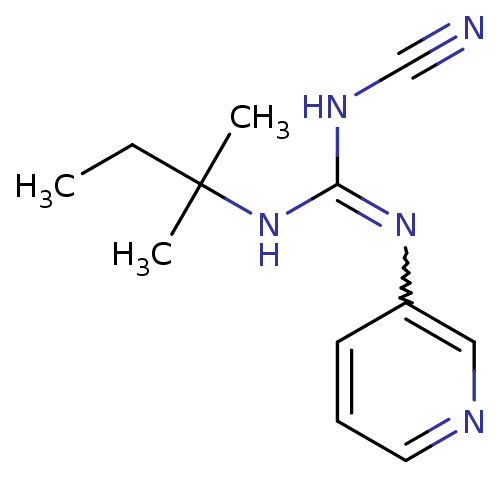
Affinity DataEC50: 0.676nMAssay Description:Activity against pig bladder KATP channel opening assessed as ability to relax spontaneous bladder contractionMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
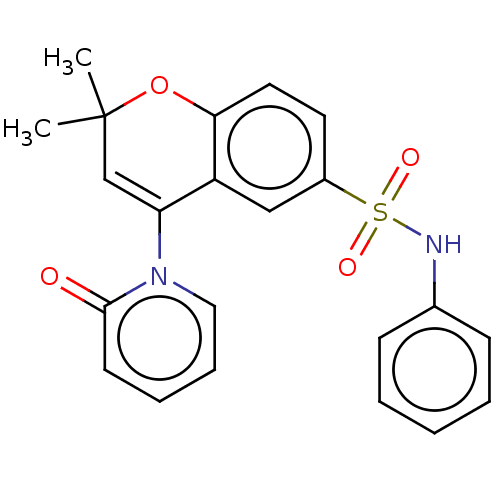
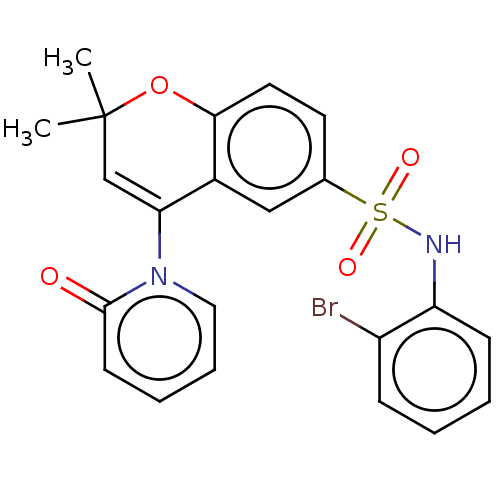
Affinity DataEC50: 0.912nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
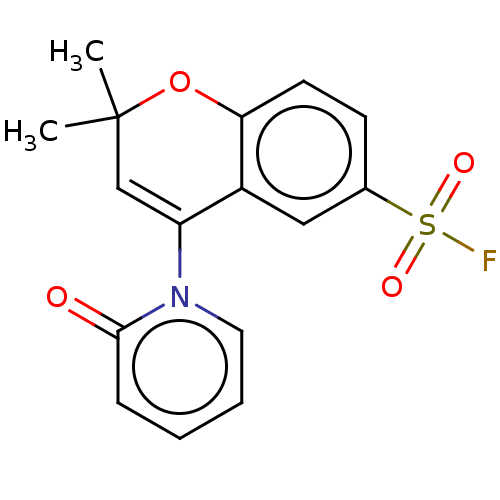
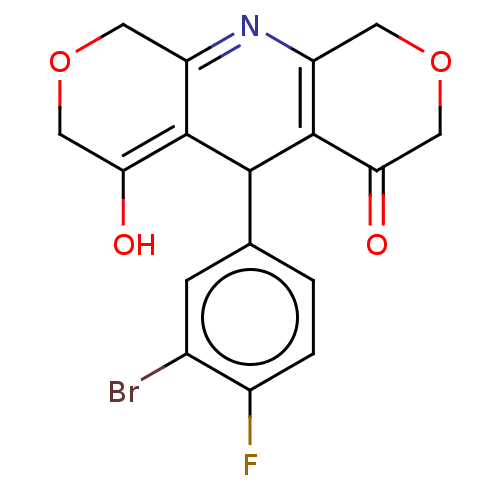
Affinity DataEC50: 1nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
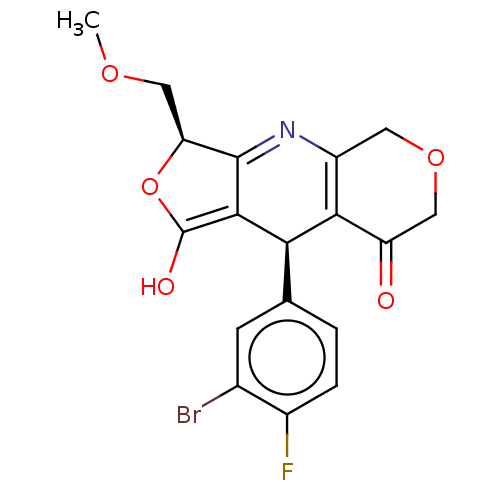
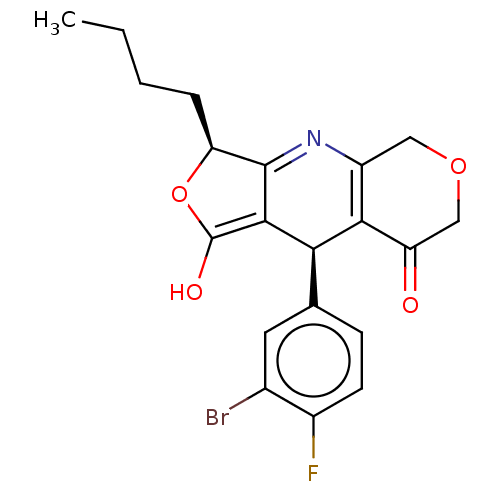
Affinity DataEC50: 1.20nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 1.80nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.80nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 2.80nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 3.70nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 3.72nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 3.89nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 6nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 7.10nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 7.40nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 8.90nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 9.80nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 9.80nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 10nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 11nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 13nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 14nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 15nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
Affinity DataKd: 15nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
Affinity DataKd: 15nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
Affinity DataKd: 17nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 17nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 18nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 19nMAssay Description:Activity against pig bladder KATP channel opening assessed as ability to relax spontaneous bladder contractionMore data for this Ligand-Target Pair
Affinity DataKd: 19nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 20nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 20nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 20nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 22nMAssay Description:Activity against pig bladder KATP channel opening assessed as ability to relax spontaneous bladder contractionMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 22.9nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 23nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKd: 23nMAssay Description:Binding affinity towards Sulfonylurea receptor 2B (SUR2B) in cultured rat aortic smooth muscle cells (SMC)More data for this Ligand-Target Pair
Affinity DataKd: 23nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 23nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 26.9nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 26.9nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataKi: 27nMAssay Description:Binding affinity was determined by displacement of [3H]P1075 from its binding sites in canine cardiac membranesMore data for this Ligand-Target Pair
Affinity DataKd: 27nMAssay Description:Binding affinity to the sulfonylurea receptor 2A (SUR2A) in purified membranes of rat-heart myocytesMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 27nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 27nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 27.5nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 28nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 28nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 28nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 29nMAssay Description:Activity against pig bladder KATP channel opening assessed as ability to relax spontaneous bladder contractionMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 30nMAssay Description:Ability to open human urinary bladder Kir6.2 channel containing SUR2B in Ltk cells by FLIPR assayMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 9/ATP-sensitive inward rectifier potassium channel 11(Human)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 35nMAssay Description:Activity against pig bladder KATP channel opening assessed as ability to relax spontaneous bladder contractionMore data for this Ligand-Target Pair