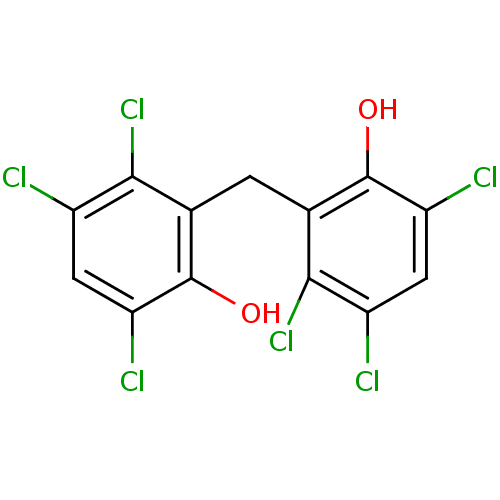
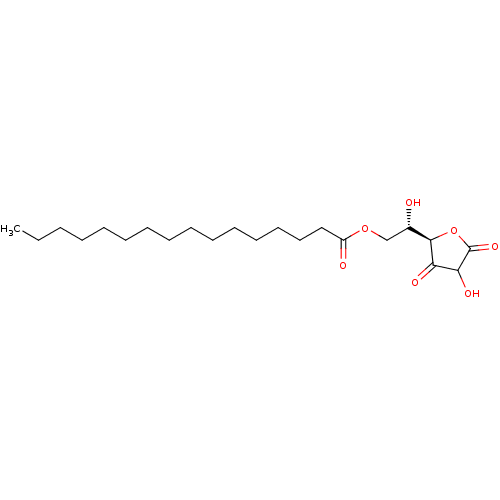
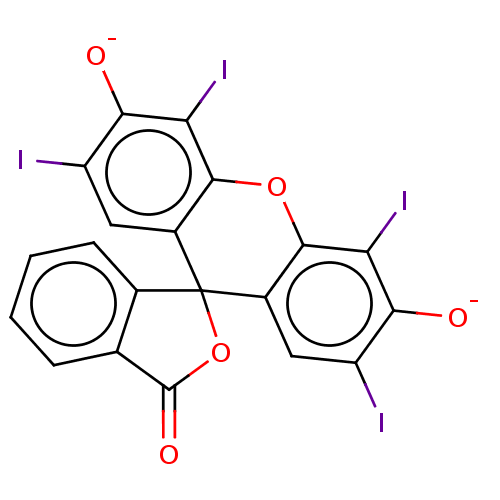
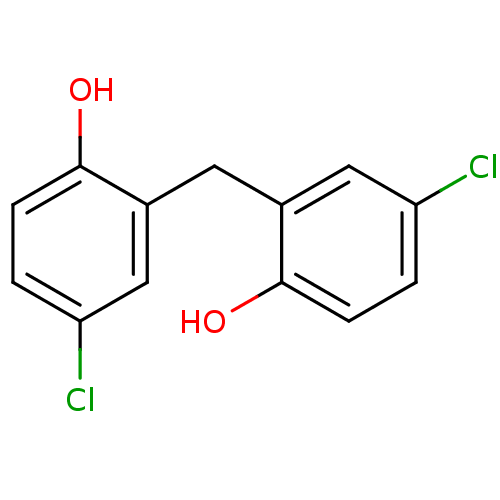
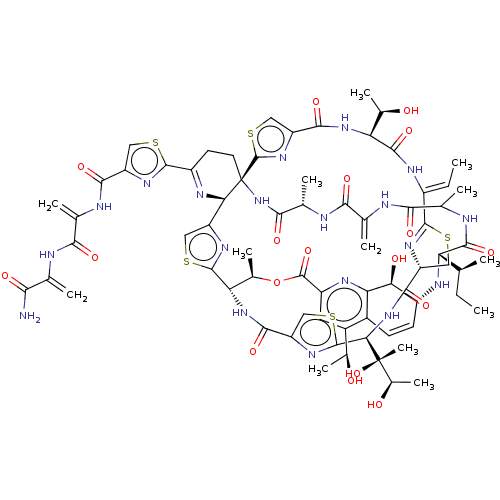
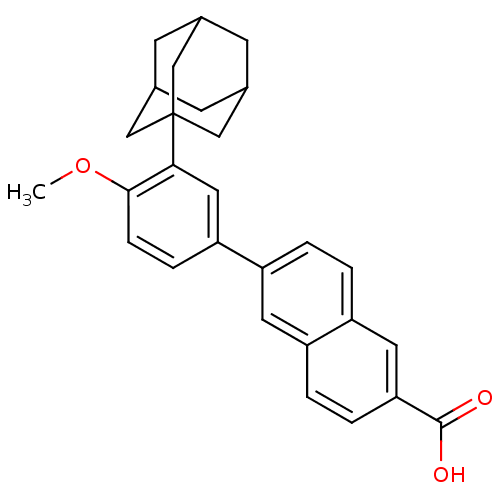
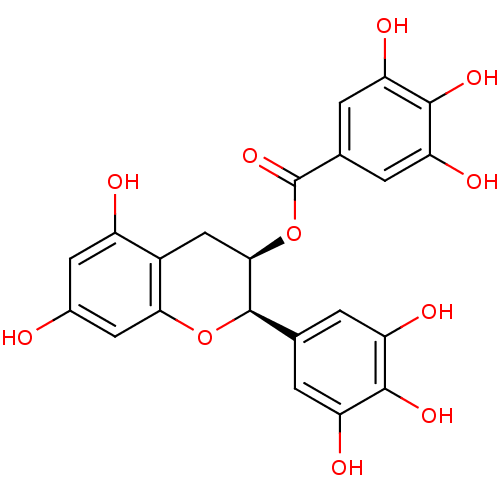
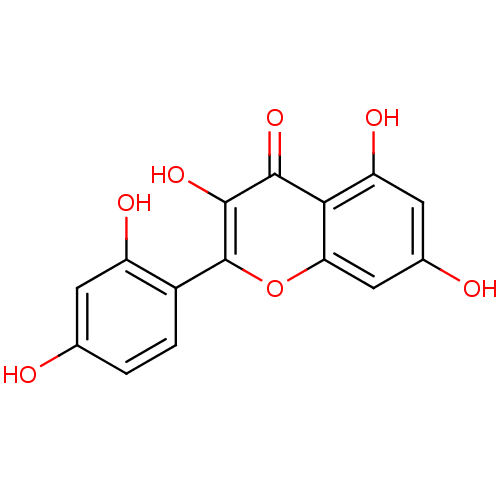
Report error Found 60 Enz. Inhib. hit(s) with Target = 'Chaperonin GroEL'
TargetChaperonin GroEL/Co-chaperonin GroES(Escherichia coli)
Indiana University School of Medicine
Curated by ChEMBL
Indiana University School of Medicine
Curated by ChEMBL
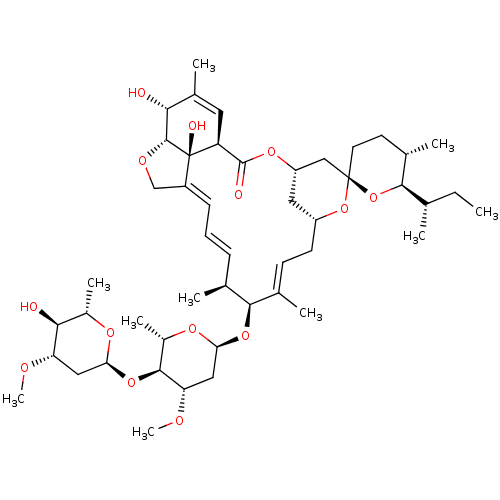
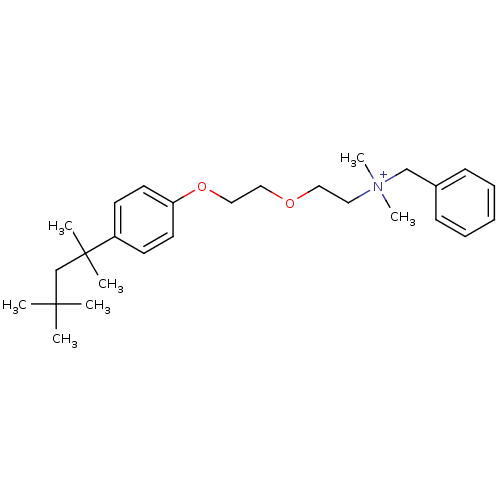
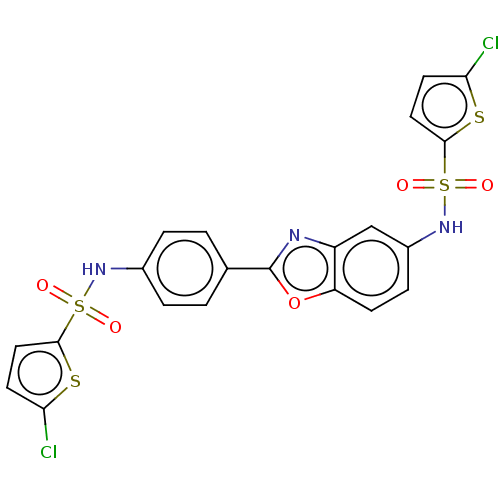
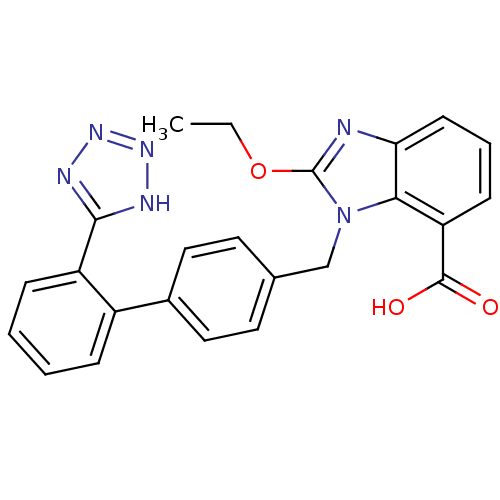
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of Escherichia coli GroEL expressed in Escherichia coliDH5alpha/Escherichia coli GroES expressed in Escherichia coli BL21 (DE3) assessed a...More data for this Ligand-Target Pair
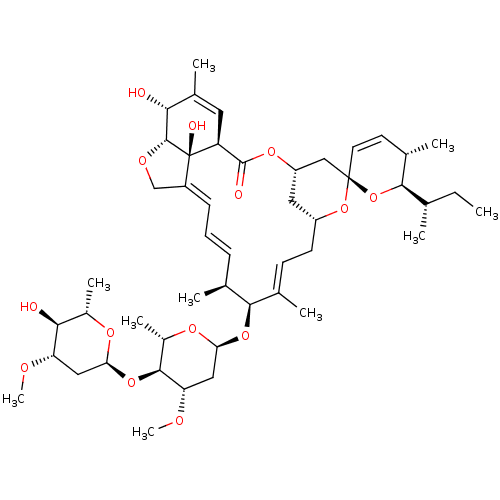
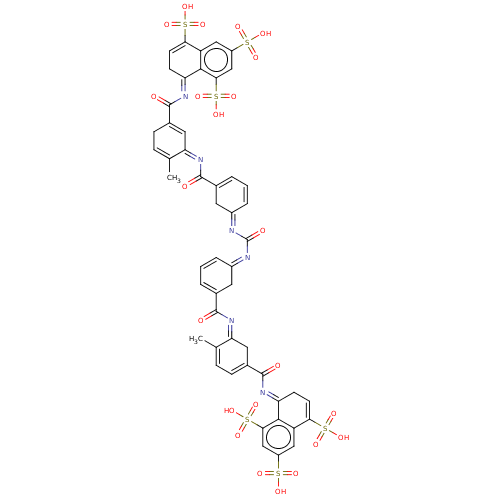
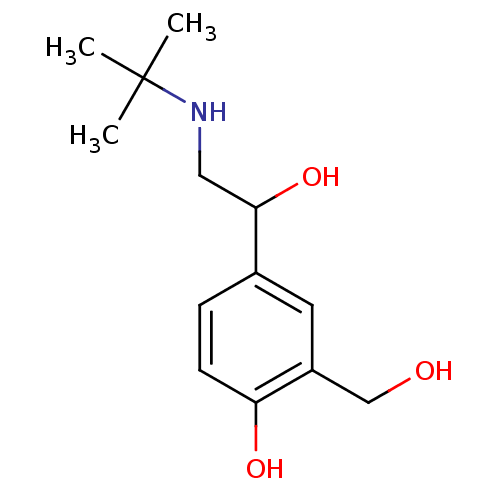
Affinity DataKd: 9.80E+3nMAssay Description:Binding affinity to Escherichia coli GroEL expressed in Escherichia coliDH5alpha by ITC assayMore data for this Ligand-Target Pair
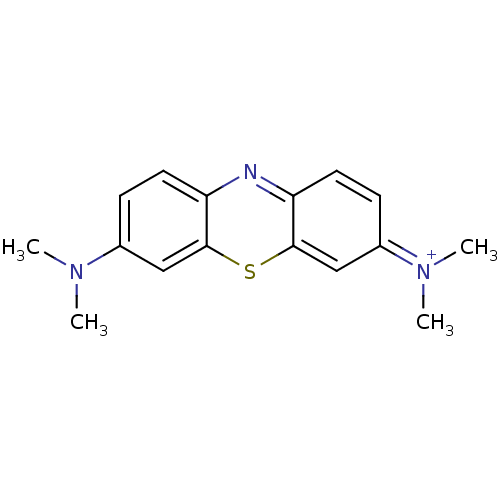
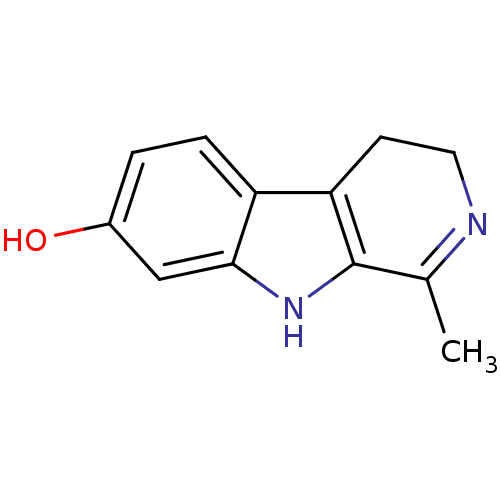
Affinity DataIC50: 1.03E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
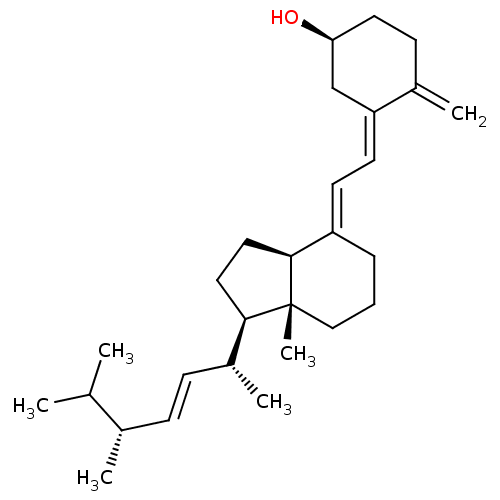
Affinity DataIC50: 1.37E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 1.52E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 1.77E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair