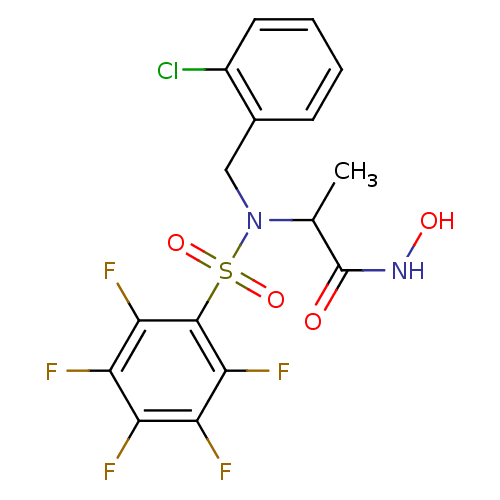
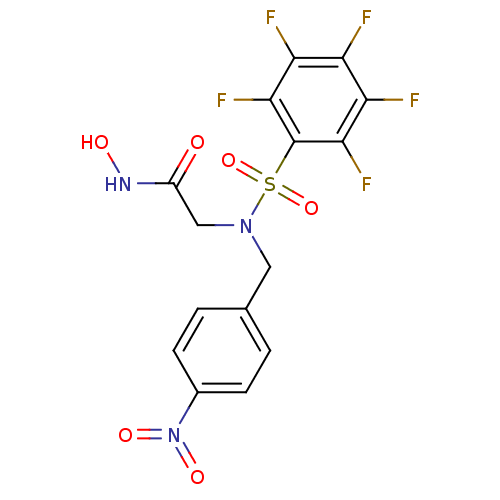
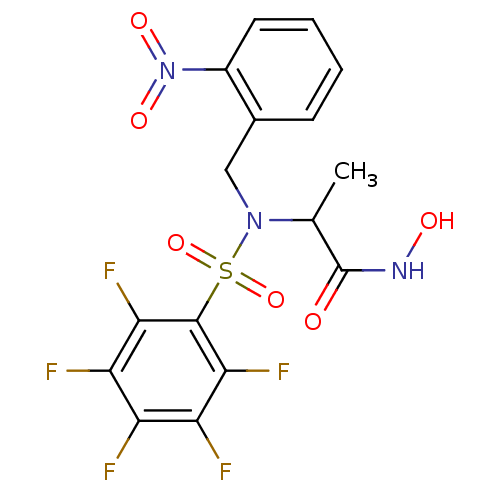
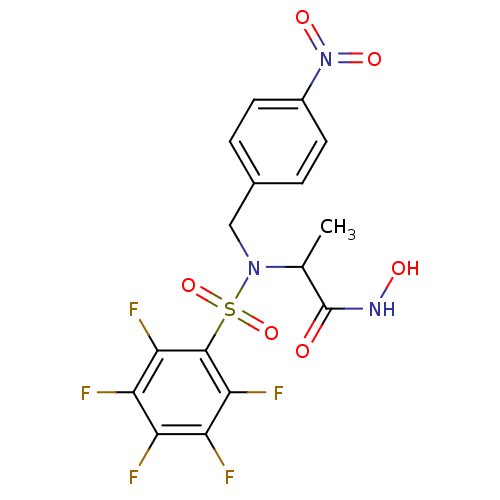
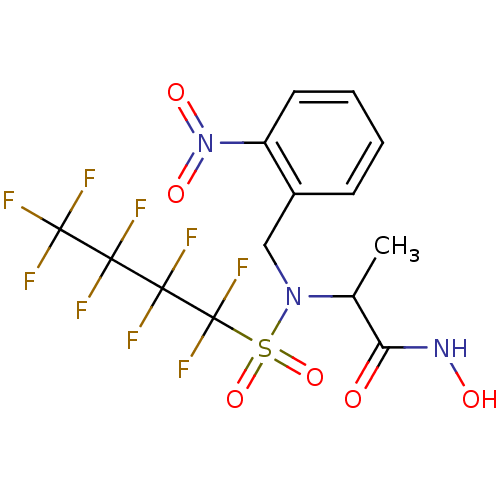
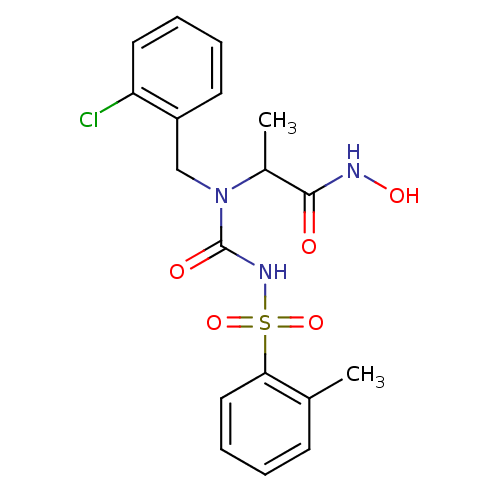
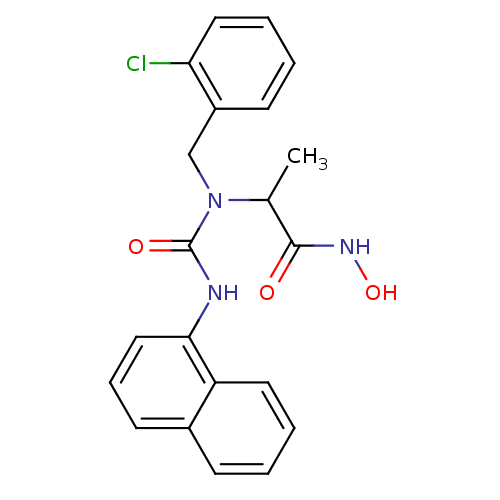
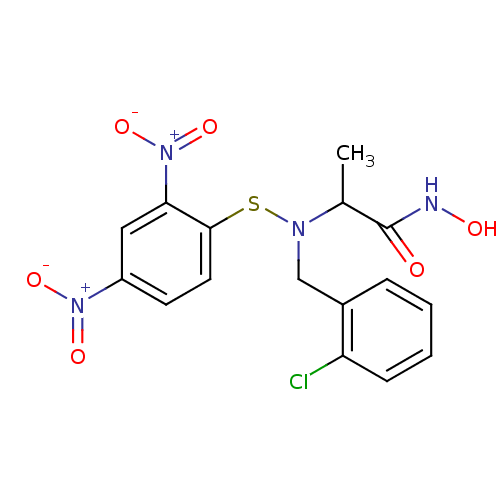
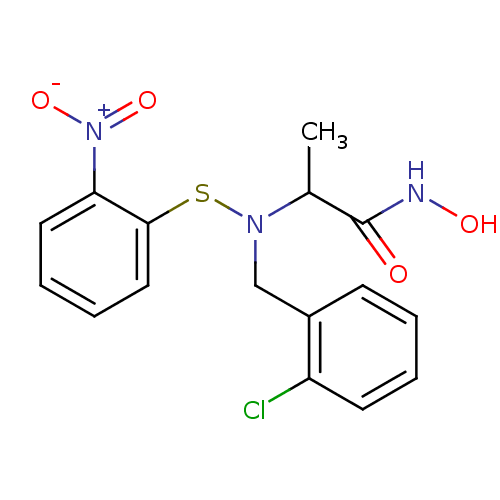
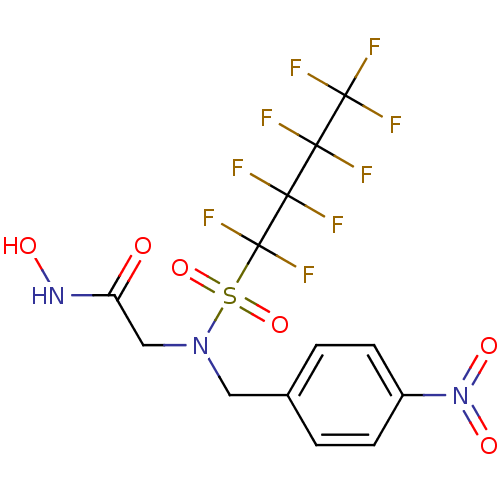
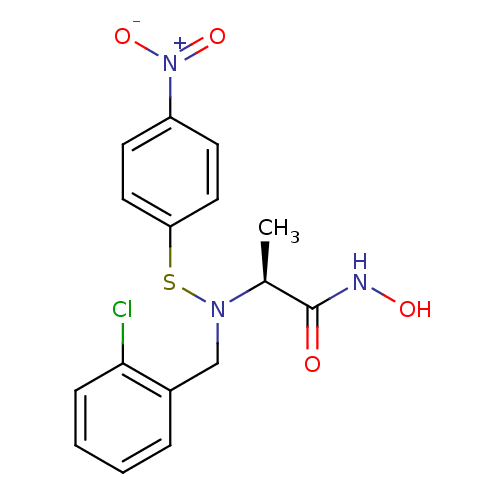
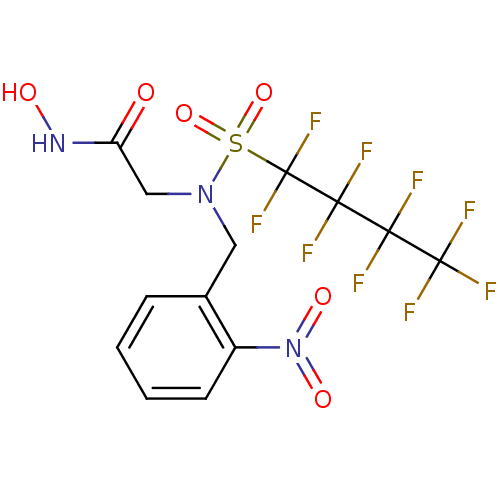
Report error Found 192 Enz. Inhib. hit(s) with Target = 'Collagenase ColG'
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Initial rates of 4-nitrophenyl acetate hydrolysis catalyzed by different CA isozymes were monitored spectrophotometrically at 400 nm. A molar absorpt...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:The rate of hydrolysis was determined from the change in absorbance at 324 nm using an extinction coefficient, 24700 M-1 cm-1 for FALGPA. Initial vel...More data for this Ligand-Target Pair