Report error Found 55 Enz. Inhib. hit(s) with Target = 'Cryptochrome-2'
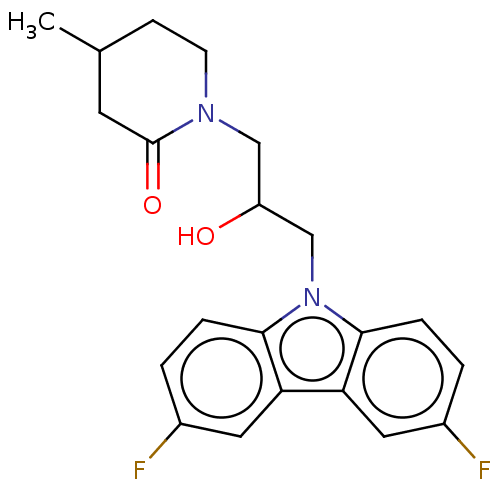
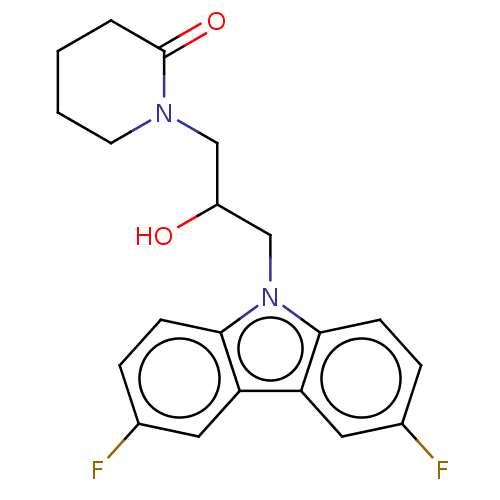
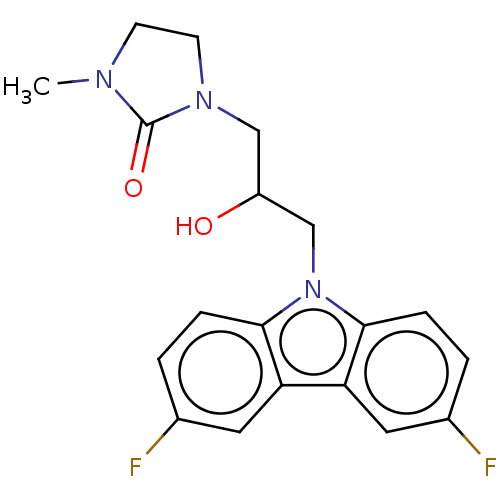
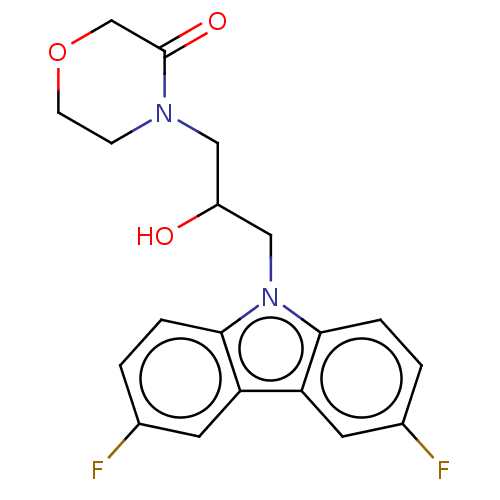
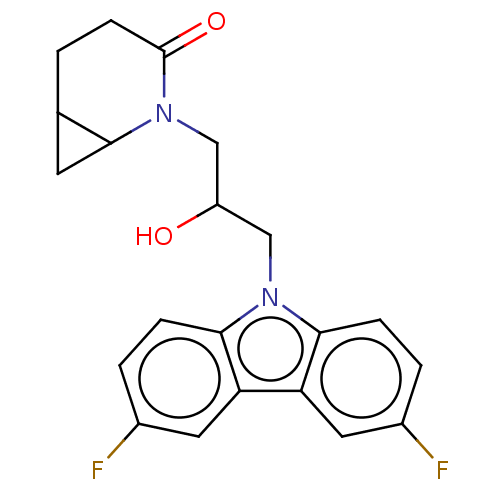
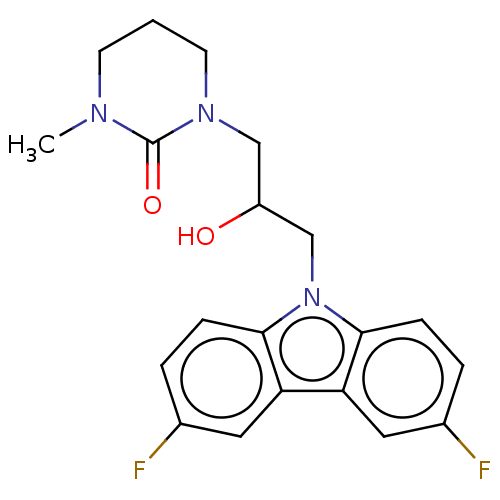
Affinity DataEC50: 79nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
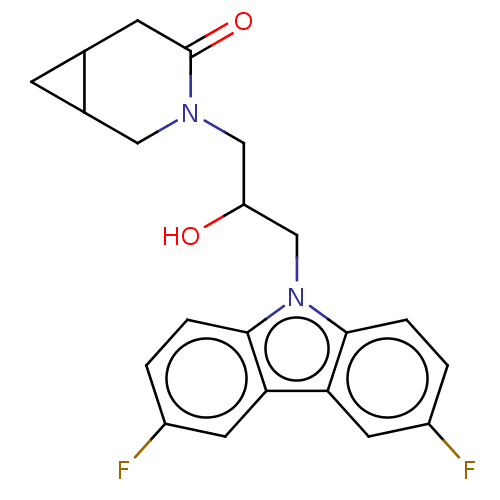
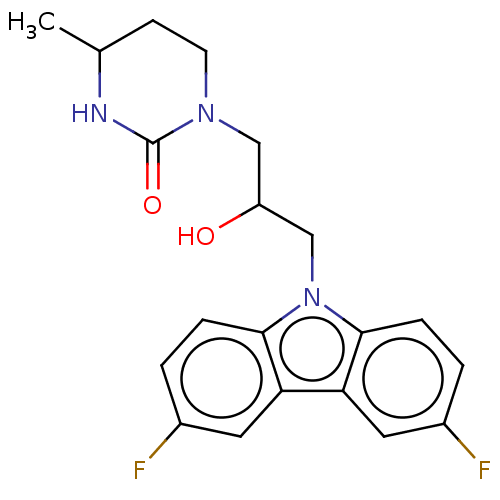
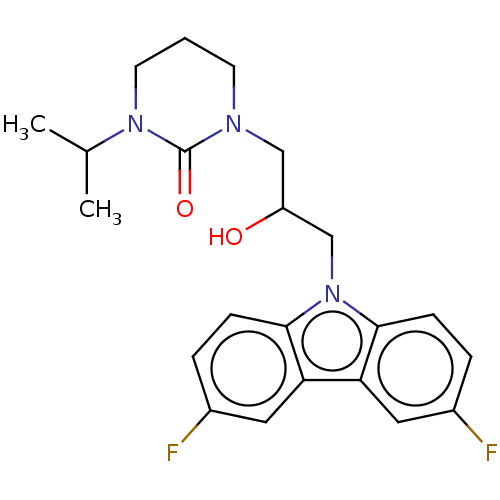
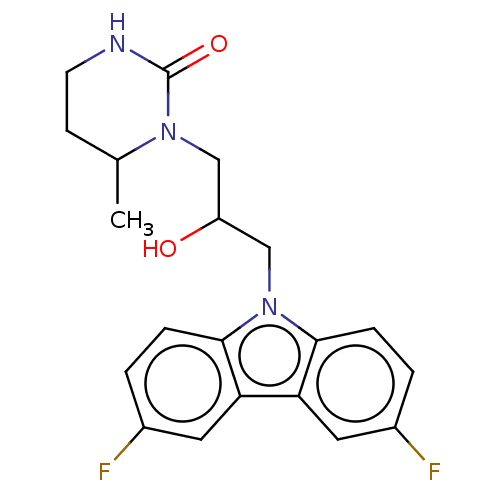
Affinity DataEC50: 79nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
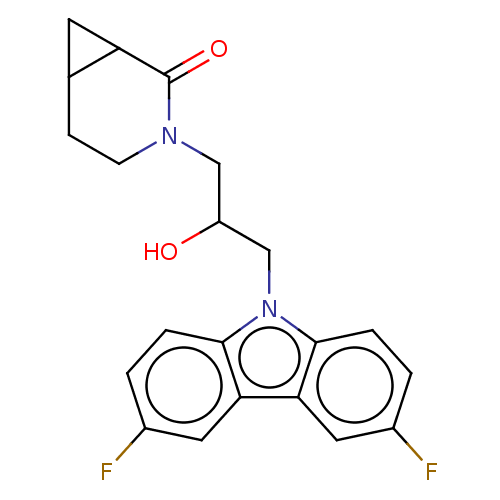
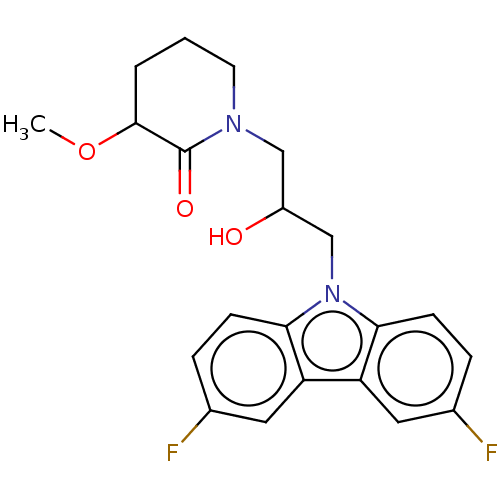
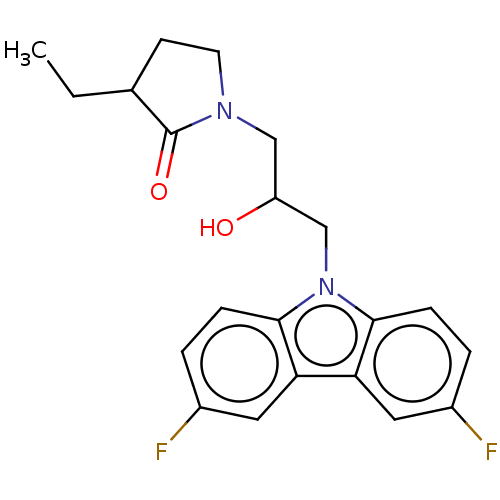
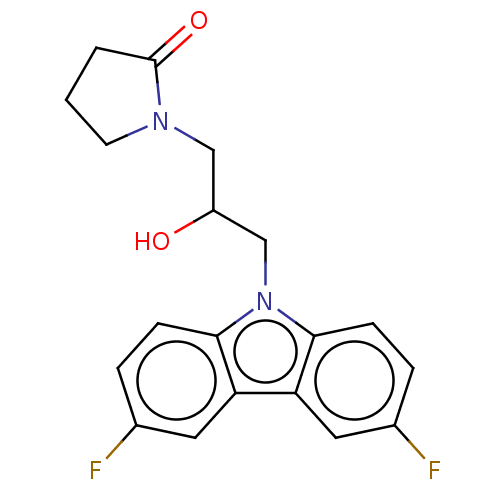
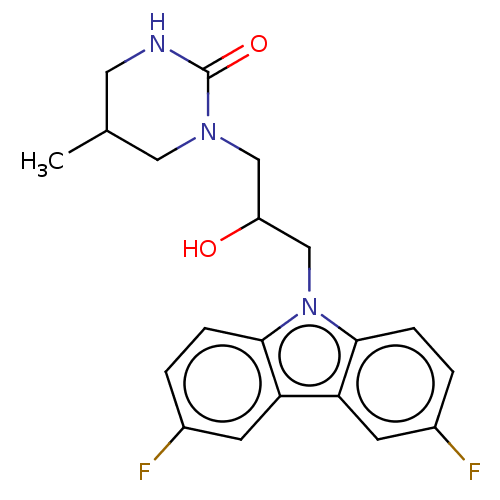
Affinity DataEC50: 128nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
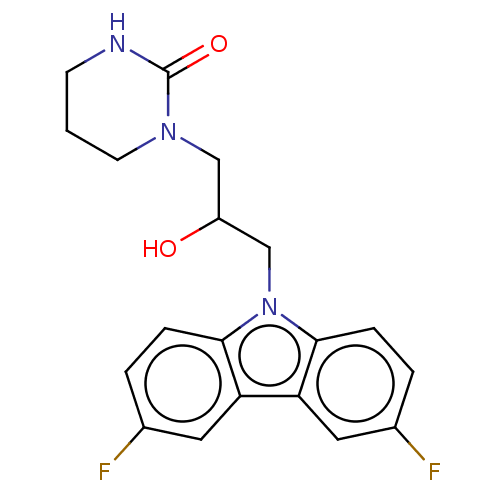
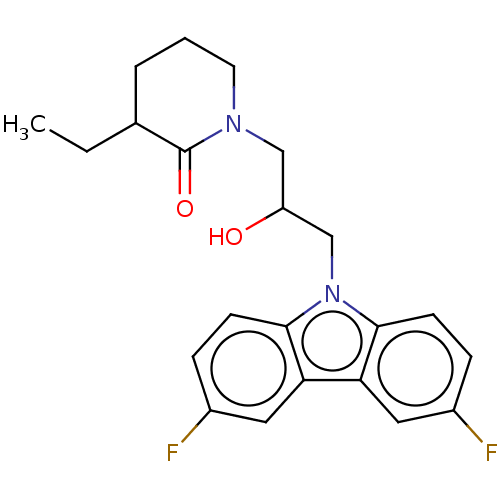
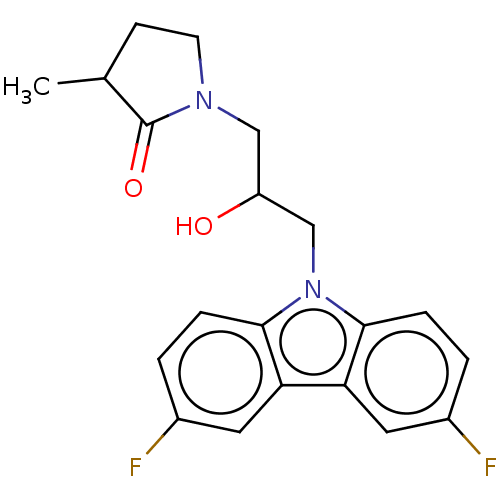
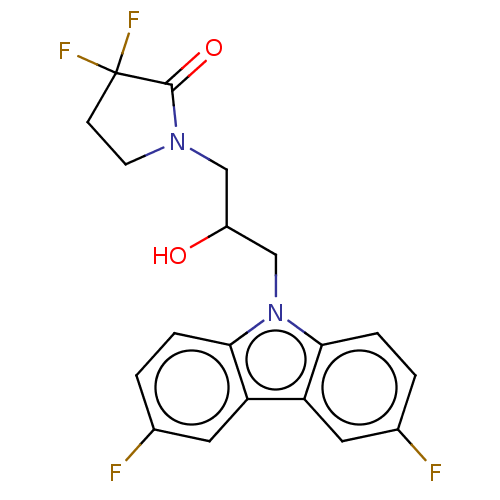
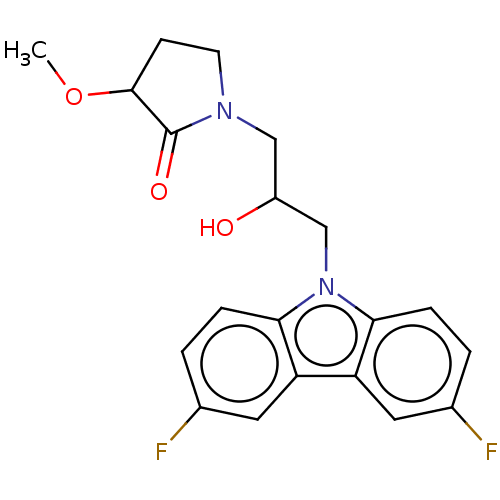
Affinity DataEC50: 149nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
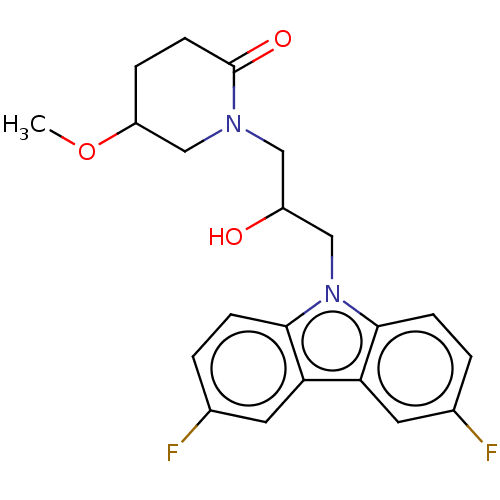
Affinity DataEC50: 163nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 164nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 167nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 171nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 188nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 192nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 194nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 199nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 208nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 224nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 246nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 261nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 330nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 332nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 363nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 397nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 406nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 417nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 417nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 424nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 447nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 463nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 487nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 497nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 528nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 541nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 547nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 565nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 605nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 620nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 625nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 667nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 716nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 719nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 742nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 833nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 855nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 870nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.03E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.06E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.12E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.14E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.15E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.19E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.36E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.49E+3nMAssay Description:Modulation activity at CRY2/PER2 in human U2OS cells harboring per2-dLuc assessed as activation of per2 activity measured every 100 mins for 5 days b...More data for this Ligand-Target Pair