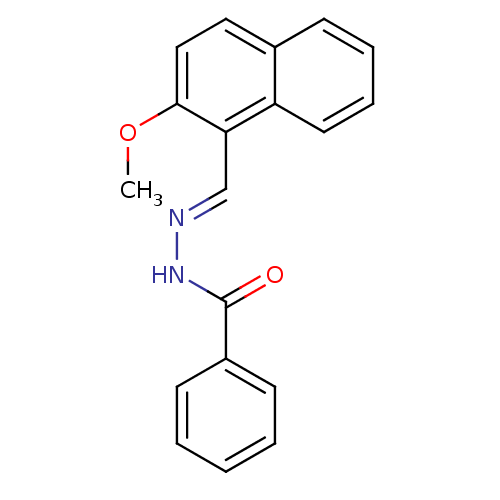
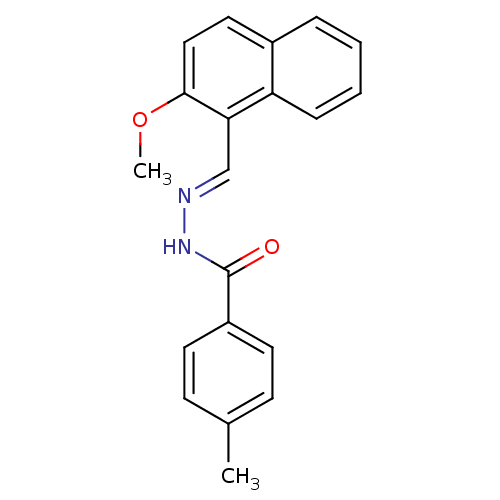
Report error Found 18 Enz. Inhib. hit(s) with Target = 'Gag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S]'
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 400nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 500nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 500nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 1.00E+3nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 2.00E+3nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 2.00E+3nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 3.00E+3nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 5.00E+3nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 7.00E+3nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 7.50E+3nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 1.20E+4nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 1.50E+4nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 1.50E+4nMpH: 8.0 T: 2°CAssay Description:To assess the effect of inhibitors, RT RNH activity was measured using a FRET-based microplate fluorescence assay. Briefly, the RNA-DNA hybrid duplex...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 2.50E+4nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 5.00E+4nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 5.00E+4nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 5.00E+4nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [600-1028,C879S]/[600-1151,C879S](Human immunodeficiency virus type 1)
Rutgers University
Rutgers University
Affinity DataIC50: 5.00E+4nMpH: 7.8 T: 2°CAssay Description:HIV-1 RT RDDP activity was generally determined by a fixed time assay. Briefly, reaction mixtures contained p51/p66 RT, template-primer, and [3H]-TTP...More data for this Ligand-Target Pair
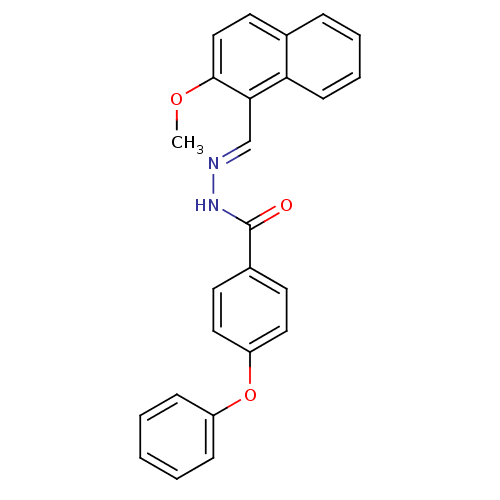

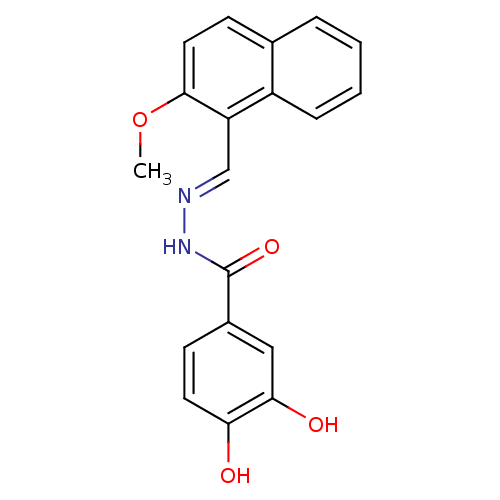
 3D Structure (crystal)
3D Structure (crystal)