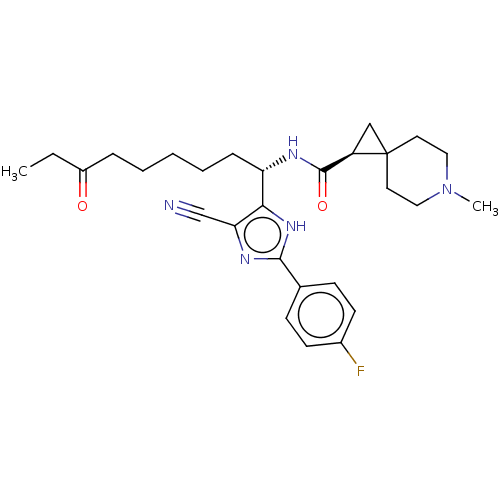
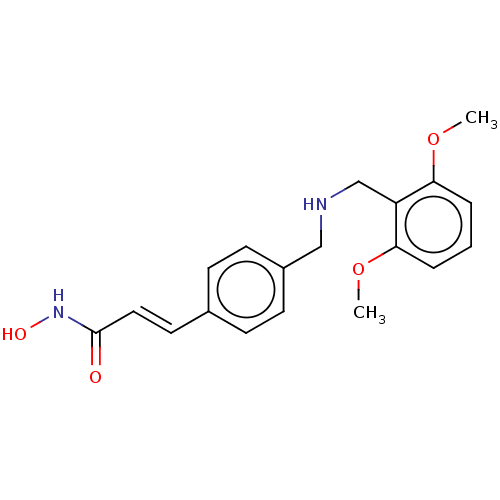
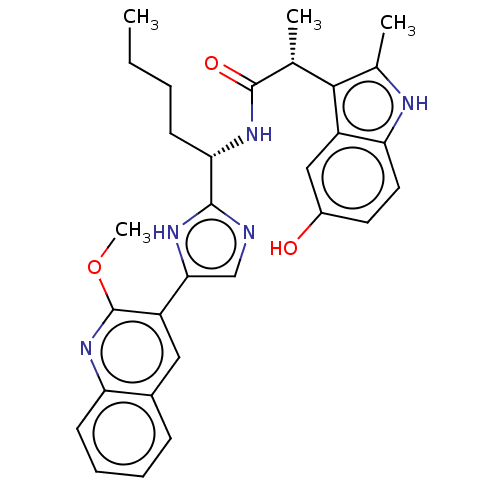
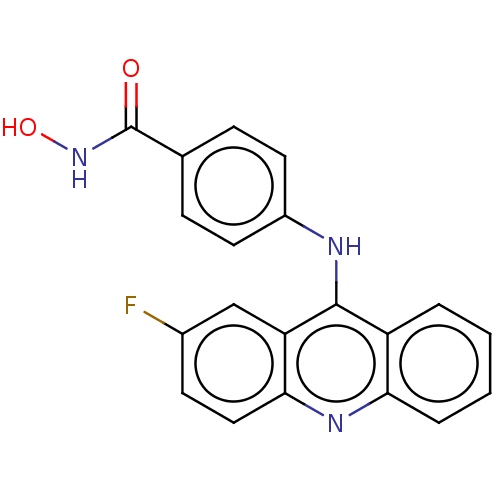
Report error Found 266 Enz. Inhib. hit(s) with Target = 'Histone deacetylase 1/2/3'
TargetHistone deacetylase 1/2/3/4/5/6/7/8/9/11/Polyamine deacetylase HDAC10(Mouse)
Institute of Technology
Curated by ChEMBL
Institute of Technology
Curated by ChEMBL
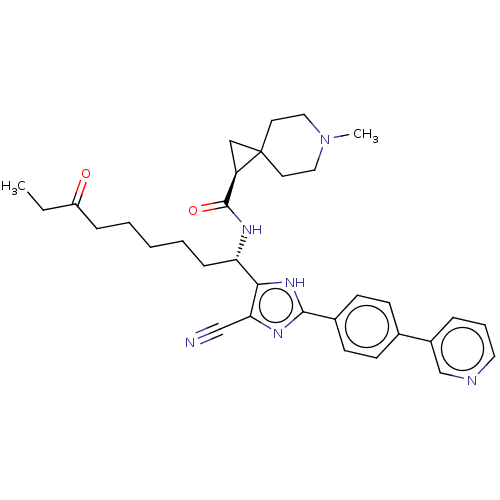
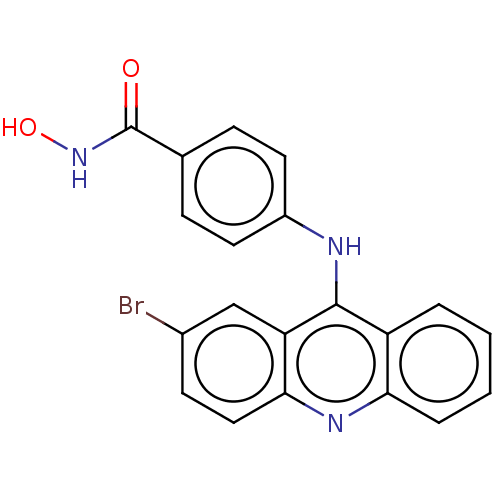
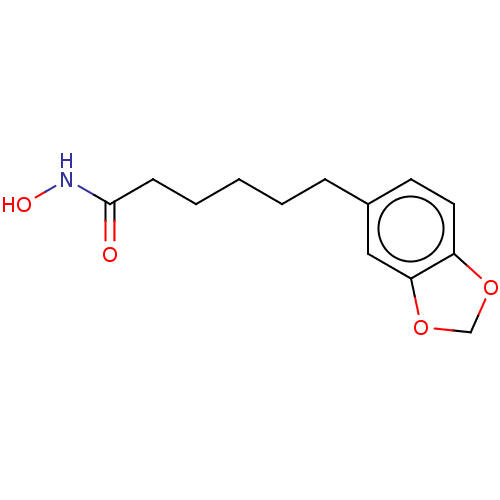
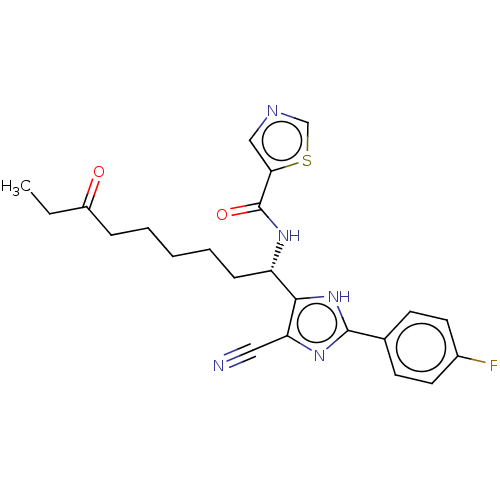
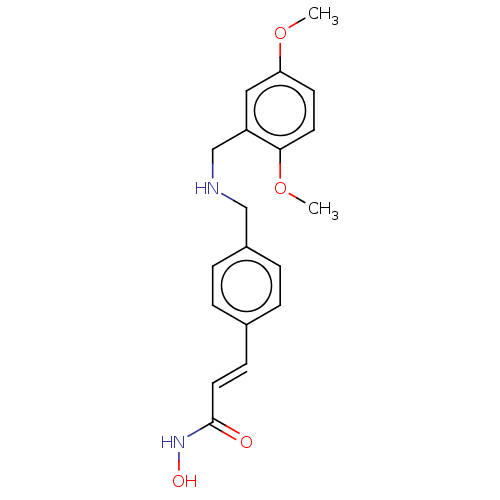
Affinity DataIC50: 4nMAssay Description:Inhibitory activity against histone deacetylases (HDAC) prepared from mouse melanoma B16/BL6 cellsMore data for this Ligand-Target Pair
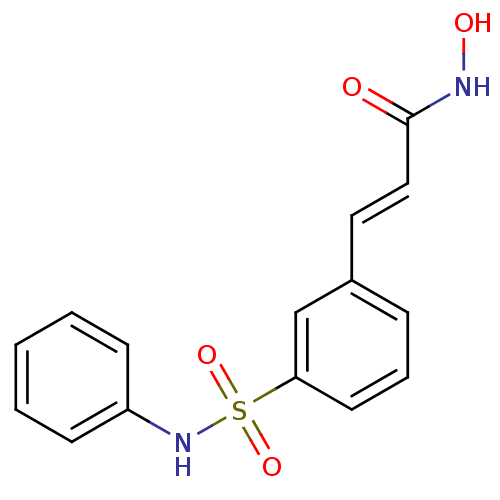
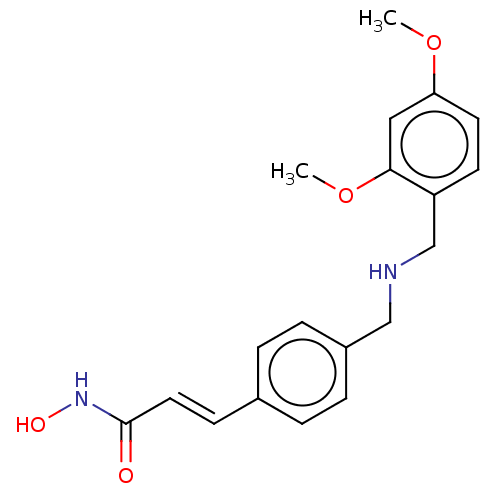
Affinity DataEC50: 4.70nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of 5% NHS by...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:Inhibition of HDAC1/HDAC2/HDAC3 in human HeLa nuclear extracts using MAL as substrate after 4 hrs by UHPLC-ESI-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of HDAC class1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 14nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of 5% NHS by...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of 5% NHS by...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of class 1 HDAC in human HeLa cell nuclear extract using Boc-Lys(Ac)-AMC as substrate preincubated for 5 mins followed by substrate additi...More data for this Ligand-Target Pair
Affinity DataEC50: 39nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of 5% NHS by...More data for this Ligand-Target Pair
Affinity DataEC50: 55nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataEC50: 77nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
TargetHistone deacetylase 1/2/3/4/5/6/7/8/9/11/Polyamine deacetylase HDAC10(Mouse)
Institute of Technology
Curated by ChEMBL
Institute of Technology
Curated by ChEMBL
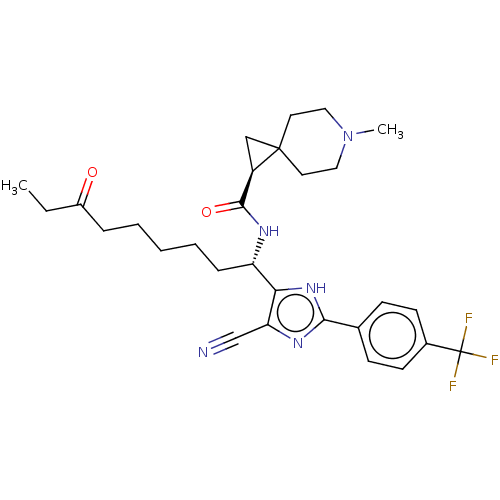
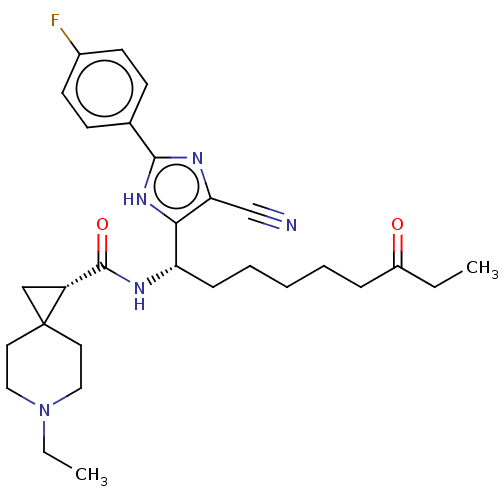
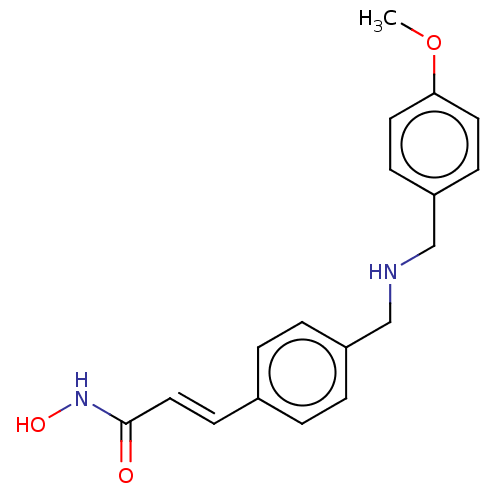
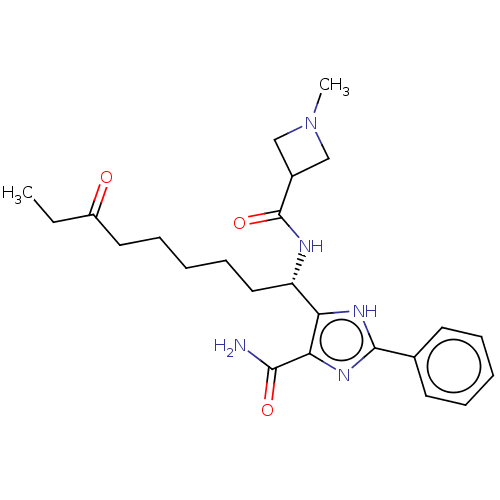
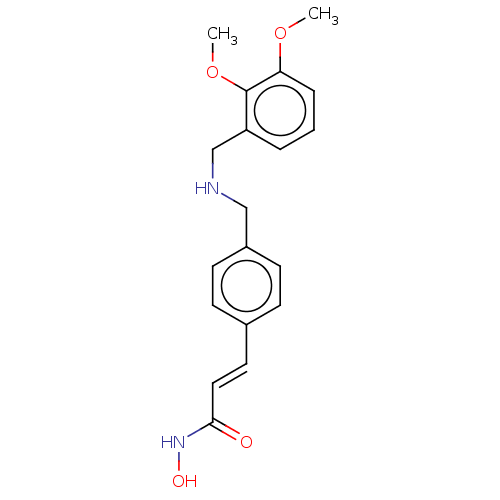
Affinity DataIC50: 84nMAssay Description:Inhibitory activity against histone deacetylases (HDAC) prepared from mouse melanoma B16/BL6 cellsMore data for this Ligand-Target Pair
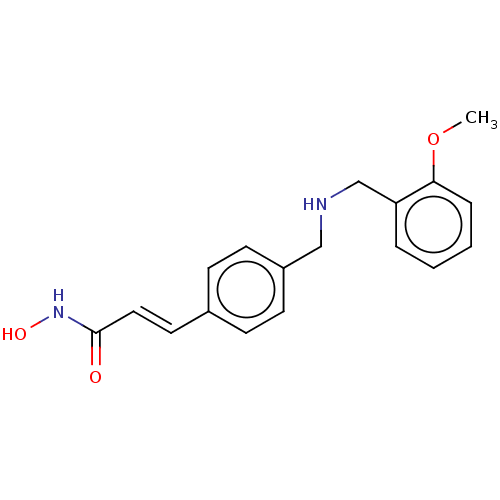
Affinity DataEC50: 97nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of class 1 HDAC in human HeLa cell nuclear extract using Boc-Lys(Ac)-AMC as substrate preincubated for 5 mins followed by substrate additi...More data for this Ligand-Target Pair
Affinity DataEC50: 105nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataEC50: 107nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of class 1 HDAC in human Jurkat 2C4 model of HIV latency assessed as reactivation of HIV latency incubated for 18 to 24 hrs in presence of...More data for this Ligand-Target Pair
TargetHistone deacetylase 1/2/3/4/5/6/7/8/9/11/Polyamine deacetylase HDAC10(Mouse)
Institute of Technology
Curated by ChEMBL
Institute of Technology
Curated by ChEMBL
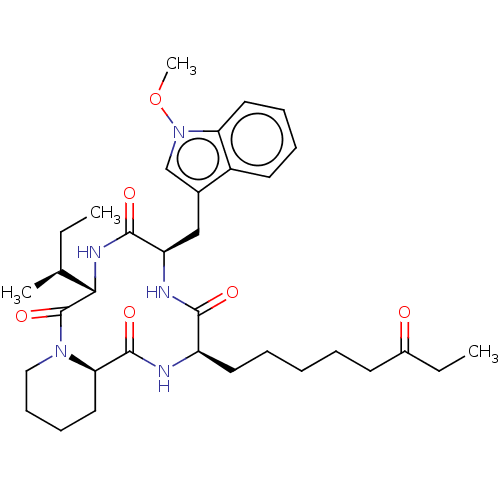
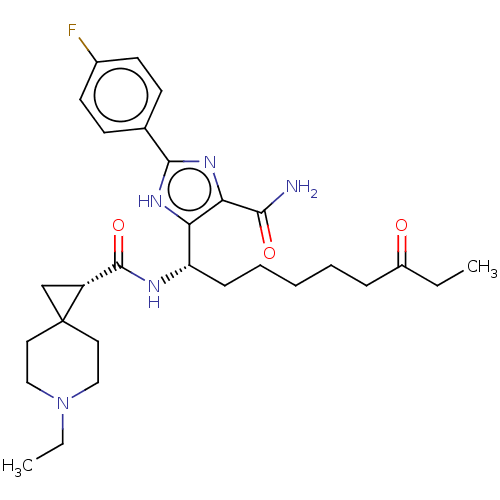
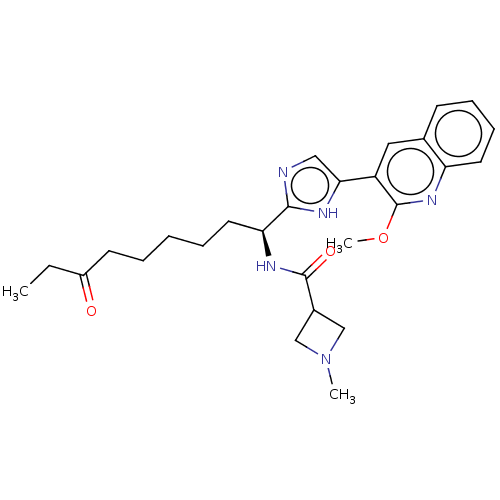
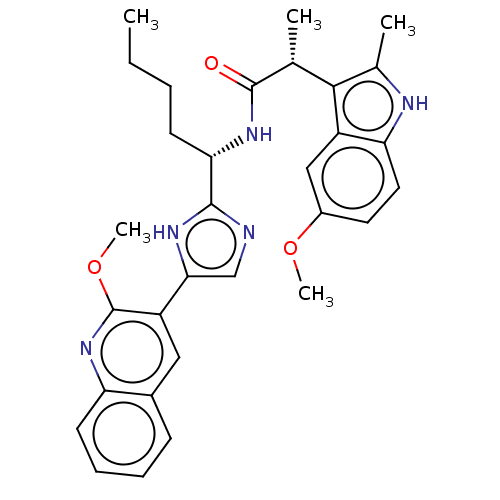
Affinity DataIC50: 141nMAssay Description:Inhibitory activity against histone deacetylases (HDAC) prepared from mouse melanoma B16/BL6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of class 1 HDAC in human HeLa cell nuclear extract using Boc-Lys(Ac)-AMC as substrate preincubated for 5 mins followed by substrate additi...More data for this Ligand-Target Pair
Affinity DataEC50: 161nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataEC50: 165nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataEC50: 166nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataEC50: 175nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataEC50: 178nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of 5% NHS by...More data for this Ligand-Target Pair
Affinity DataIC50: 188nMAssay Description:Inhibition of HDAC1/HDAC2/HDAC3 in human HeLa nuclear extracts using MAL as substrate after 4 hrs by UHPLC-ESI-MS/MS analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 189nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 193nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataIC50: 195nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of class 1 HDAC in human Jurkat 2C4 model of HIV latency assessed as reactivation of HIV latency incubated for 18 to 24 hrs in presence of...More data for this Ligand-Target Pair
Affinity DataEC50: 233nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataIC50: 235nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of class 1 HDAC in human Jurkat 2C4 model of HIV latency assessed as reactivation of HIV latency incubated for 18 to 24 hrs in presence of...More data for this Ligand-Target Pair
Affinity DataEC50: 247nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Inhibition of class 1 HDAC in human HeLa cell nuclear extract using Boc-Lys(Ac)-AMC as substrate preincubated for 5 mins followed by substrate additi...More data for this Ligand-Target Pair
Affinity DataEC50: 254nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 0.1% ...More data for this Ligand-Target Pair
Affinity DataIC50: 258nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 259nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataEC50: 269nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Inhibition of HDAC1/2/3/6 in human Cal27 cells using Boc-Lys(epsilon-Ac)-AMC as substrate preincubated for 18 hrs followed by substrate addition meas...More data for this Ligand-Target Pair
Affinity DataEC50: 276nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataIC50: 278nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 292nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 296nMAssay Description:Inhibition Class 1 histone deacetylase in human HeLa nuclear extracts using Fluor-de- Lys-green substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of class 1 HDAC in human Jurkat 2C4 model of HIV latency assessed as reactivation of HIV latency incubated for 18 to 24 hrs in presence of...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of Class 1 histone deacetylase in human THP-1 cells using Ac-LGK(Ac)-AMC as substrate incubated for 3 hrs followed by 50 uM SAHA addition ...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Inhibition of HDAC1/2/3/6 in human A2780cisR cells using Boc-Lys(epsilon-Ac)-AMC as substrate preincubated for 18 hrs followed by substrate addition ...More data for this Ligand-Target Pair
Affinity DataEC50: 322nMAssay Description:Inhibition of Class 1 HDAC in VSV-G-pseudotyped HIV infected human 2C4 cell Jurkat model assessed as reactivation of HIV latency in presence of 5% NH...More data for this Ligand-Target Pair
Affinity DataEC50: 324nMAssay Description:Inhibition of class 1 HDAC in human Jurkat model of HIV latency assessed as reactivation of HIV latency incubated for 20 hrs in presence of SAHA and ...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:Inhibition of HDAC1/2/3/6 in human A2780cisR cells using Boc-Lys(epsilon-Ac)-AMC as substrate preincubated for 18 hrs followed by substrate addition ...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:Inhibition of HDAC1/2/3/6 in human Cal27CisR cells using Boc-Lys(epsilon-Ac)-AMC as substrate preincubated for 18 hrs followed by substrate addition ...More data for this Ligand-Target Pair
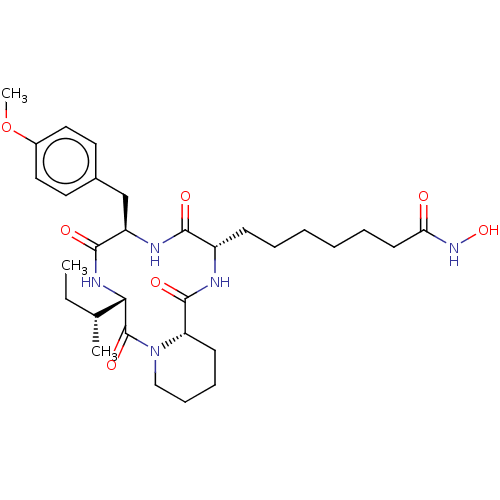
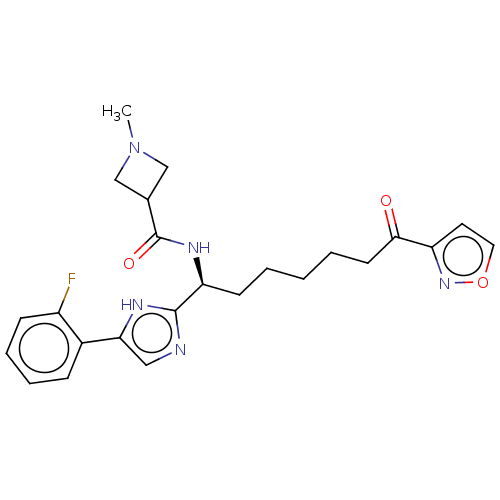
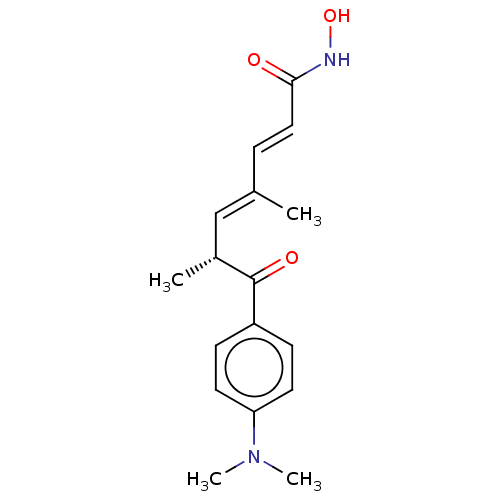
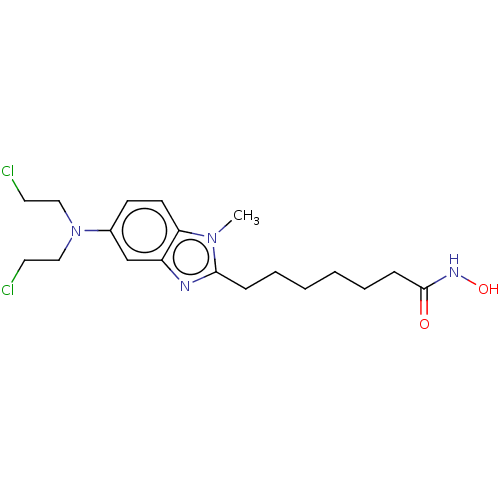
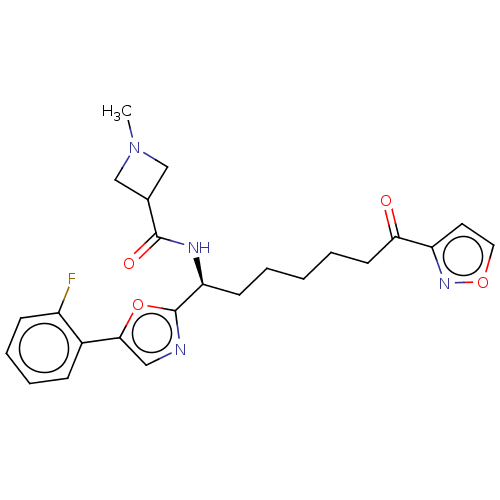

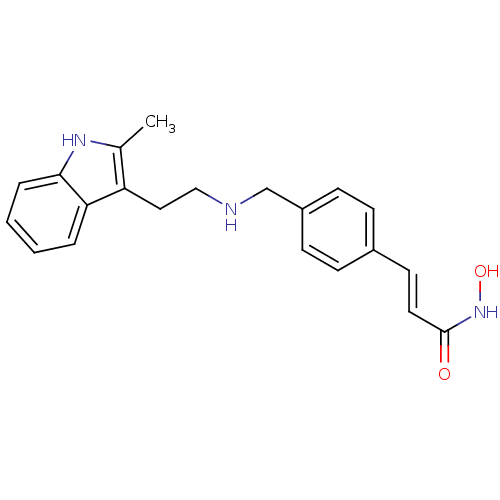
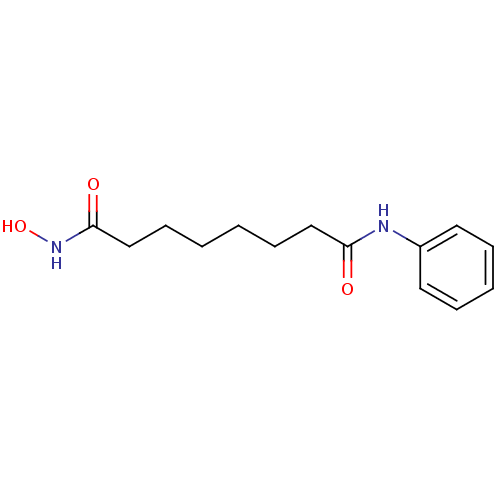
 3D Structure (crystal)
3D Structure (crystal)