Report error Found 2625 Enz. Inhib. hit(s) with Target = 'Metabotropic glutamate receptor 1'
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
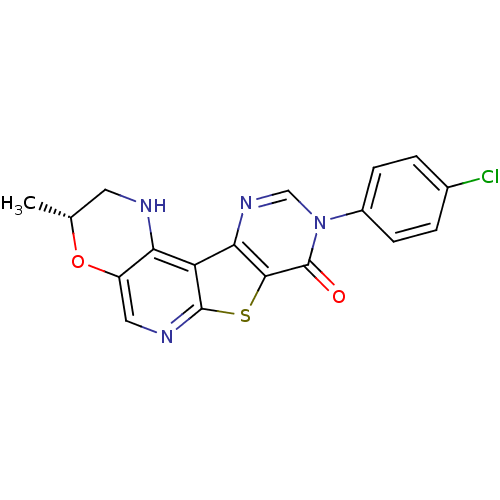
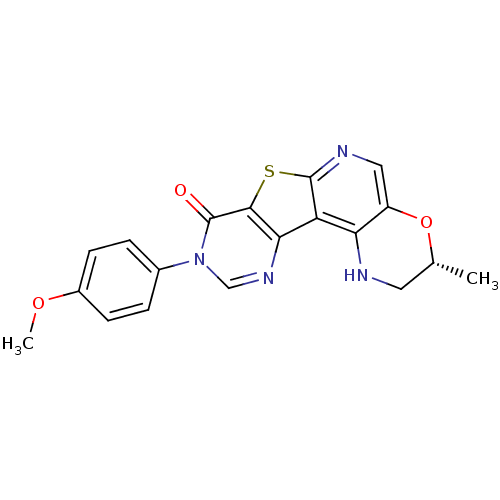
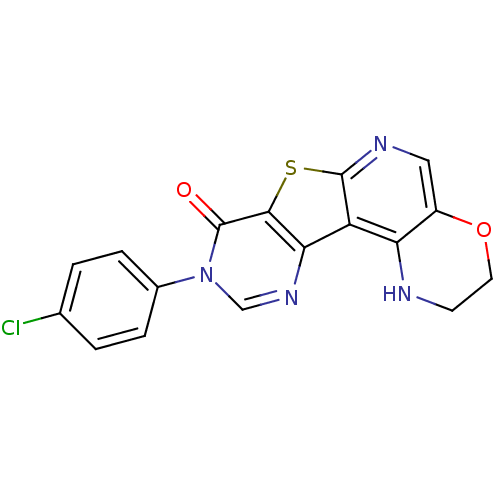
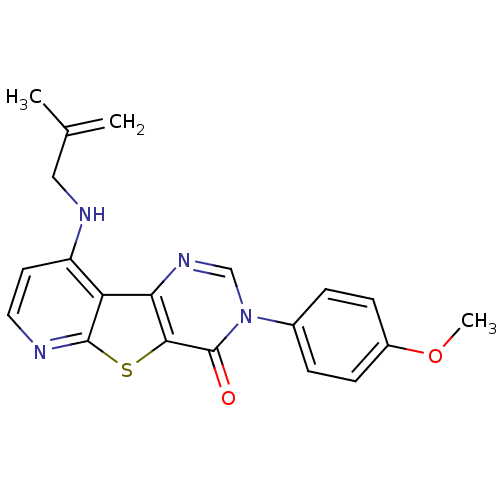
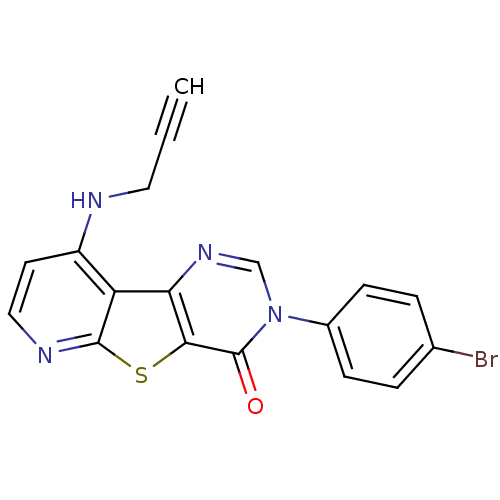
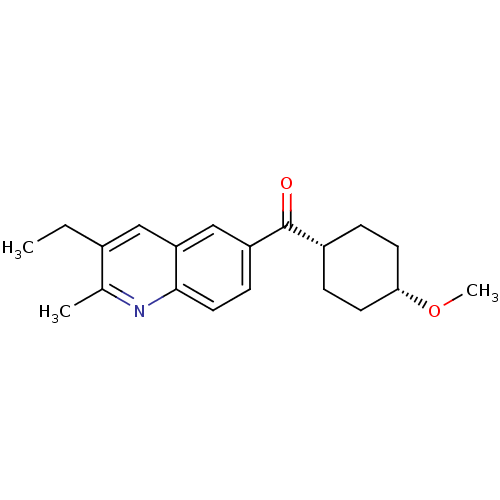
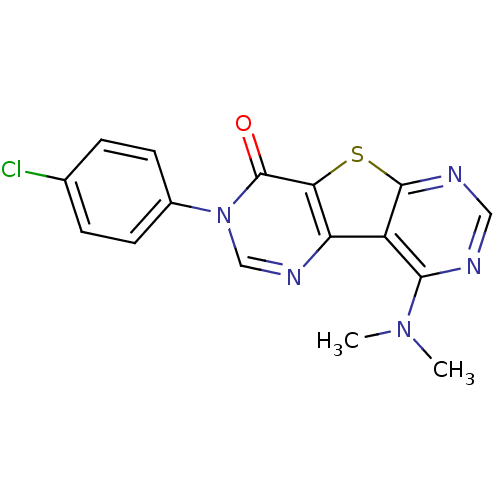
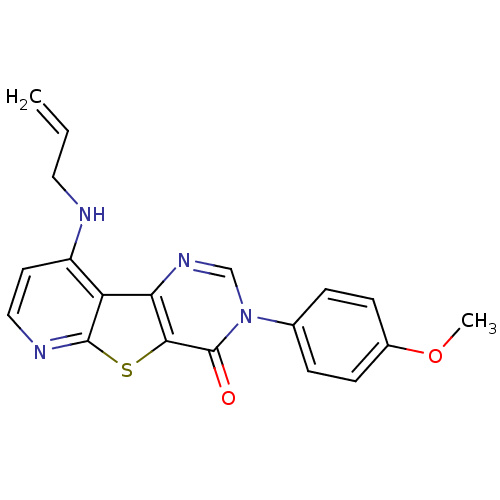
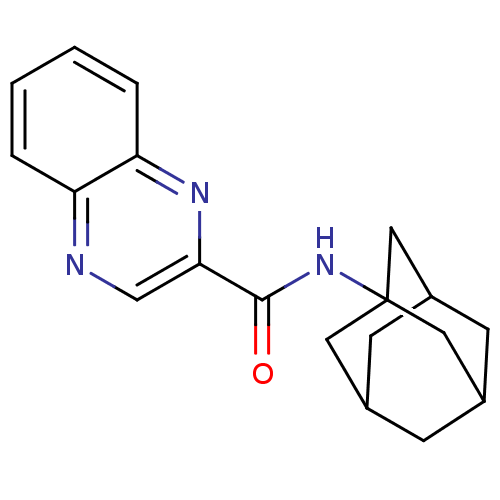
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.200nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataKi: 0.340nMAssay Description:Binding affinity to mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Binding affinity to mGluR1 by PET analysisMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Binding affinity to mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Antagonist activity at human metabotropic glutamate receptor 1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:Antagonist activity at human mGluR1a expressed in CHO cells assessed by measuring intracellular calcium by FLIPR assayMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.400nMAssay Description:Antagonist activity at rat metabotropic glutamate receptor 1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.400nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.550nMAssay Description:Inhibitory concentration against human metabotropic glutamate receptorMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.550nMAssay Description:Inhibitory concentration against human metabotropic glutamate receptorMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.560nMAssay Description:Inhibitory concentration against human metabotropic glutamate receptorMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.600nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.600nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.700nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.700nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.700nMAssay Description:Antagonist activity at human mGluR1a expressed in CHO cells assessed by measuring intracellular calcium by FLIPR assayMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.800nMAssay Description:Antagonist activity at human metabotropic glutamate receptor 1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 0.870nMAssay Description:In vitro affinity for cloned rat metabotropic glutamate 1 receptors stably expressed on CHO cells determined using [3H]-R214127 as radioligandMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataKi: 0.870nMAssay Description:Binding affinity to mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataKi: 0.870nMAssay Description:Binding affinity to mGluR1 by PET analysisMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 0.900nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataIC50: 0.900nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibitory concentration against human metabotropic glutamate receptorMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataIC50: 1nMAssay Description:Inhibition of rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataIC50: 1nMAssay Description:Antagonist activity at rat mGluR1 expressed in 1321N1 cellsMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1nMAssay Description:Displacement of [3H]R214127 from mGluR1 in rat cerebellum membranesMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1nMAssay Description:Displacement of [3H]9-Dimethylamino-3-(4-methoxy-phenyl)-3H-pyrido[3',2':4,5]thieno[3,2-d]pyrimidin-4-one from mGluR1 in rat cerebellum membraneMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human mGluR1 expressed in 1321N1 cells assessed as effect on L-glutamate-induced calcium mobilizationMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1.10nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1.10nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1.20nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1.21nMAssay Description:Antagonist activity at human mGlu1 receptorMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1.40nMAssay Description:Antagonist activity at human mGluR1a expressed in CHO cells assessed by measuring intracellular calcium by FLIPR assayMore data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Mouse)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1.40nMAssay Description:Binding affinity to mouse mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Human)
National Institute of Radiological Sciences
Curated by ChEMBL
National Institute of Radiological Sciences
Curated by ChEMBL
Affinity DataIC50: 1.5nMAssay Description:Antagonist activity at human mGluR1More data for this Ligand-Target Pair
TargetMetabotropic glutamate receptor 1(Rat)
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Johnson and Johnson Pharmaceutical Research and Development, Beerse
Curated by PDSP Ki Database
Affinity DataKi: 1.5nMAssay Description:Antagonist activity at rat mGluR1More data for this Ligand-Target Pair
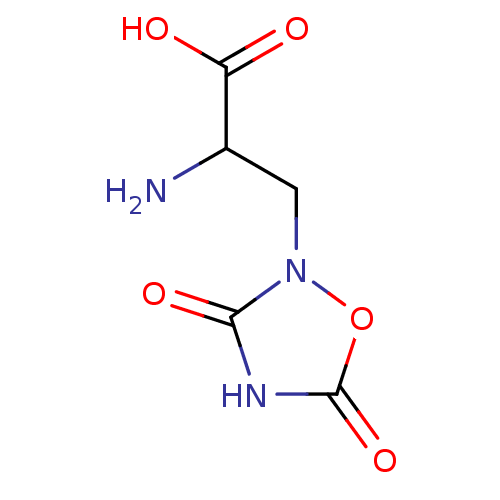
 3D Structure (crystal)
3D Structure (crystal)