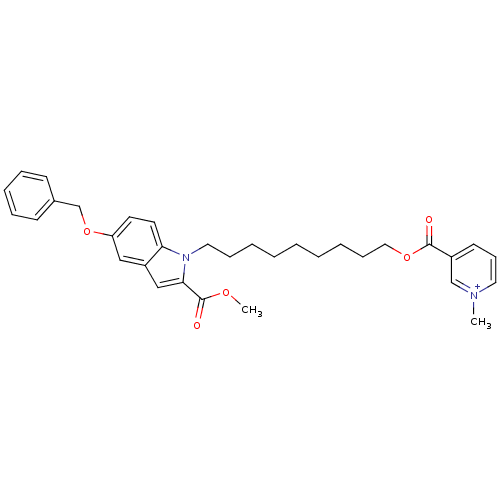
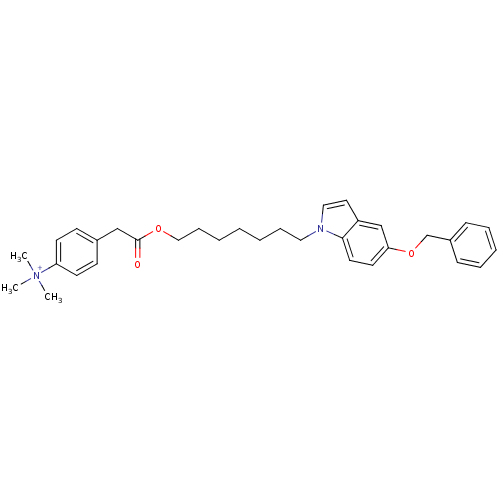
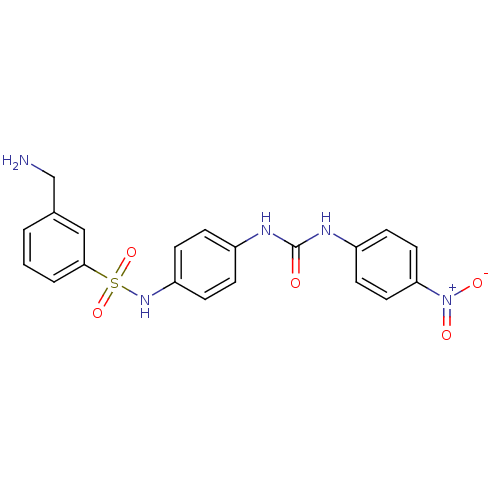
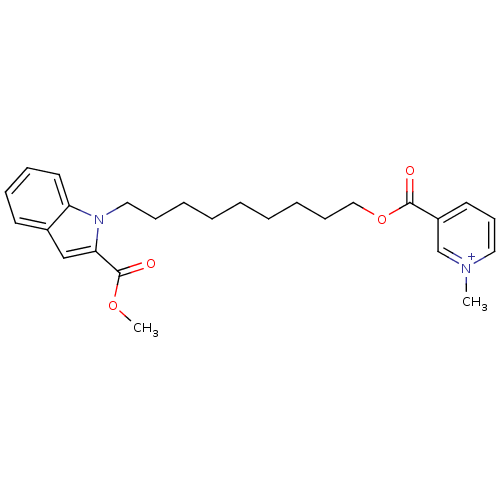
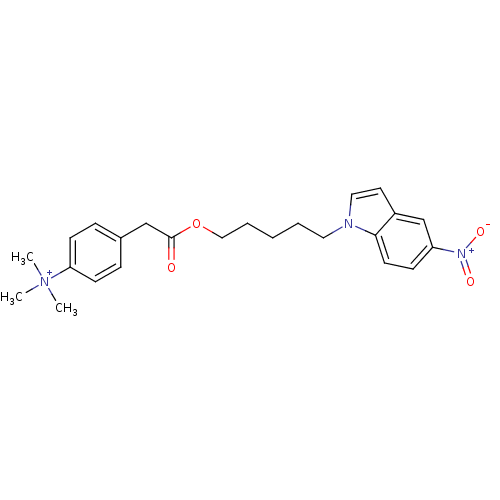
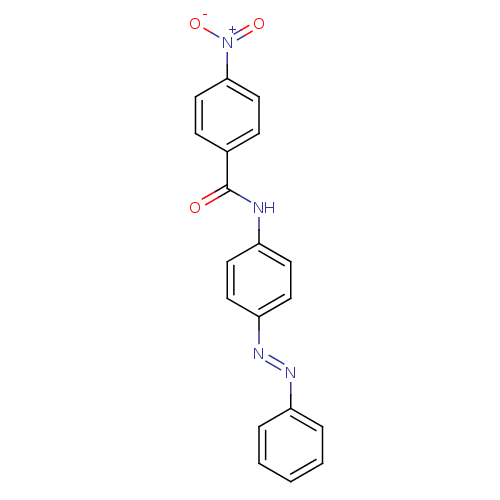
Report error Found 175 Enz. Inhib. hit(s) with Target = 'NH(3)-dependent NAD(+) synthetase'
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
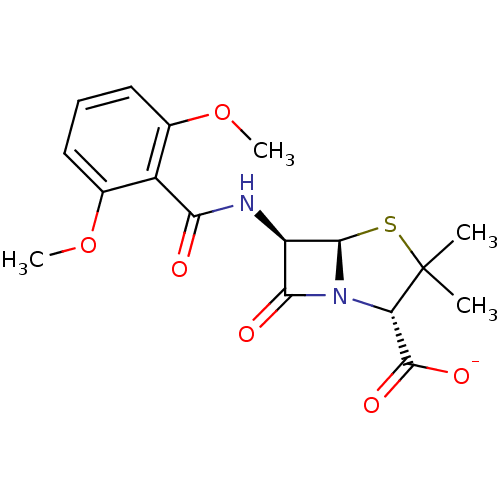
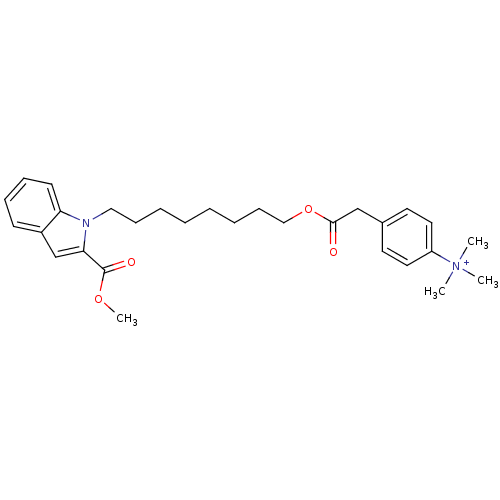
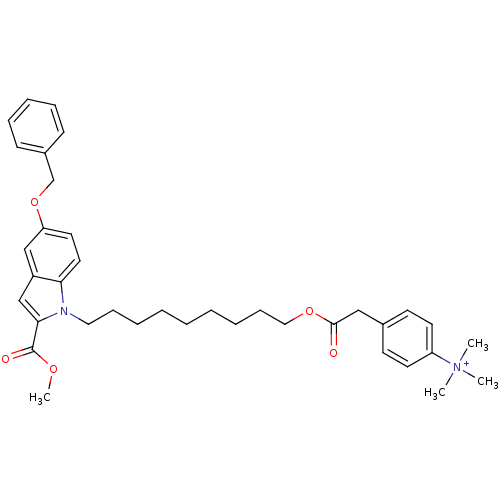
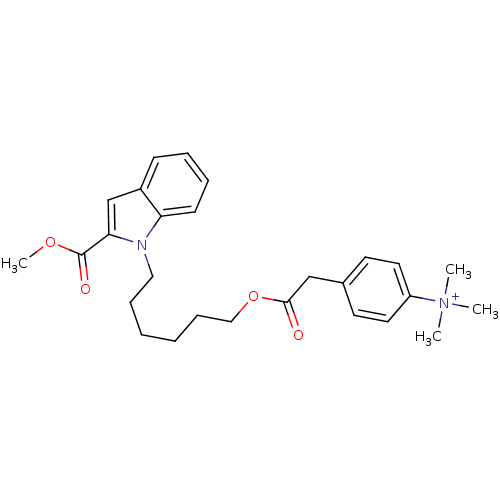
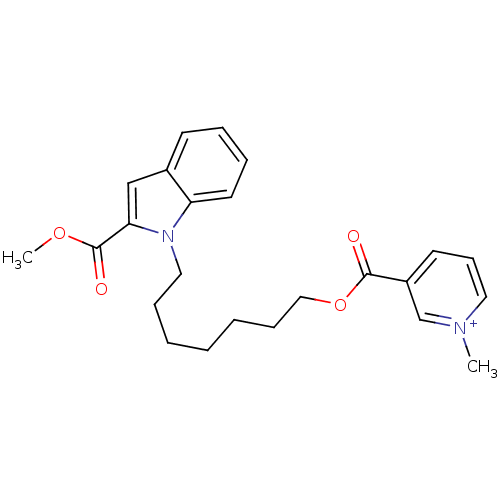
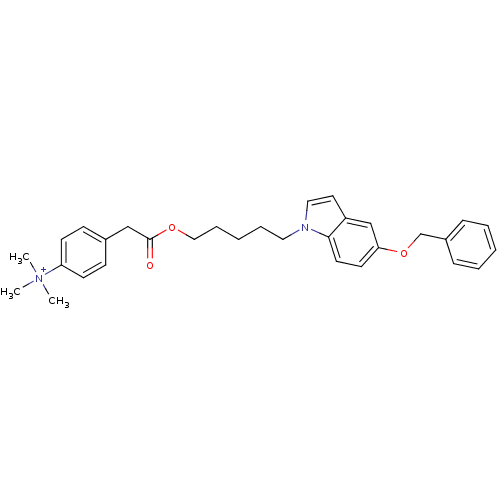
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
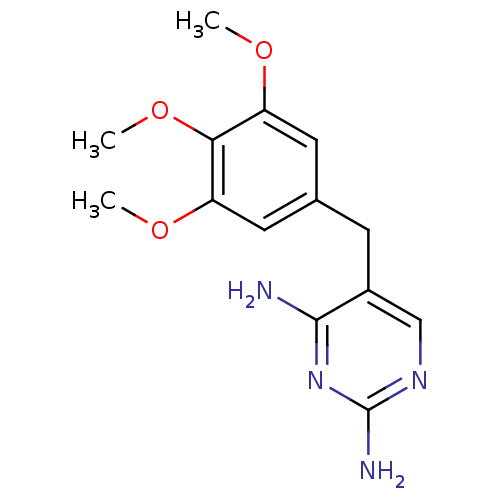
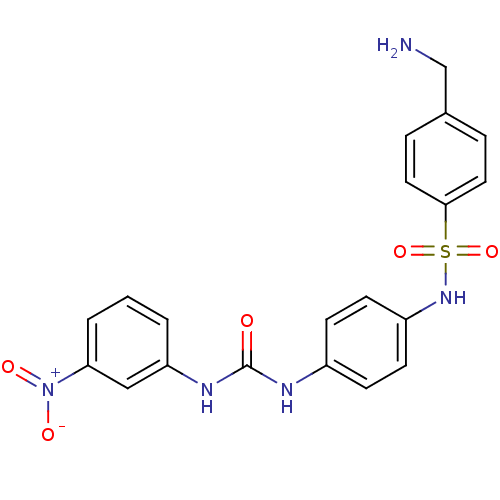
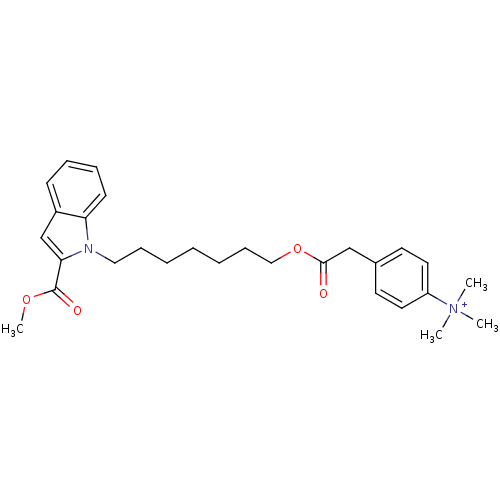
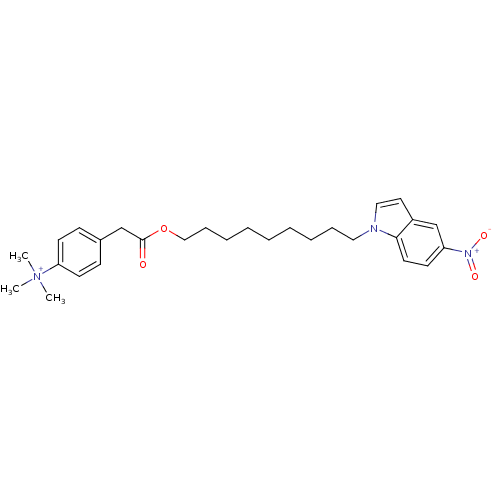
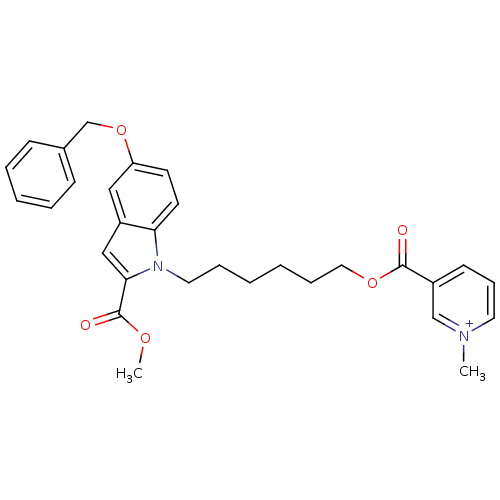
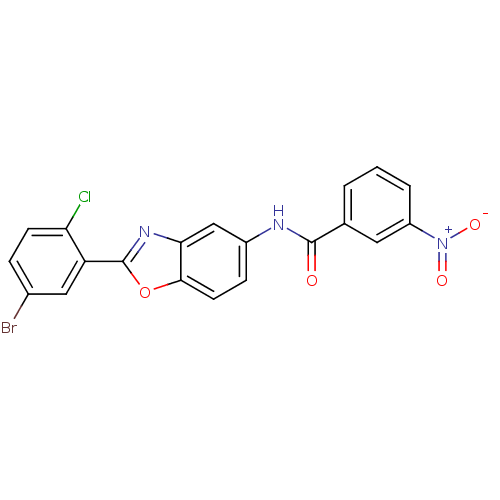
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
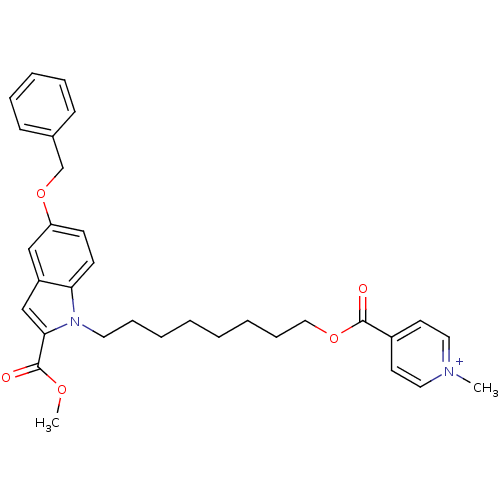
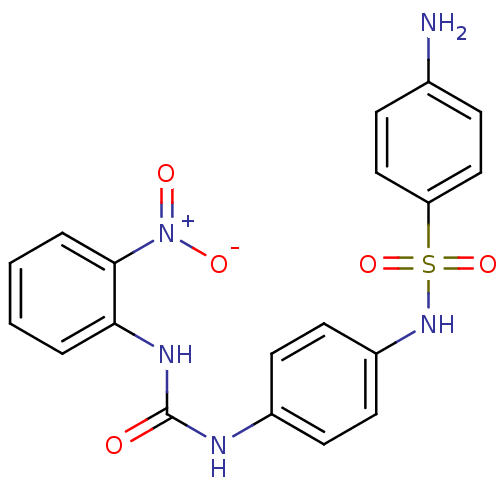
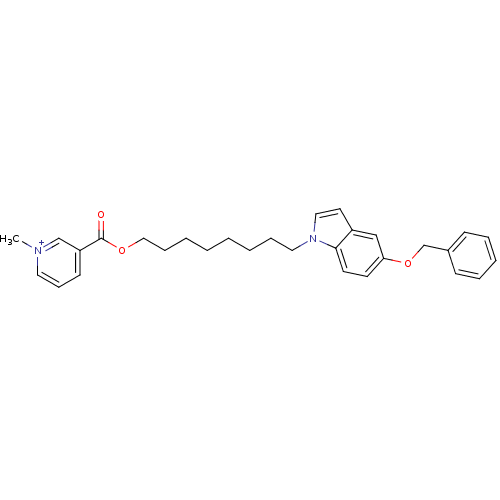
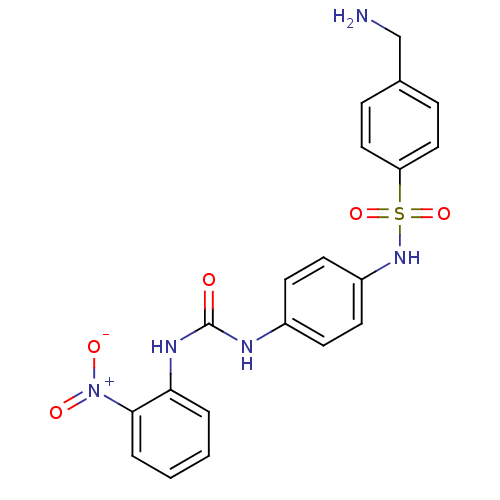
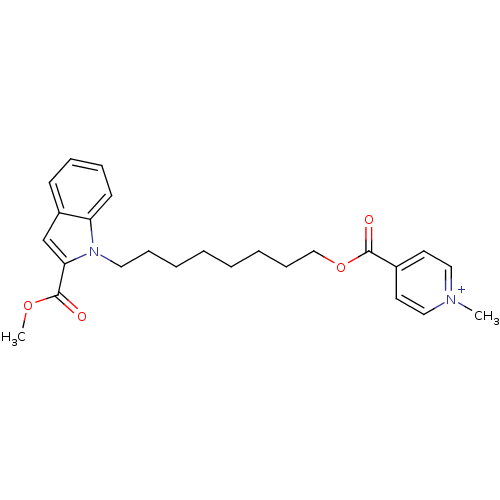
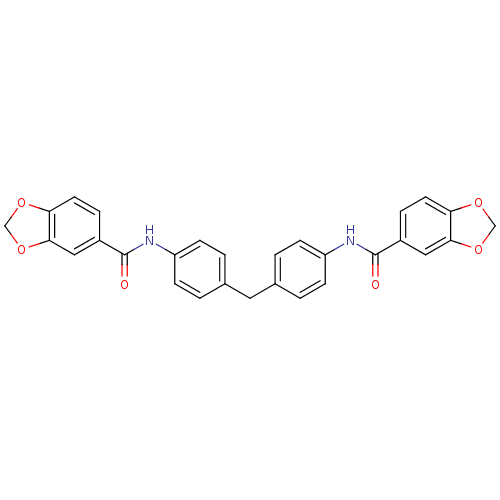
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
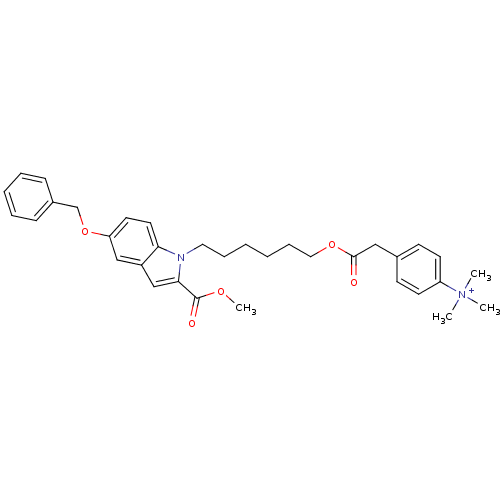
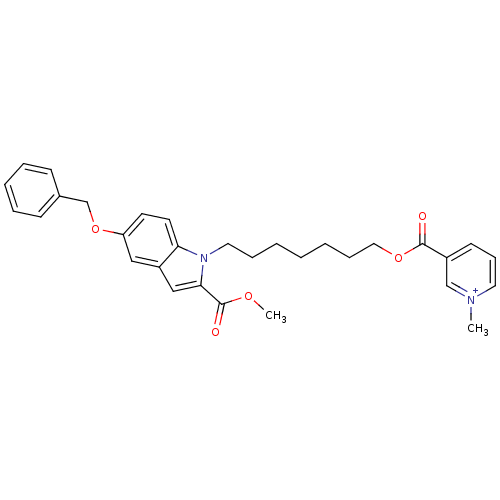
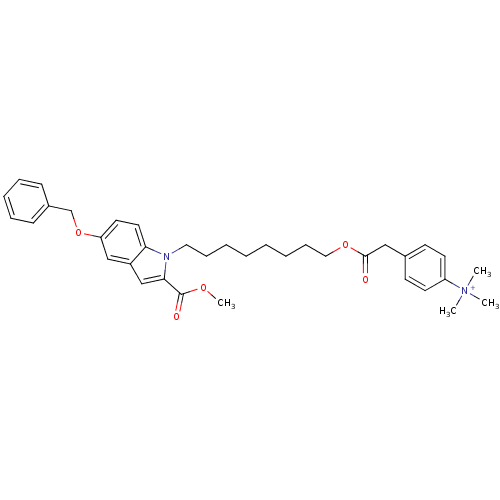
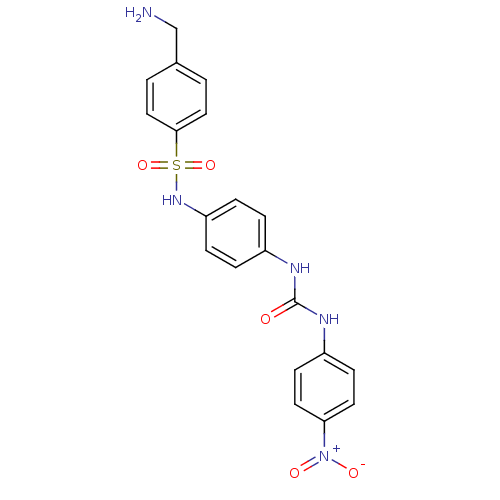
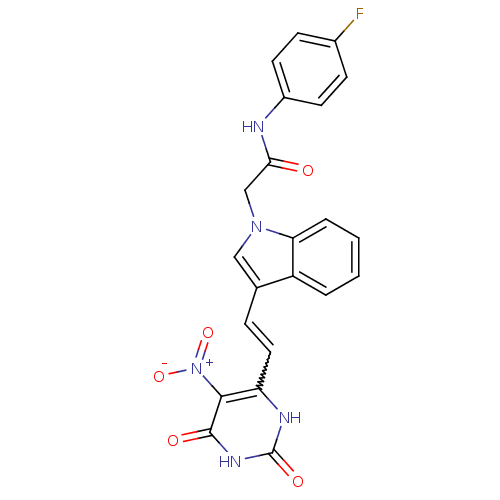
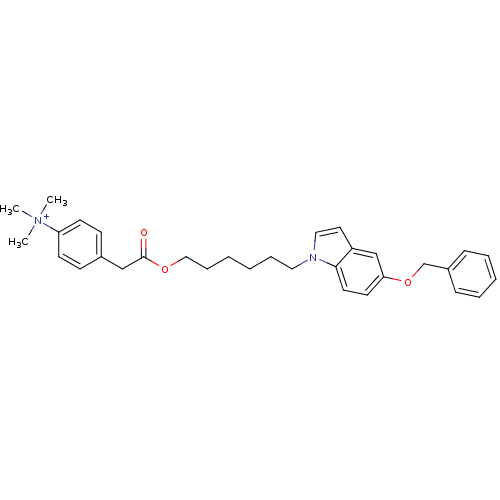
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.71E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 1.90E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.25E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.27E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.32E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 3.70E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 4.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.54E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.18E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 6.30E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.57E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 6.90E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
TargetNH(3)-dependent NAD(+) synthetase(Bacillus subtilis (strain 168))
University of Alabama At Birmingham
Curated by ChEMBL
University of Alabama At Birmingham
Curated by ChEMBL
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibitory activity against Bacillus subtilis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.60E+4nMAssay Description:Inhibition of Bacillus anthracis NAD synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.75E+4nMpH: 8.5 T: 2°CAssay Description:Inhibitory concentration of the compound measured against B. anthracis Nicotinamide adenine dinucleotide synthetase (NADs) using HPLC Enzyme AssayMore data for this Ligand-Target Pair