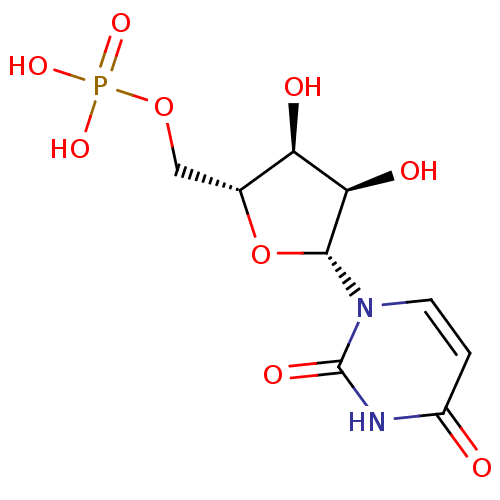
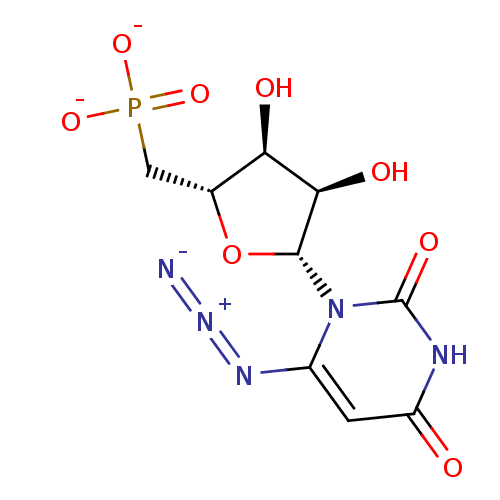
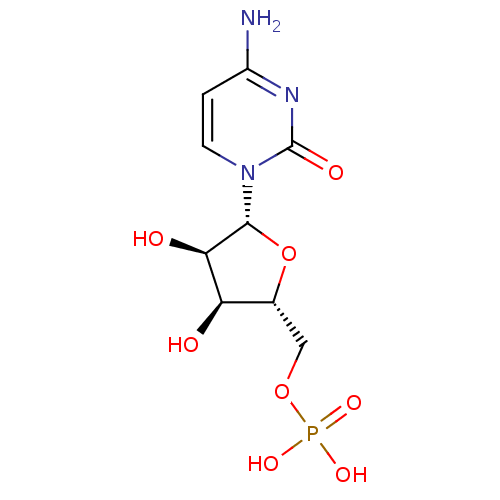
Report error Found 33 Enz. Inhib. hit(s) with Target = 'Orotidine 5'-phosphate decarboxylase'
Affinity DataKi: 0.00880nMAssay Description:Inhibition of Saccharomyces cerevisia uridine 5'-monophosphate synthaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.00900nM ΔG°: -63.0kJ/moleT: 2°CAssay Description:Inhibition of Saccharomyces cerevisia uridine 5'-monophosphate synthase after overnight incubation at room temperature by VP-ITC microcalorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 5nM ΔG°: -47.4kJ/moleT: 2°CAssay Description:Inhibition of Saccharomyces cerevisia uridine 5'-monophosphate synthase after overnight incubation at room temperature by VP-ITC microcalorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 64nMAssay Description:Inhibition of yeast ODCaseMore data for this Ligand-Target Pair
Affinity DataKi: 64nMAssay Description:Inhibition of Saccharomyces cerevisiae ODCase at 25 degreeC by competitive binding assayMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 190nM ΔG°: -42.2kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 200nM ΔG°: -42.1kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 360nM ΔG°: -40.5kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:Inhibition of yeast ODCaseMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 630nMAssay Description:Irreversible inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylaseMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 840nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase at 55 degreeC by competitive binding assayMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 840nM ΔG°: -38.2kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 840nM ΔG°: -38.2kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 6.20E+3nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCaseMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.10E+4nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCaseMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.11E+4nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.14E+4nM ΔG°: -31.1kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.24E+4nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase at 55 degreeC by competitive binding assayMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.24E+4nM ΔG°: -30.8kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 2.90E+4nM ΔG°: -28.5kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 2.90E+4nM ΔG°: -28.5kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 2.90E+4nM ΔG°: -28.5kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 2.90E+4nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase at 55 degreeC by competitive binding assayMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 3.70E+4nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylase by VP-ITC microcalorimeterMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.34E+5nM ΔG°: -24.3kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.34E+5nM ΔG°: -24.3kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 3.30E+5nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 3.60E+5nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylase by VP-ITC microcalorimeterMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 6.45E+5nM ΔG°: -20.0kJ/molepH: 7.5 T: 2°CAssay Description:An enzyme assay method using isothermal titration calorimetry (ITC) was developed to investigate the inhibition kinetics of ODCase. Titrations were p...More data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.20E+6nMAssay Description:Irreversible inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylaseMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 1.20E+6nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum ODCase by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 3.32E+6nMAssay Description:Inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylase by VP-ITC microcalorimeterMore data for this Ligand-Target Pair
TargetOrotidine 5'-phosphate decarboxylase(Methanobacterium thermoautotrophicum)
Toronto General Research Institute
Toronto General Research Institute
Affinity DataKi: 3.00E+7nMAssay Description:Irreversible inhibition of Methanobacterium thermoautotrophicum 5'-monophosphate decarboxylaseMore data for this Ligand-Target Pair


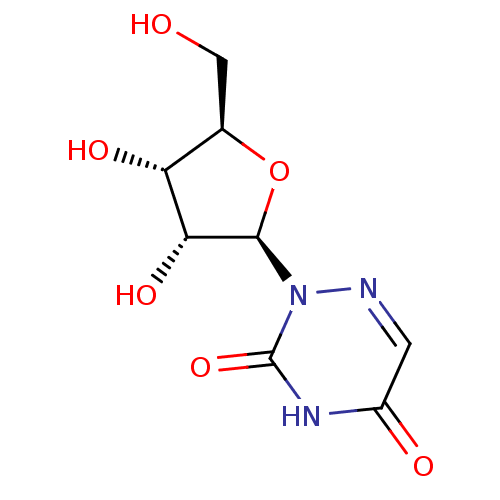

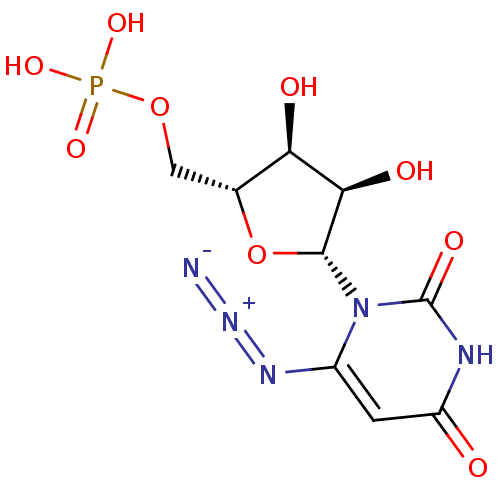
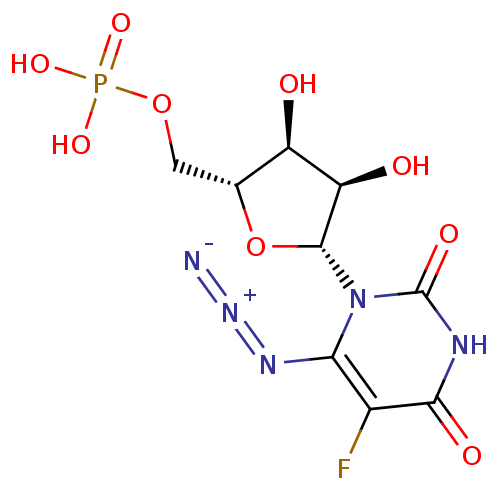
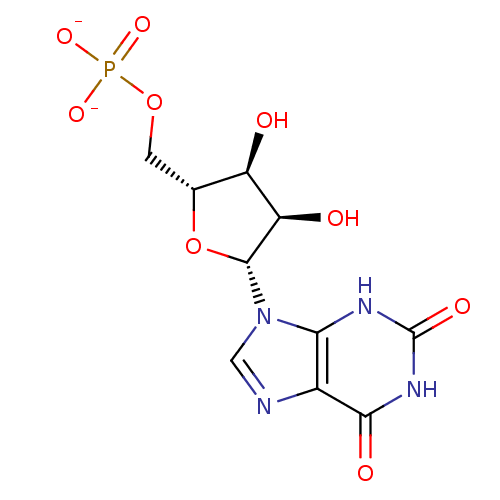


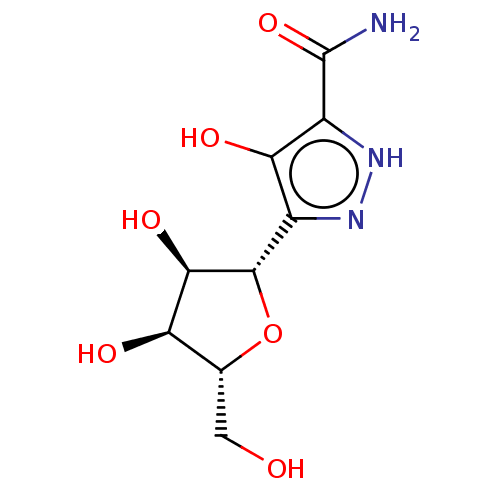

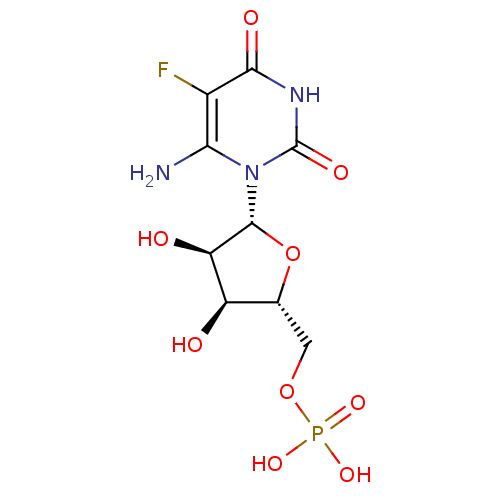



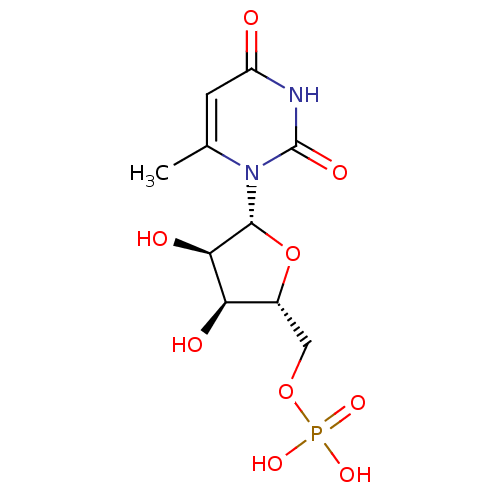
 3D Structure (crystal)
3D Structure (crystal)