Report error Found 39 Enz. Inhib. hit(s) with Target = 'Pepsin A-5'
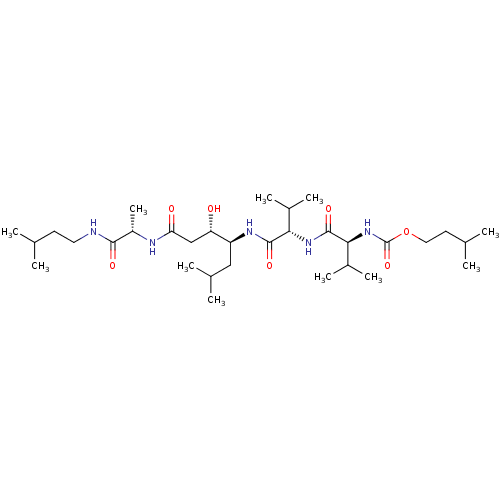
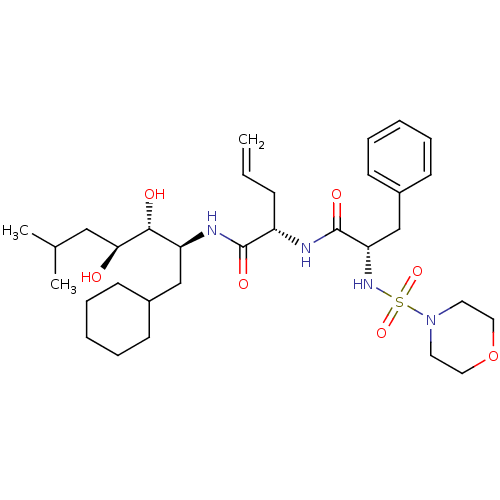
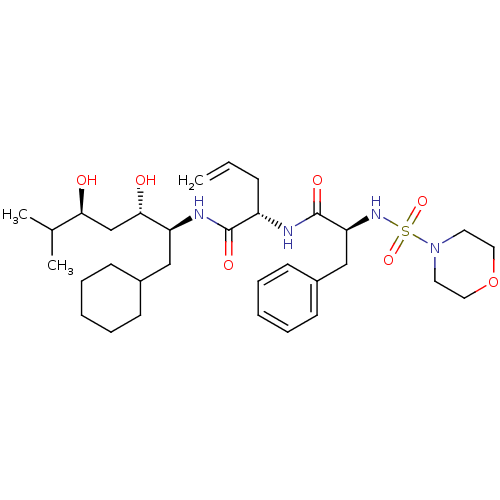
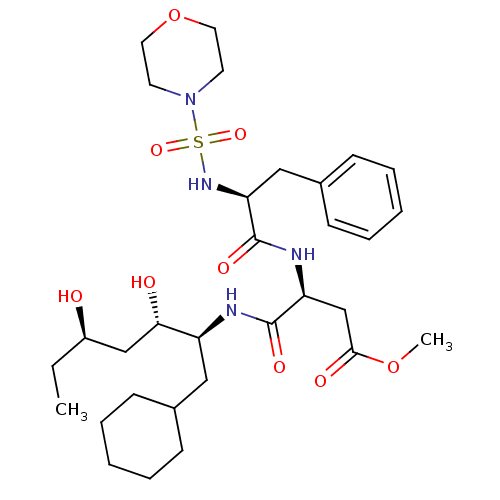
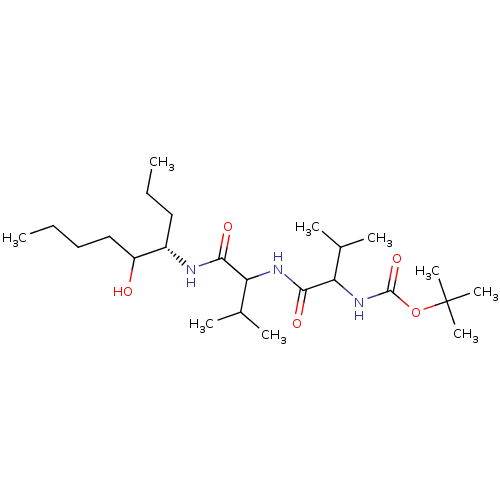
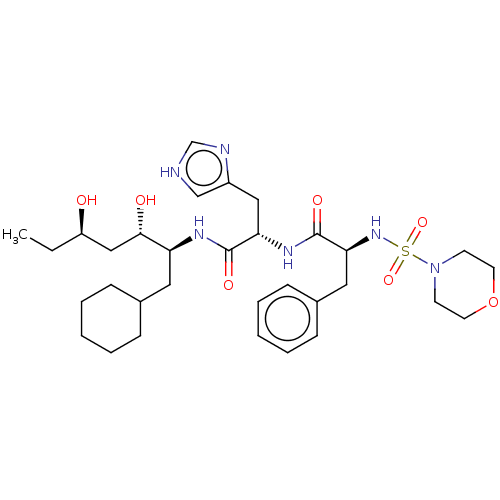
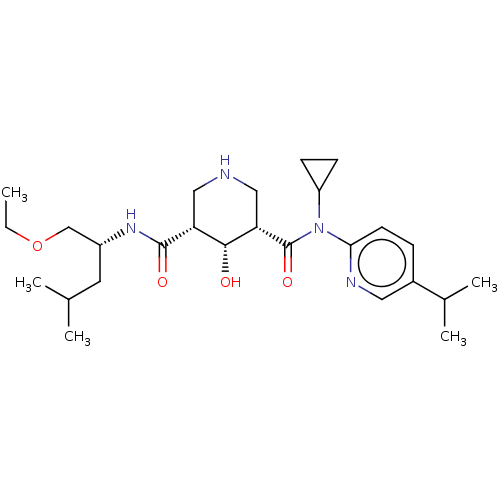
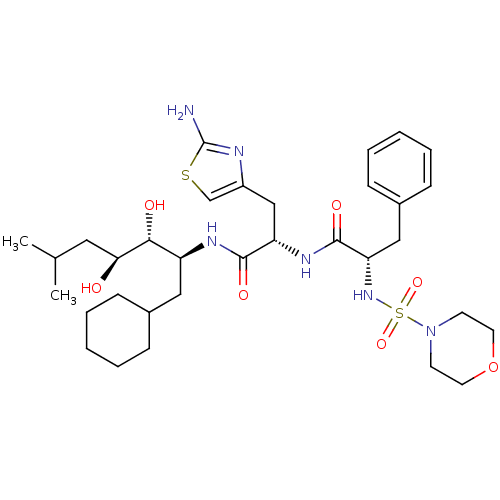
Affinity DataIC50: 4.20nMpH: 3.1Assay Description:Inhibition of human pepsin at pH 3.1More data for this Ligand-Target Pair
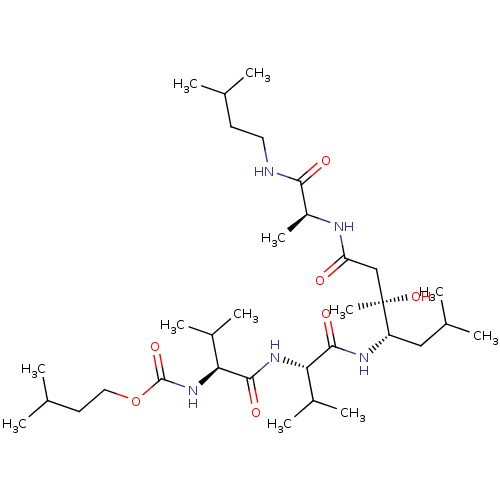
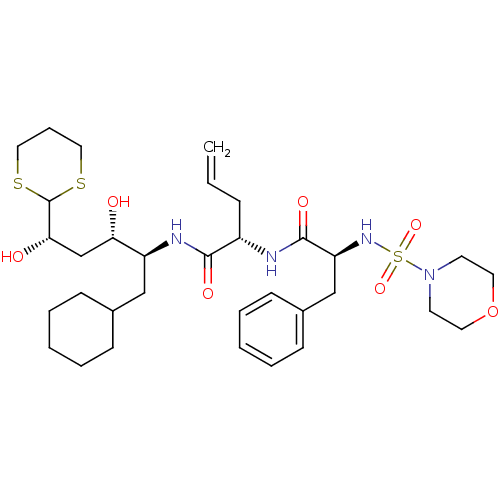
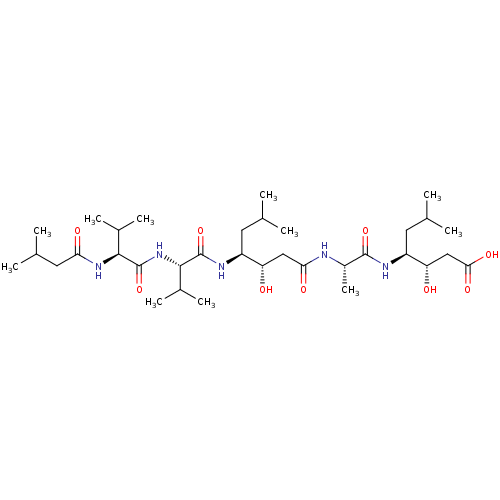
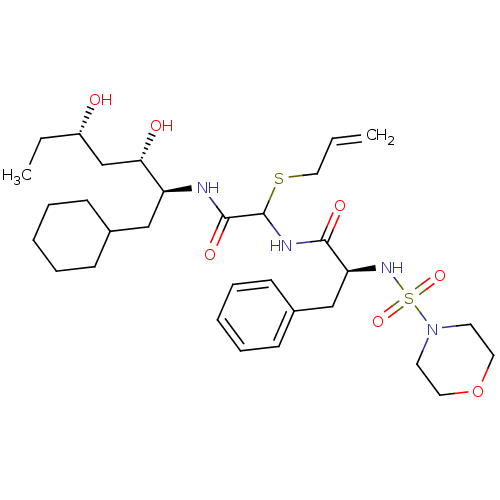
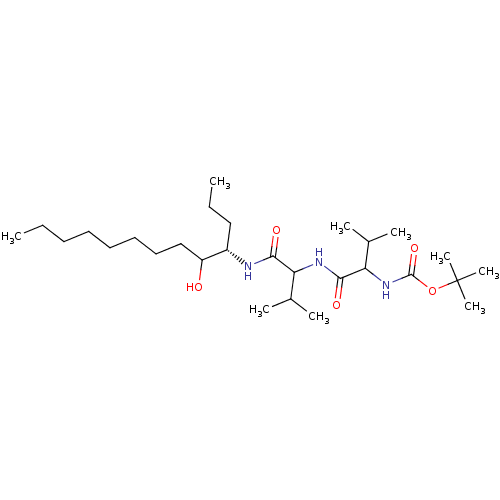
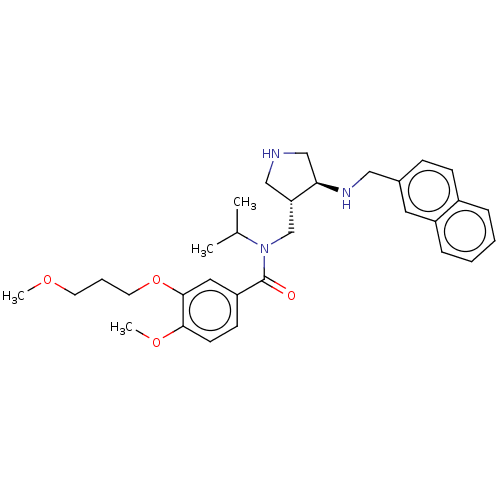
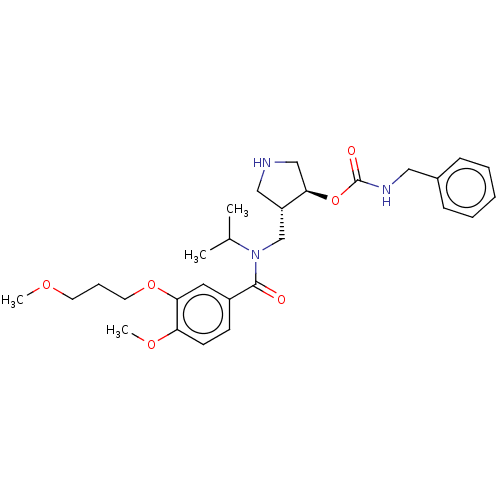
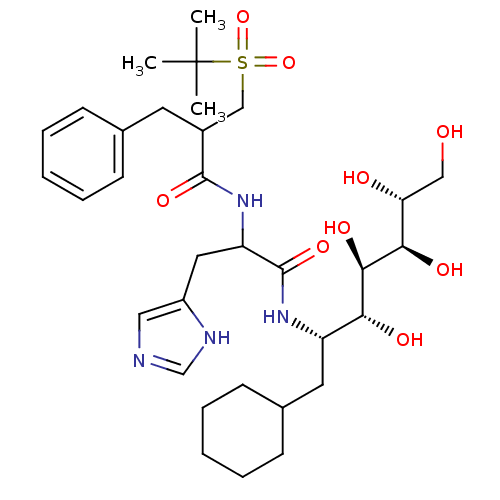
Affinity DataIC50: 6nMpH: 3.1Assay Description:Inhibition of human pepsin at pH 3.1More data for this Ligand-Target Pair
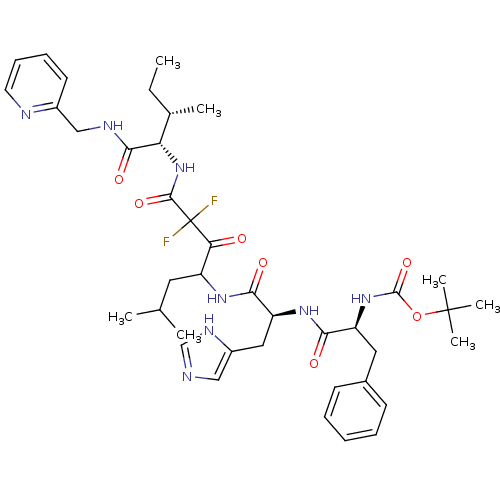
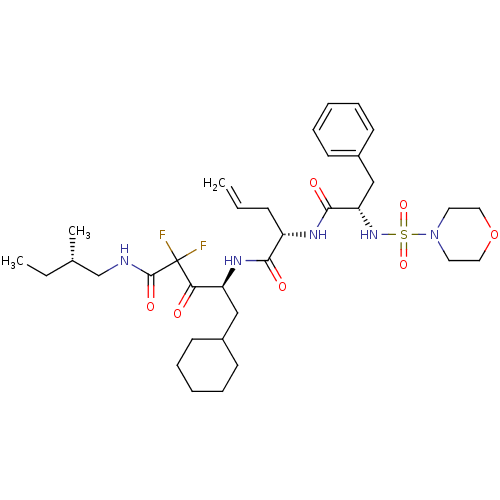
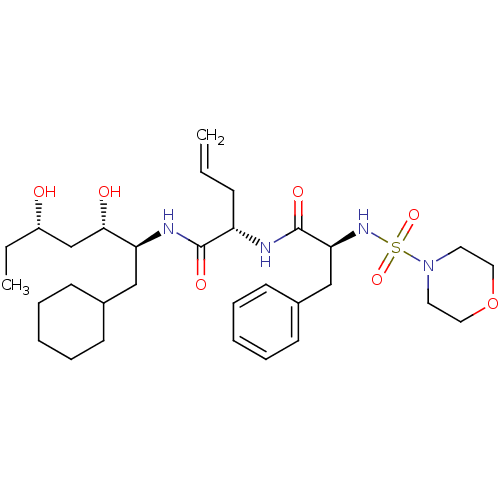
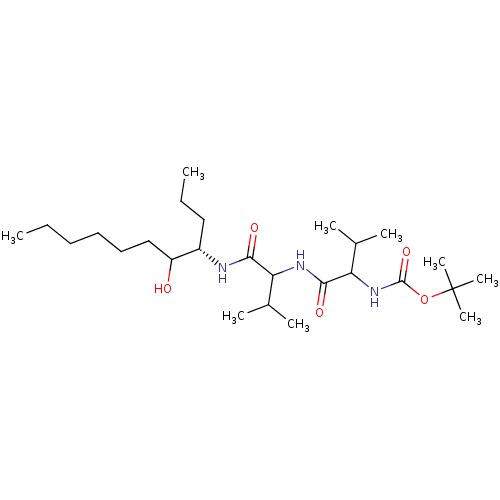
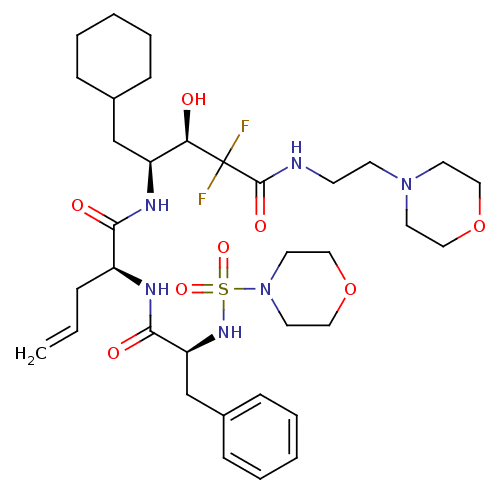
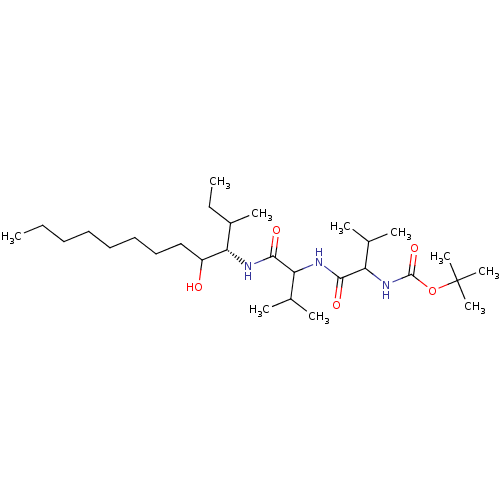
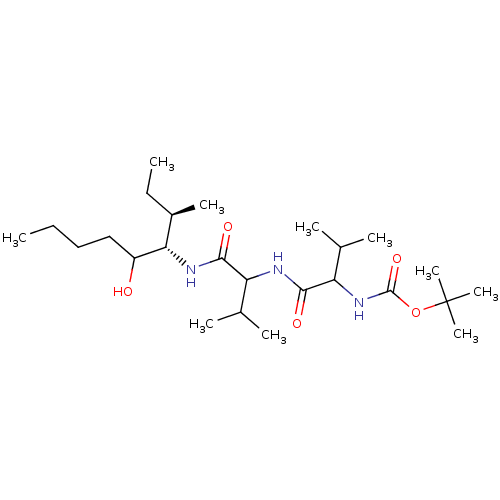
Affinity DataKi: 11nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
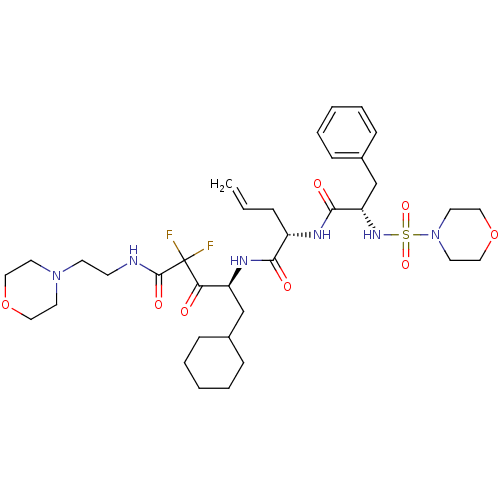
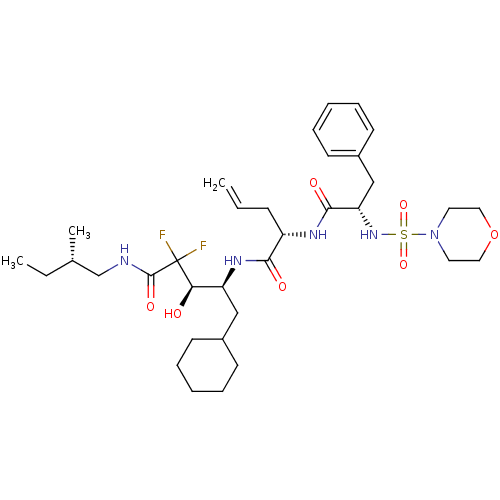
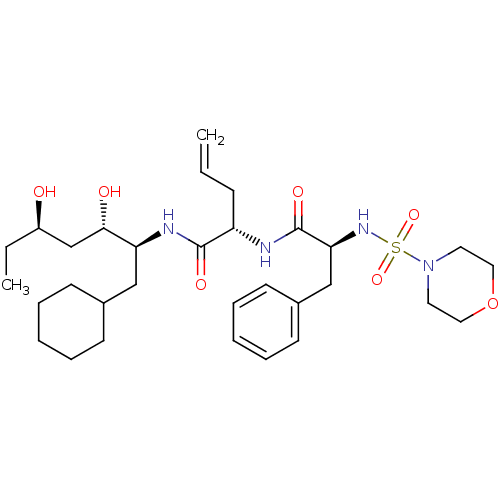
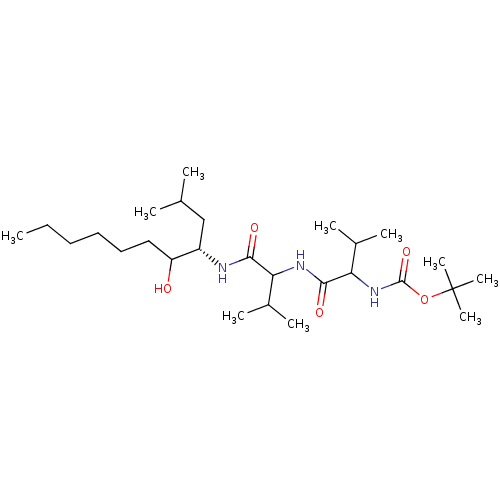
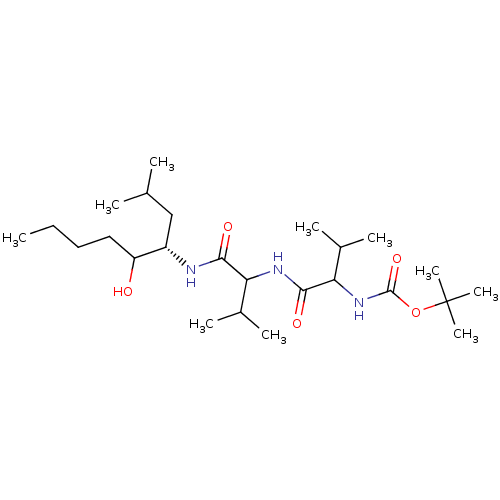
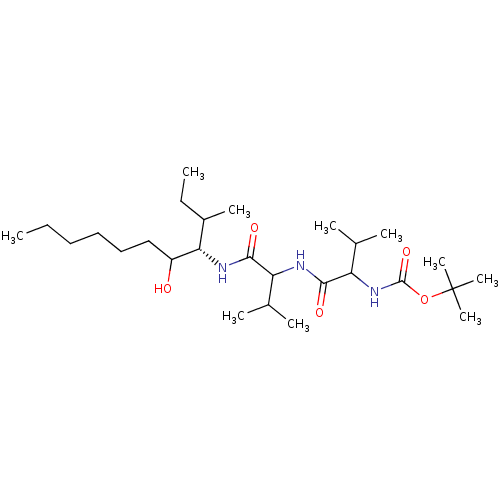
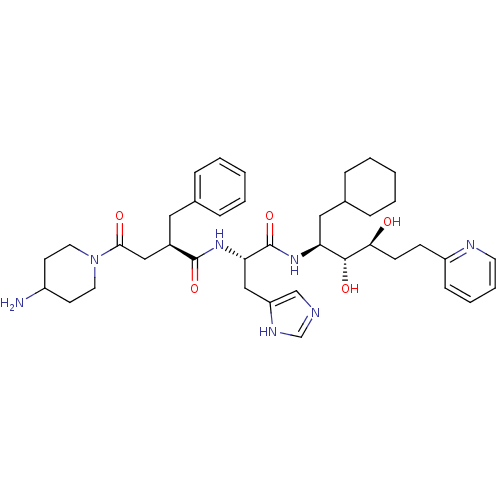
Affinity DataKi: 13nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataKi: 69nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 185nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 516nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 830nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 1.07E+3nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of pepsinMore data for this Ligand-Target Pair
Affinity DataKi: 1.24E+3nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+3nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Pepsin inhibition using synthetic heptapeptide substrate Phe-Gly-His-Phe-(N02)-Phe-Ala- Phe-OMeMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibitory concentration against pepsinMore data for this Ligand-Target Pair
Affinity DataKi: >6.00E+3nMAssay Description:Pepsin inhibition using synthetic heptapeptide substrate Phe-Gly-His-Phe-(N02)-Phe-Ala- Phe-OMeMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human pepsin AMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human recombinant pepsinogen A expressed in Escherichia coli using fluorescence-quenched Dabcyl-E-R-Nle-F-L-S-F-P-EDANS substrate by fl...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of pepsin A (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+4nMAssay Description:Inhibitory activity against pepsin as oxidation of o-phenylenediamine by Horse radish peroxidase (Pepsin sensitive)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Selectivity against human pepsinMore data for this Ligand-Target Pair
Affinity DataIC50: 2.02E+5nMAssay Description:Inhibitory activity against pepsin.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of pepsinMore data for this Ligand-Target Pair