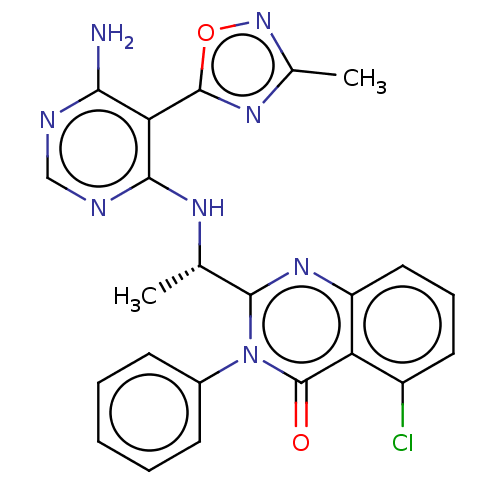
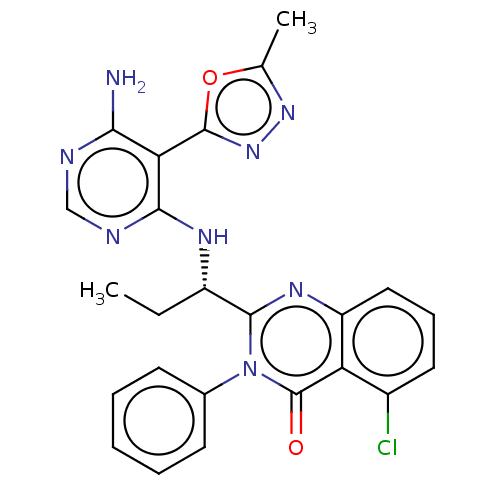
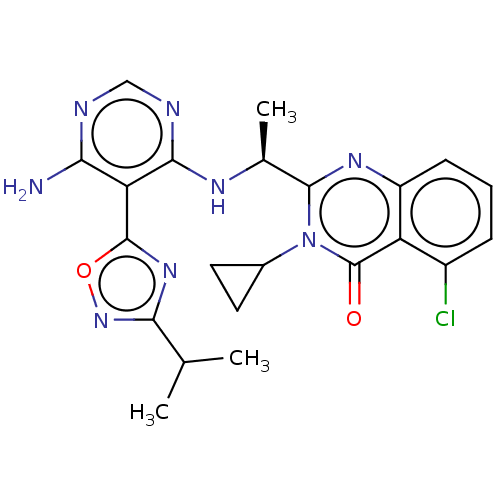
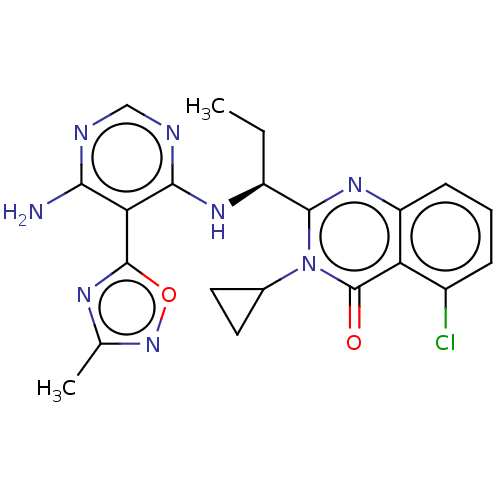
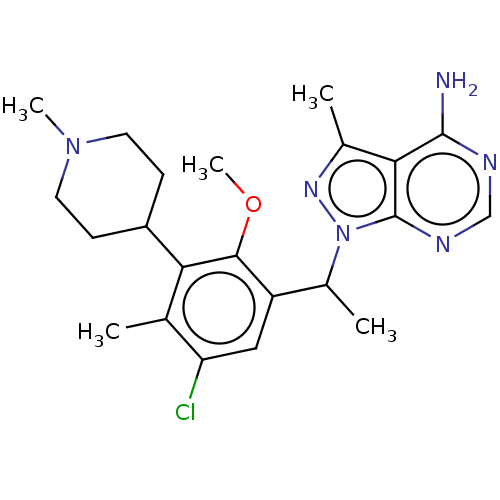
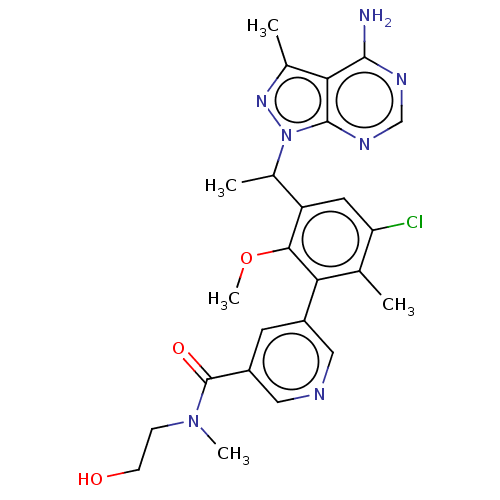
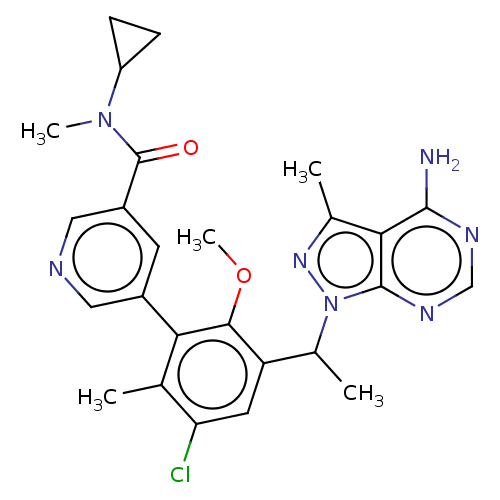
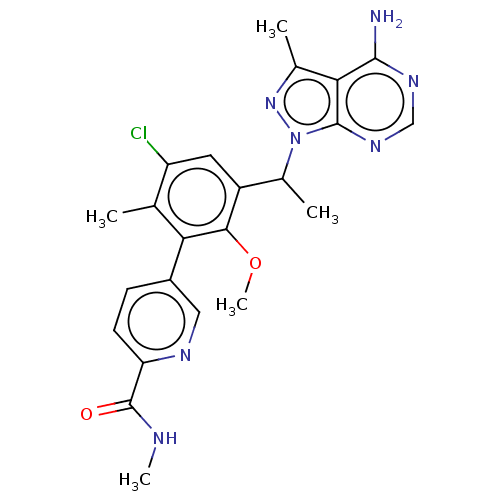
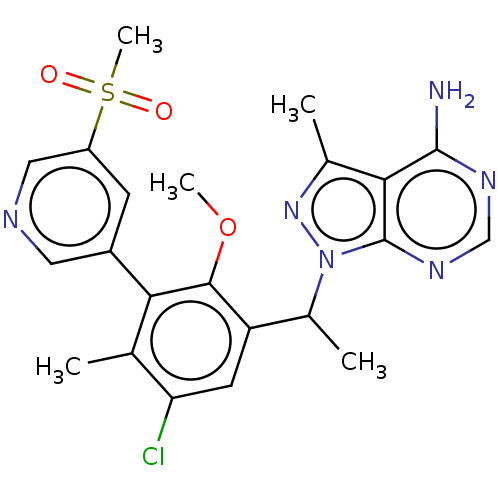
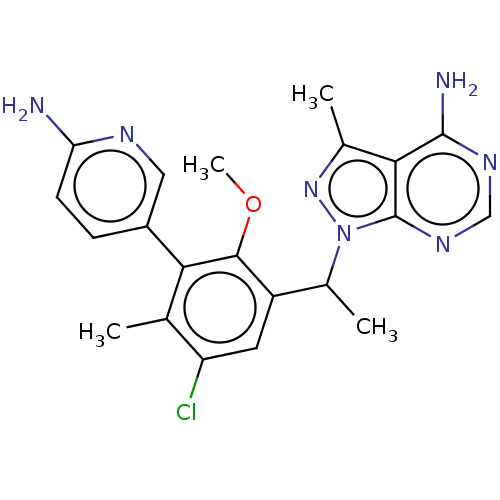
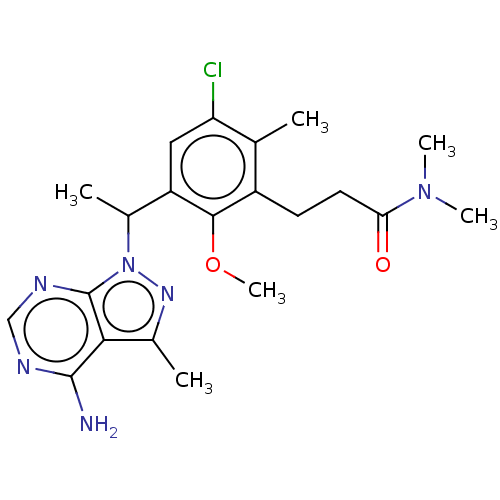
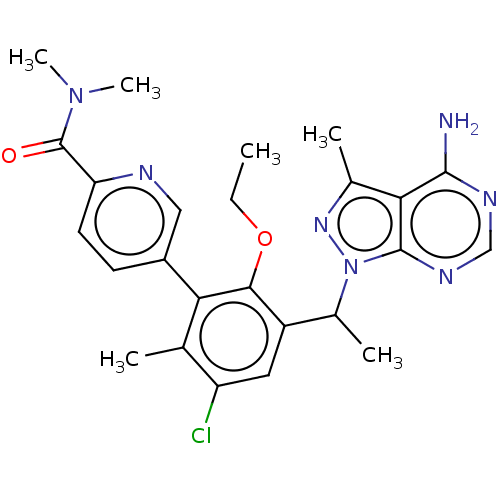
Report error Found 409 Enz. Inhib. hit(s) with Target = 'Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform'
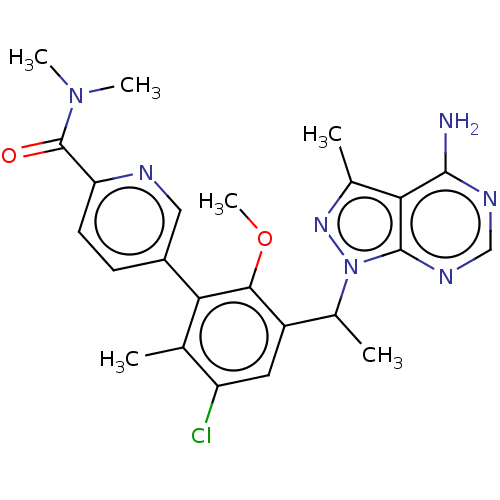
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
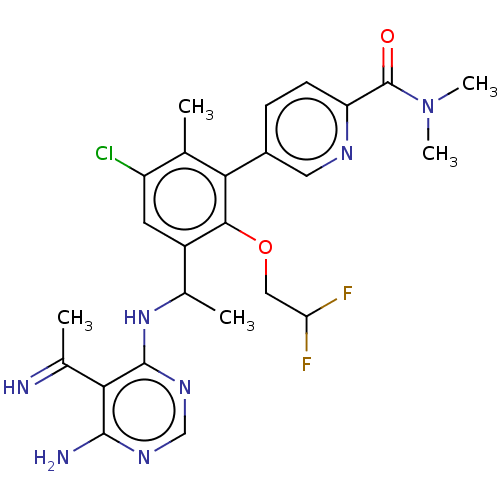
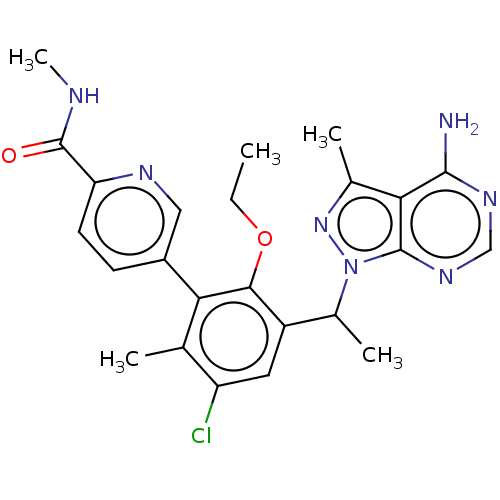
Affinity DataIC50: 2nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
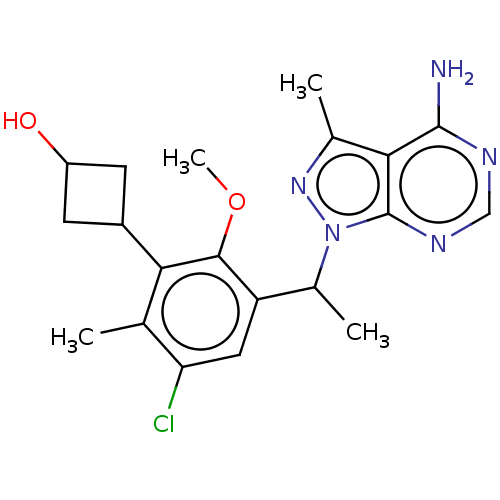
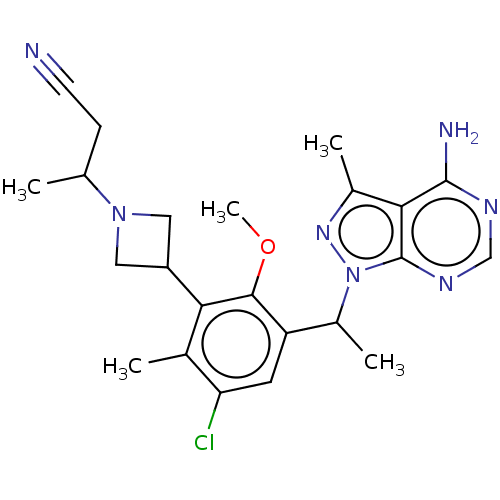
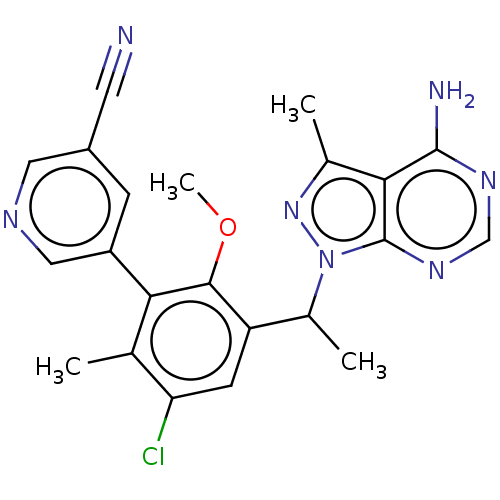
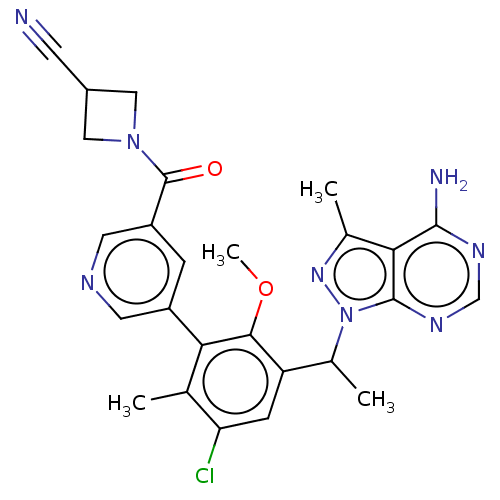
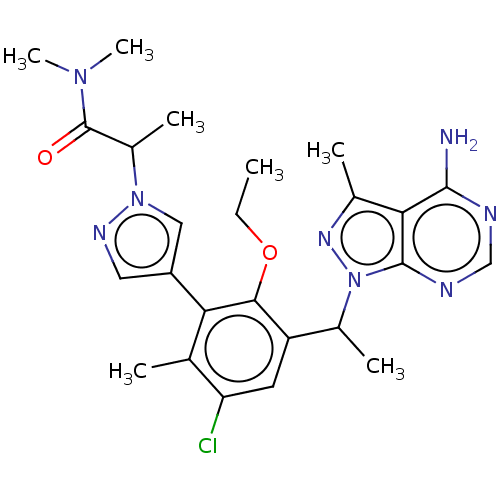
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
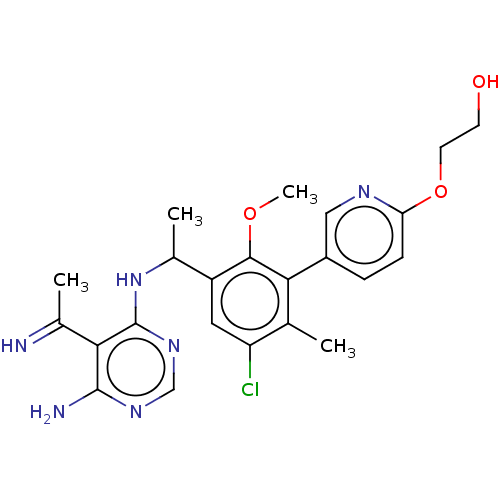
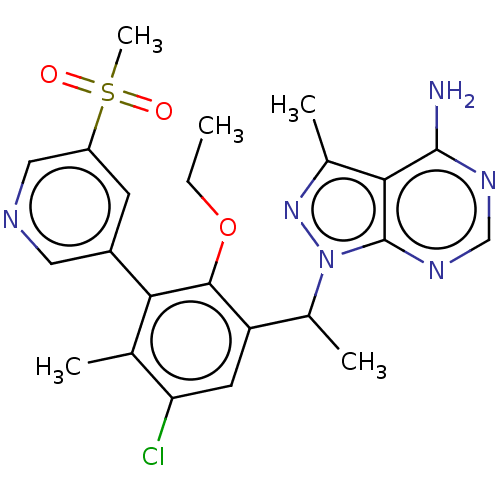
Affinity DataIC50: 2nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
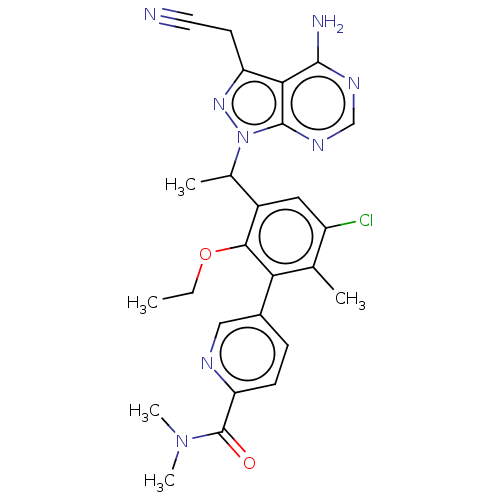
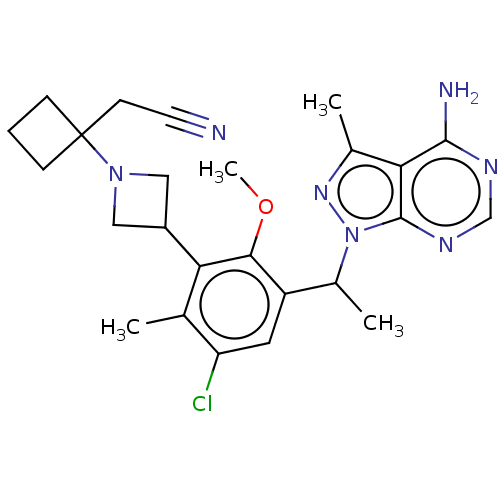
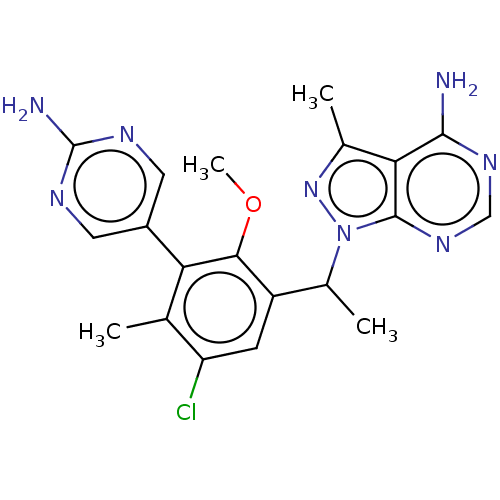
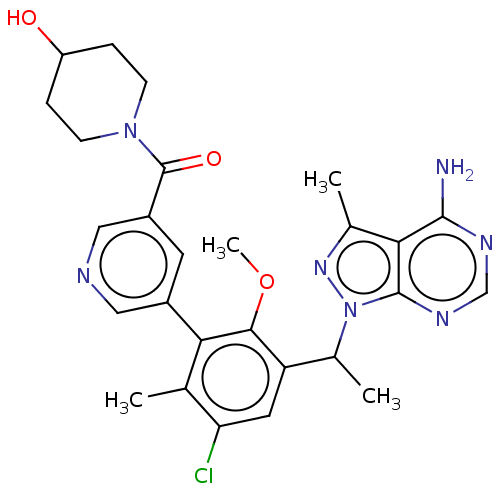
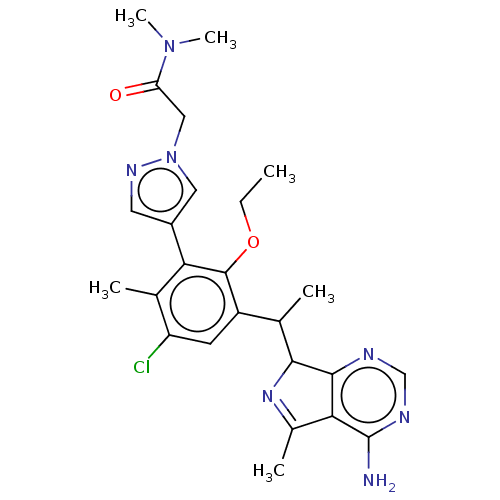
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
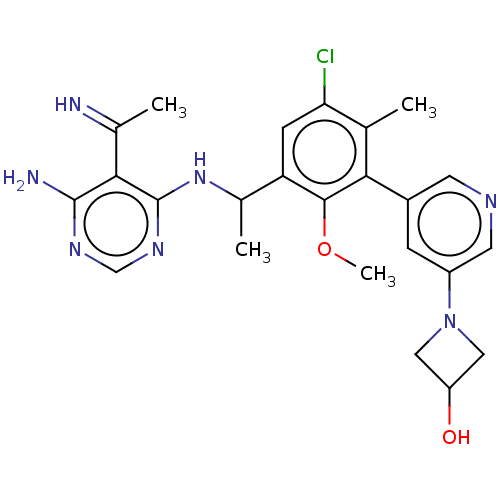
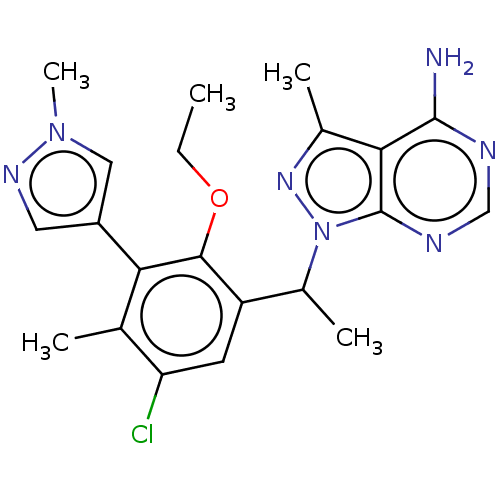
Affinity DataIC50: 3nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
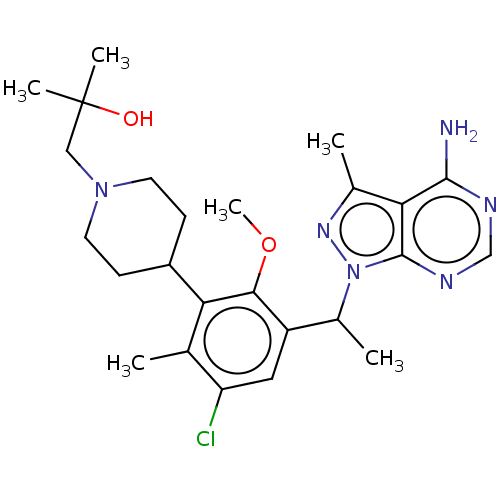
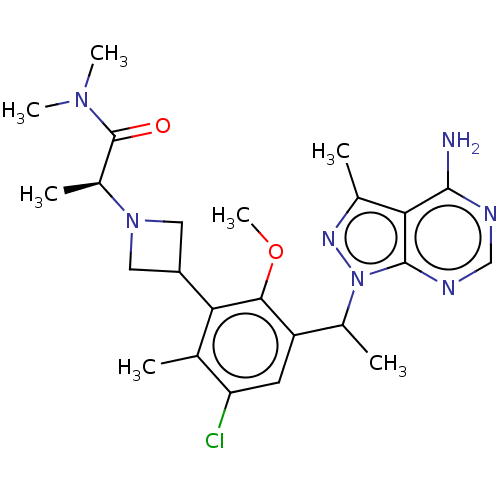
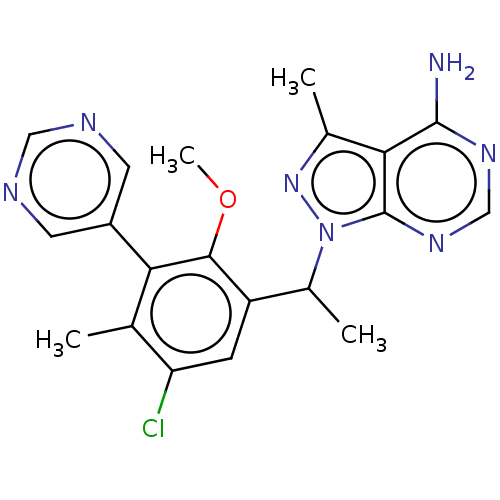
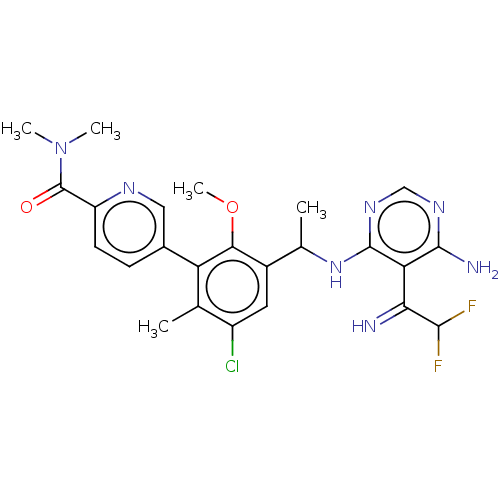
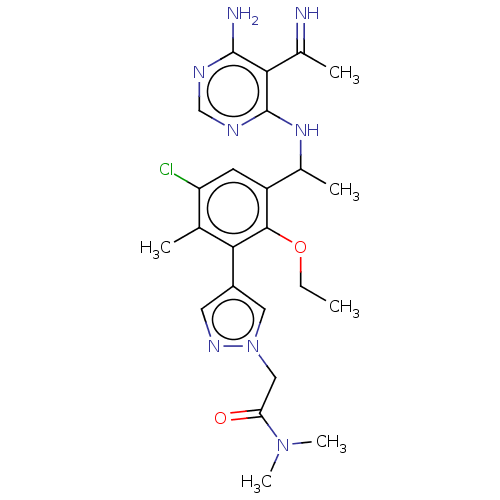
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
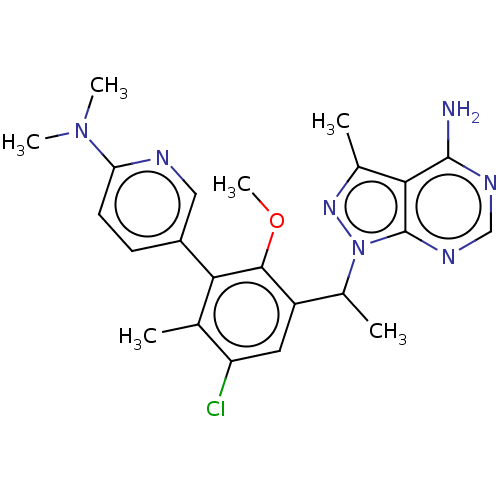
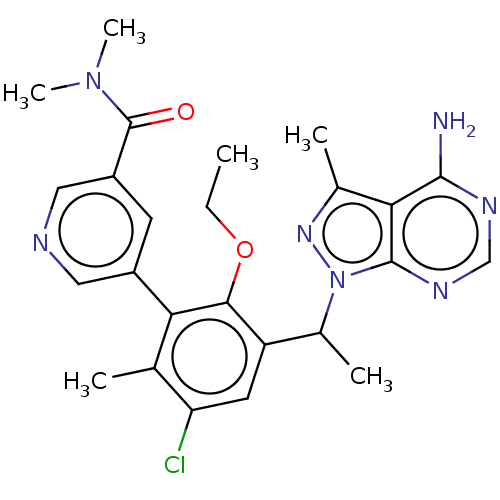
Affinity DataIC50: 5nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 6nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 6nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 8nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 8nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 9nMT: 2°CAssay Description:PI3K (p110α/p85α) (h) is incubated in assay buffer containing 10 μM phosphatidylinositol 4,5-bisphosphate and MgATP (concentration as ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta/delta isoform(Human)
Calitor Sciences
US Patent
Calitor Sciences
US Patent
Affinity DataIC50: 10nMpH: 6.7 T: 2°CAssay Description:Lipid kinase substrate, phosphoinositol-4,5-bisphosphate (PIP2), are purchased from Echelon Biosciences (Salt Lake City, Utah). PI3K isoforms α,...More data for this Ligand-Target Pair