Report error Found 12 Enz. Inhib. hit(s) with Target = 'Pyruvate dehydrogenase E1 component subunit beta, mitochondrial'
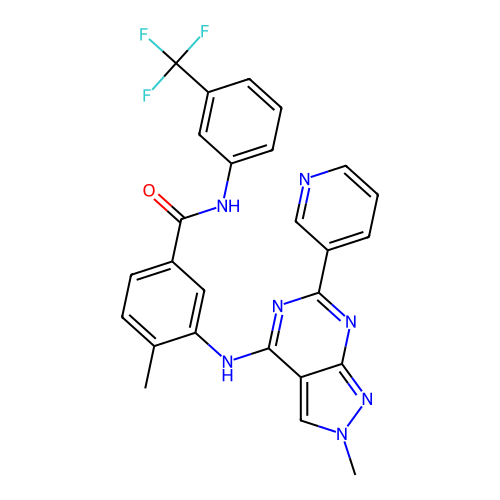
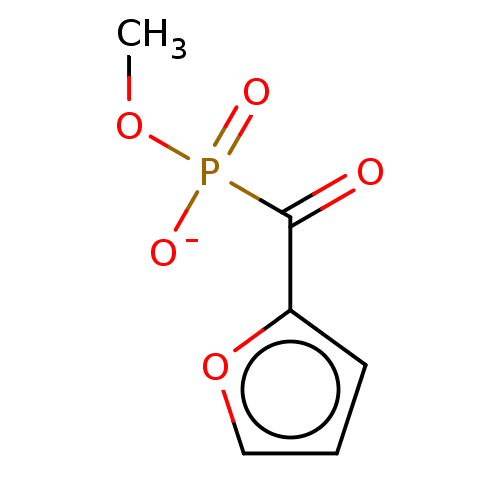
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Homo sapiens)
Johann Wolfgang Goethe University
Curated by ChEMBL
Johann Wolfgang Goethe University
Curated by ChEMBL
Affinity DataKd: 2.94E+3nMAssay Description:Binding affinity to human PDHB incubated for 45 mins by Kinobead based pull down assayMore data for this Ligand-Target Pair
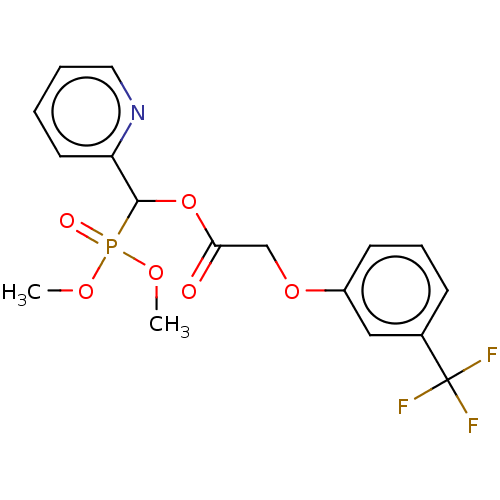
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Homo sapiens)
Johann Wolfgang Goethe University
Curated by ChEMBL
Johann Wolfgang Goethe University
Curated by ChEMBL
Affinity DataKd: 6.36E+3nMAssay Description:Binding affinity to human PDHB incubated for 45 mins by Kinobead based pull down assayMore data for this Ligand-Target Pair
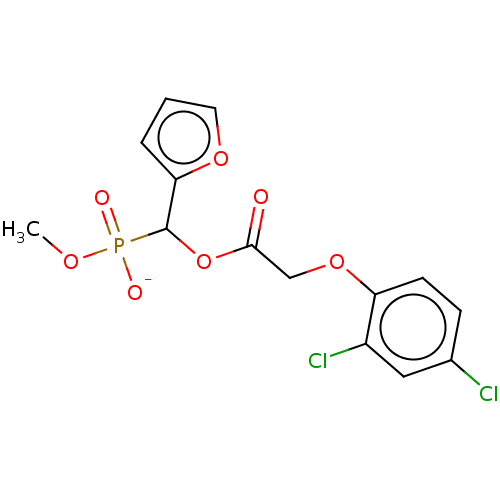
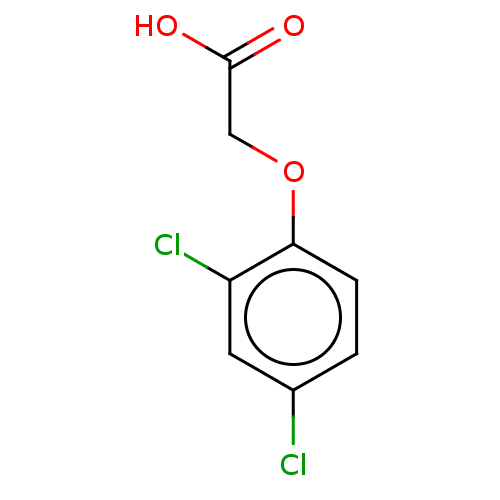
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 6.96E+3nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
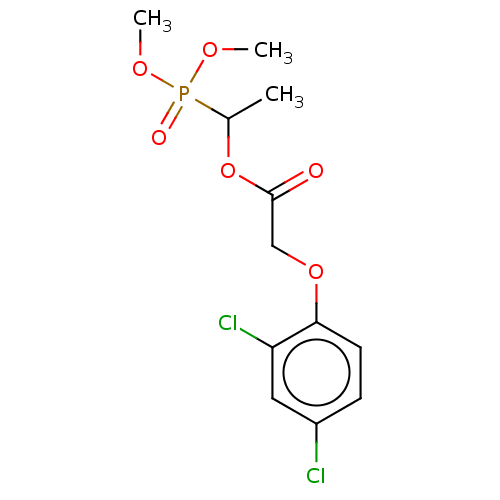
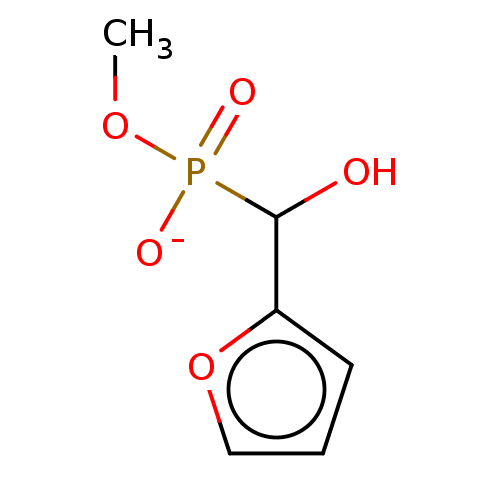
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 1.82E+4nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
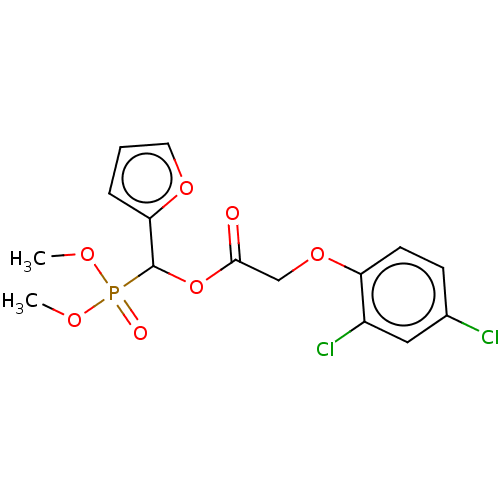
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 1.82E+4nMAssay Description:Inhibition of PDHc in Pisum sativum Zhongwan-2 (pea) shoot mitochondria assessed as decrease in rate of NADH formation using sodium pyruvate as subst...More data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 2.83E+4nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 3.89E+5nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 4.42E+5nMAssay Description:Inhibition of PDHc in Pisum sativum Zhongwan-2 (pea) shoot mitochondria assessed as decrease in rate of NADH formation using sodium pyruvate as subst...More data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 2.30E+6nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 4.53E+6nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair
TargetPyruvate dehydrogenase E1 component subunit beta, mitochondrial(Garden pea)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 4.67E+6nMAssay Description:Inhibition of Pisum sativum (pea) PDHc by spectrophotometric assayMore data for this Ligand-Target Pair