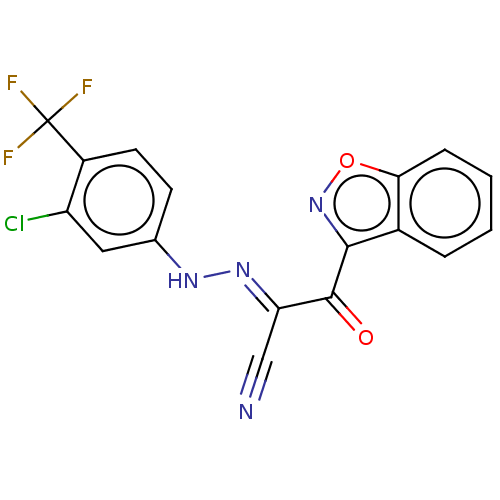
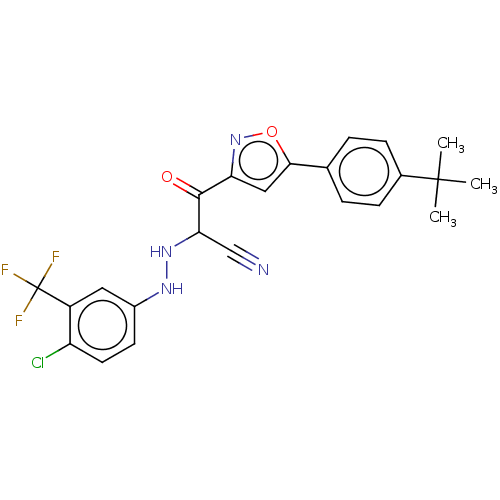
Report error Found 190 Enz. Inhib. hit(s) with Target = 'Rap guanine nucleotide exchange factor 4'
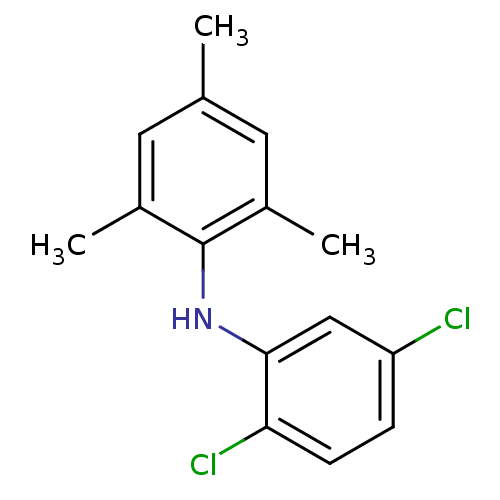
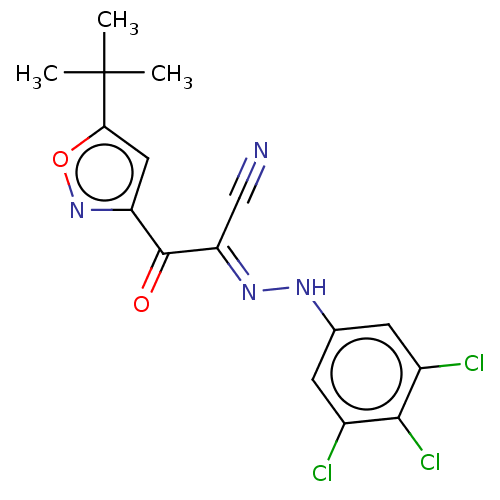
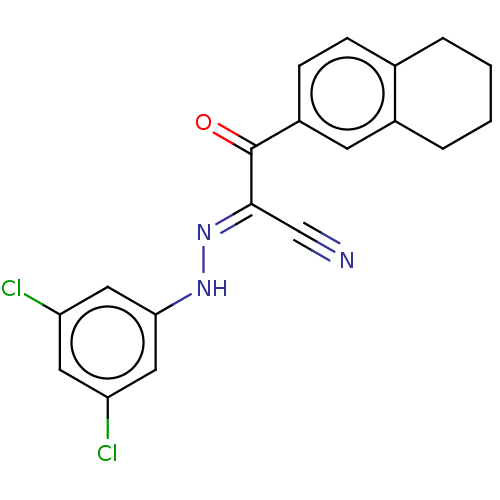
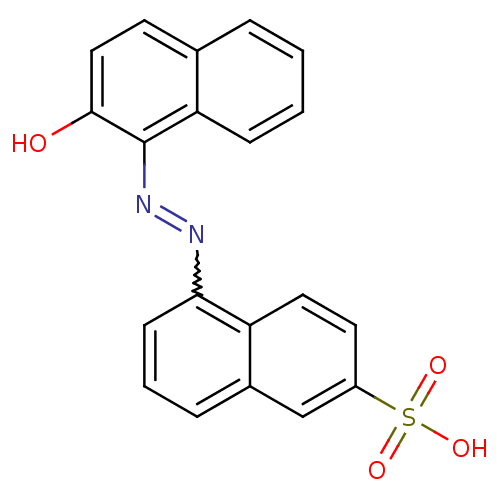
Affinity DataEC50: 300nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
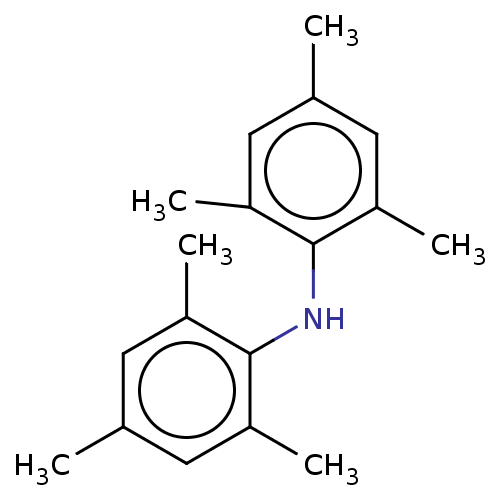
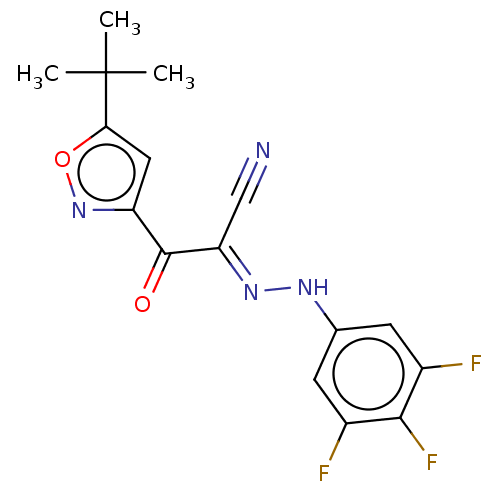
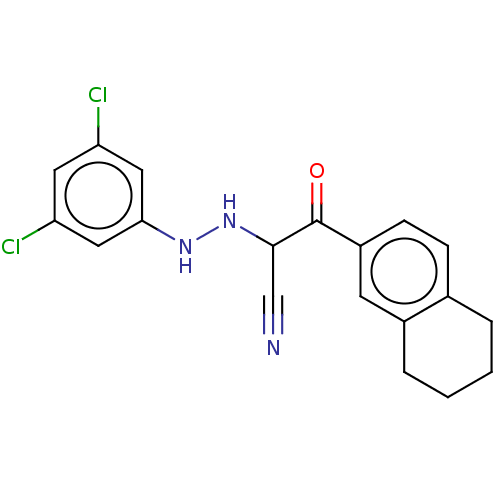
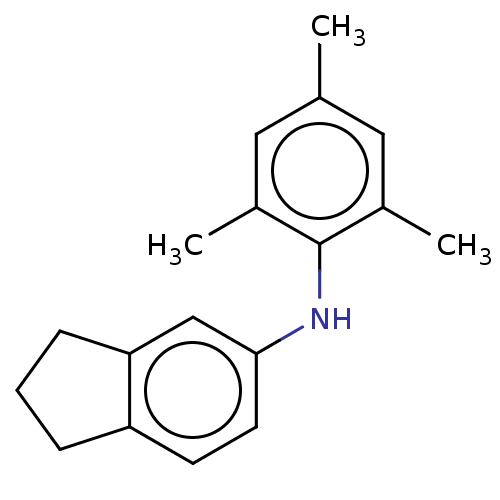
Affinity DataIC50: 300nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
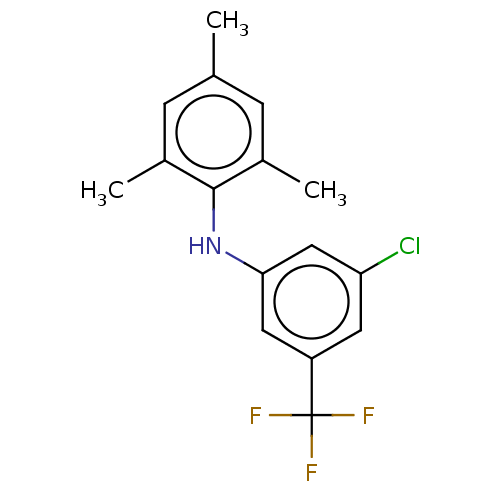
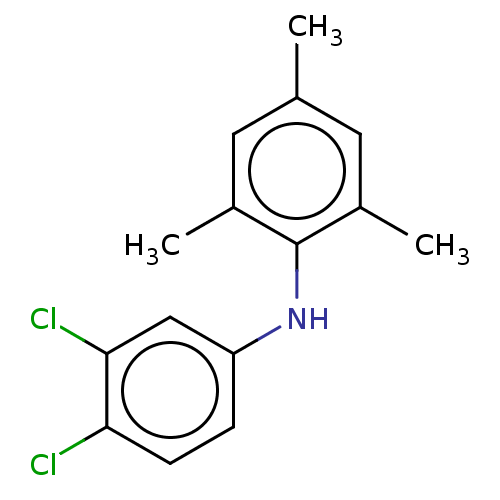
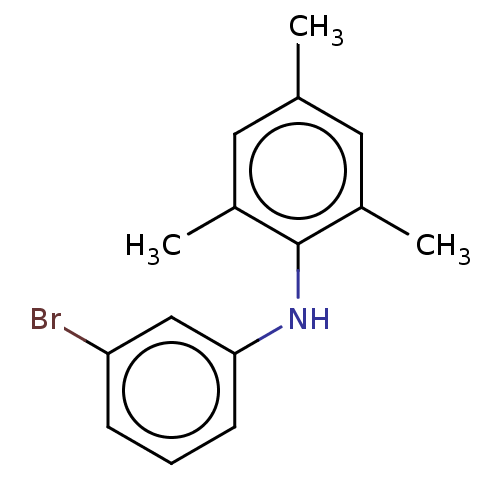
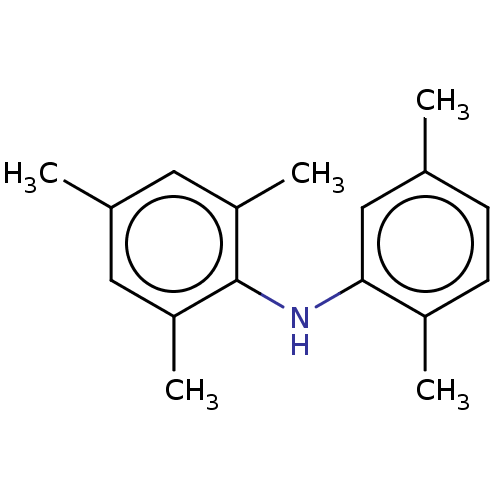
Affinity DataIC50: 400nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
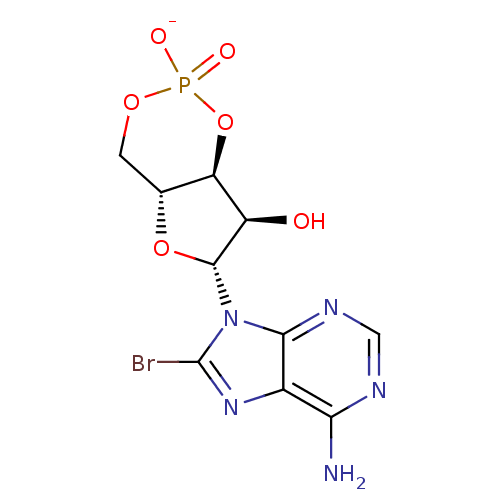
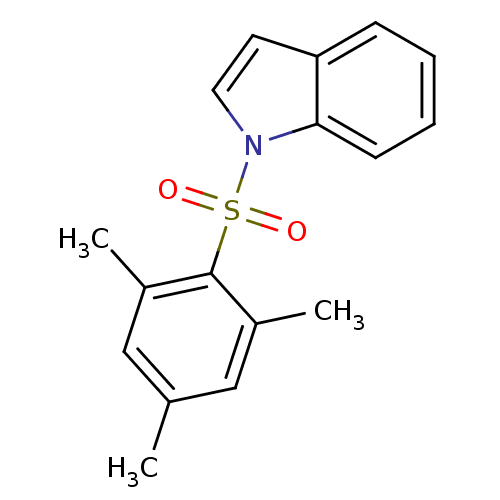
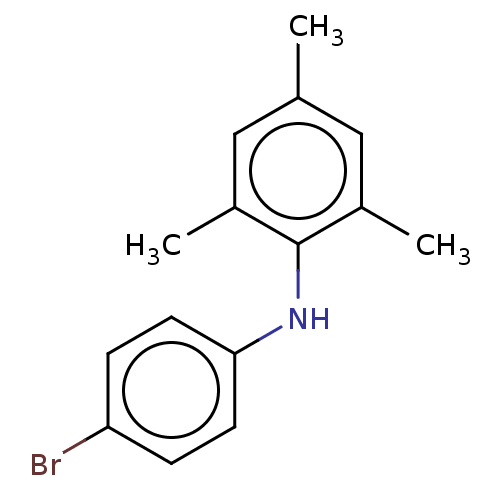
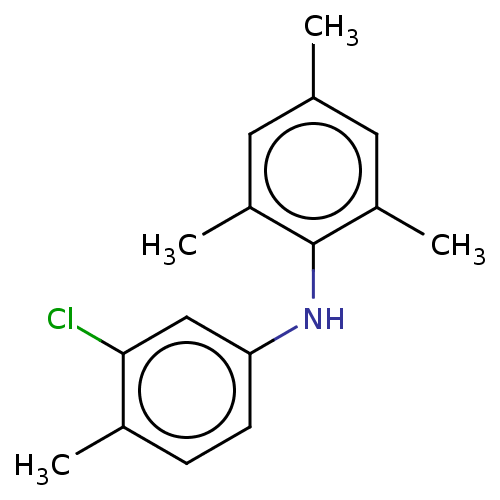
Affinity DataIC50: 400nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of EPAC2 (unknown origin)More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 600nMAssay Description:Inhibition of EPAC2 (unknown origin)More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 600nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 700nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 700nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 780nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 900nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Displacement of 8-NBD-cAMP from mouse EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of EPAC2 (unknown origin) assessed as reduction in cAMP-mediated guanine nucleotide exchange factor activity in presence of C-terminal tru...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:Antagonist activity at recombinant full length EPAC2 (unknown origin) assessed as inhibition of cAMP-mediated Rap1b-BODIPY GDP nucleotide exchange ac...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:From the biological results discussed above, compounds 10, 14-15, 23-24, 26-27, and 31-32 were identified as potent EPAC1 inhibitors with IC50 values...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of EPAC2 (unknown origin) assessed as reduction in cAMP-mediated guanine nucleotide exchange factor activity in presence of C-terminal tru...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of EPAC2 (unknown origin) assessed as reduction in cAMP-mediated guanine nucleotide exchange factor activity in presence of C-terminal tru...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Displacement of 8-NBD-cAMP from EPAC2 (unknown origin) by fluorescence plate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.90E+3nMAssay Description:From the biological results discussed above, compounds 10, 14-15, 23-24, 26-27, and 31-32 were identified as potent EPAC1 inhibitors with IC50 values...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 1.90E+3nMAssay Description:Antagonist activity at recombinant EPAC2 (unknown origin) assessed 8-NBD-cAMP substrate fluorescence intensity by substrate competing assayMore data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 2.06E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of 8-NBD-cAMP binding to EPAC2 (unknown origin) by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
TargetRap guanine nucleotide exchange factor 4(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataIC50: 2.20E+3nMAssay Description:From the biological results discussed above, compounds 10, 14-15, 23-24, 26-27, and 31-32 were identified as potent EPAC1 inhibitors with IC50 values...More data for this Ligand-Target Pair