Report error Found 61 Enz. Inhib. hit(s) with Target = 'Serpin H1'
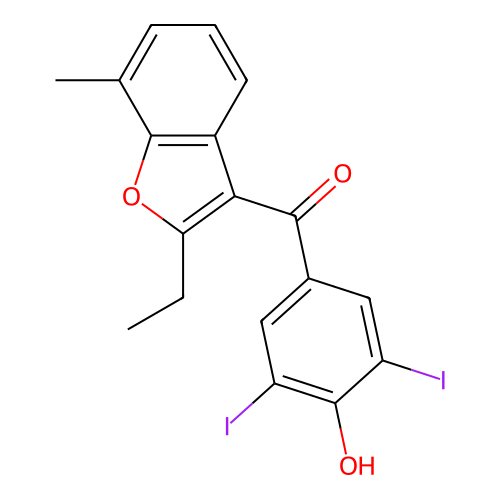
Affinity DataIC50: 580nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
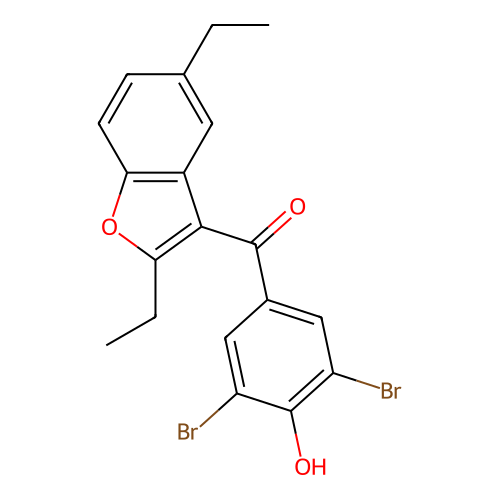
Affinity DataIC50: 640nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
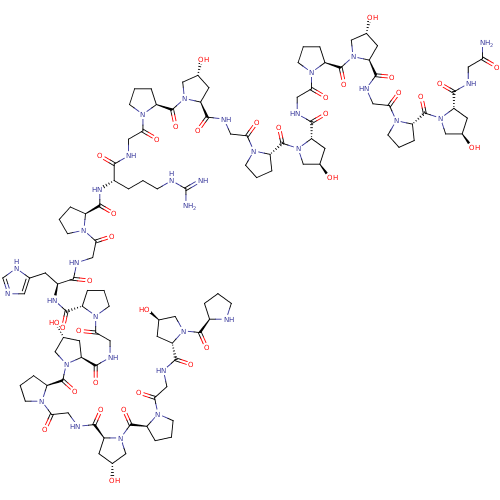
Affinity DataIC50: 690nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
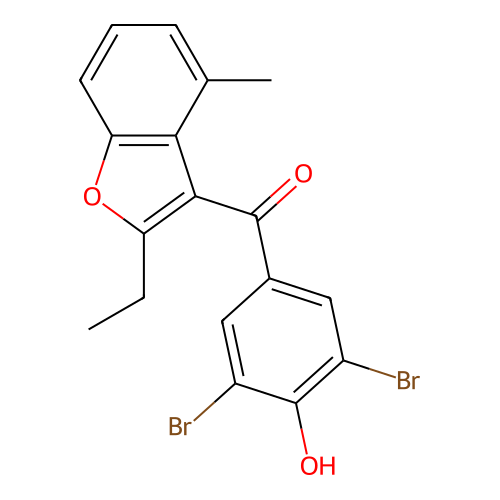
Affinity DataIC50: 940nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 940nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibitory concentration against Hsp47 collagen chaperone in turbidity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 5.52E+3nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibitory concentration against Hsp47 collagen chaperone in turbidity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.40E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.50E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.80E+3nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.59E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibitory concentration against Hsp47 collagen chaperone in turbidity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibitory concentration against Hsp47 collagen chaperone in turbidity assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3.14E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.21E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3.71E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.78E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.08E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 4.83E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.91E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.92E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 7.10E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 7.30E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataEC50: 7.31E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.90E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair
Affinity DataEC50: 8.08E+4nMAssay Description:Fibrils were formed by adding 180 ul of phosphate-buffered saline (PBS, pH 7.4) solution to 20 ul of collagen dissolved in acidic solution (collagen ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.50E+4nMAssay Description:Binding affinity to mouse recombinant HSP47 expressed in Escherichia coli JM109 after 10 mins by surface plasmon resonance assayMore data for this Ligand-Target Pair