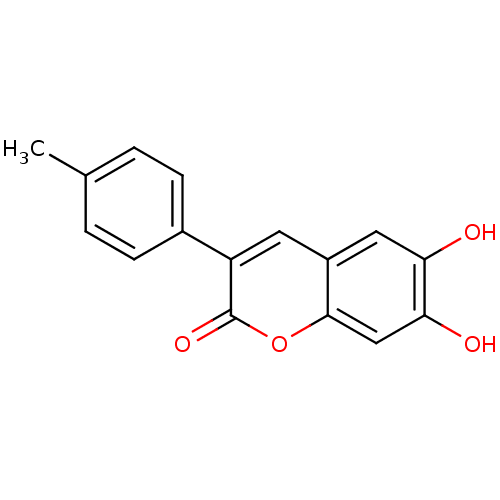
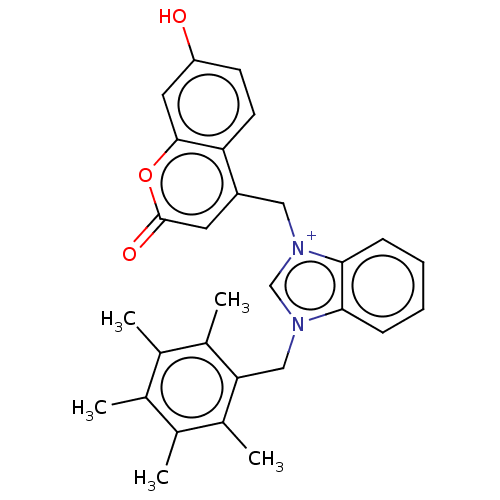
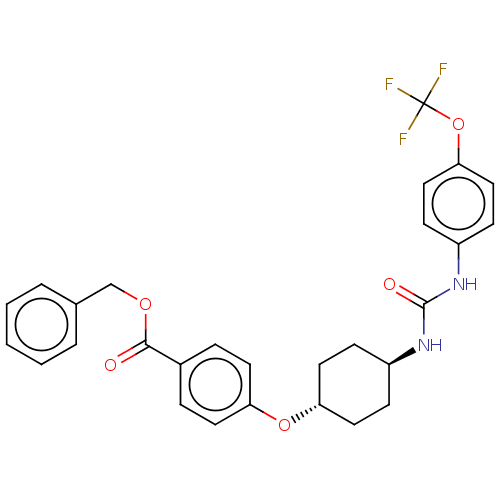
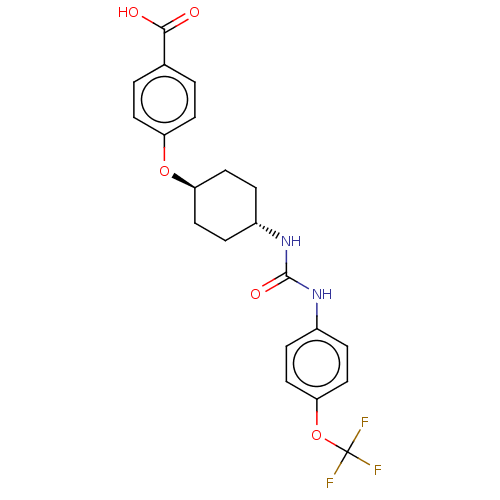
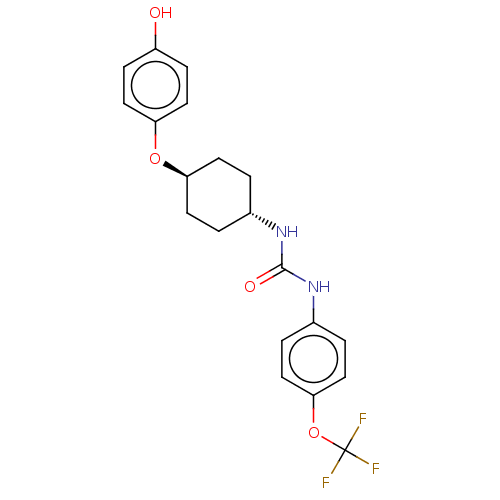
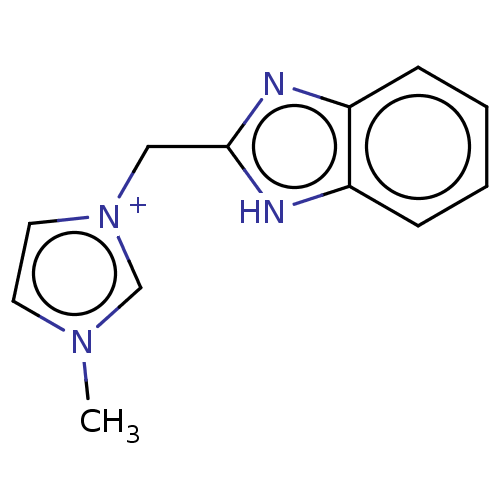
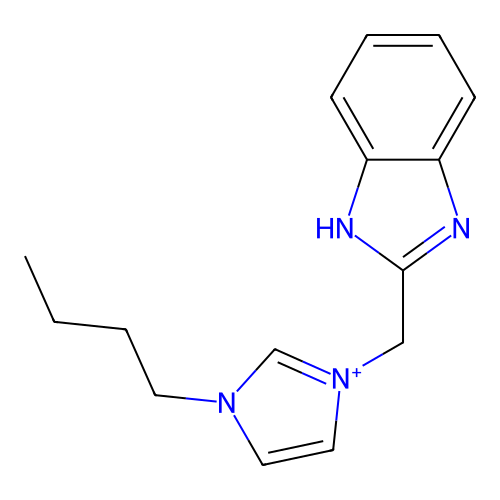
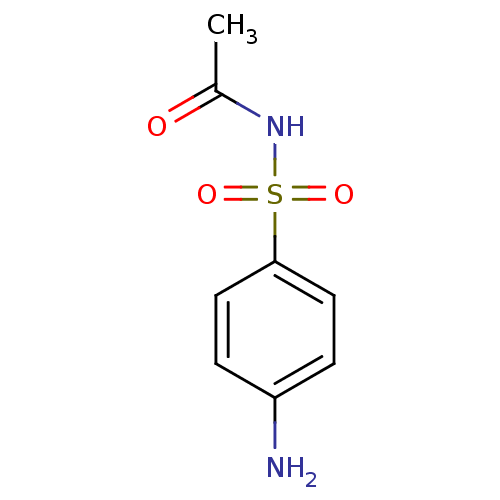
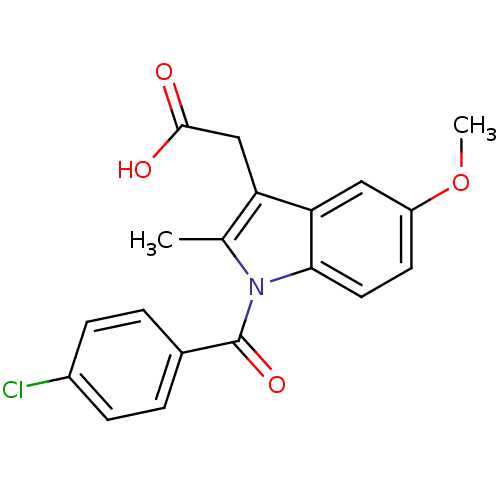
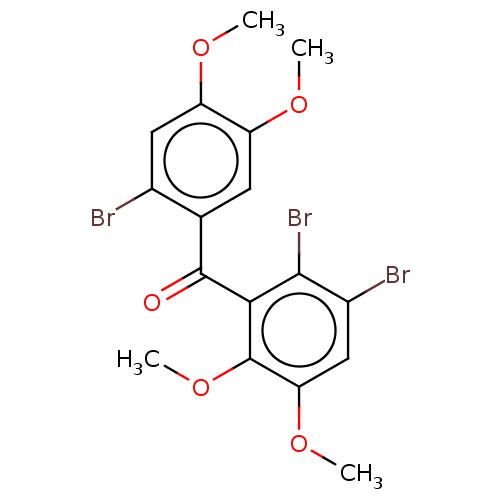
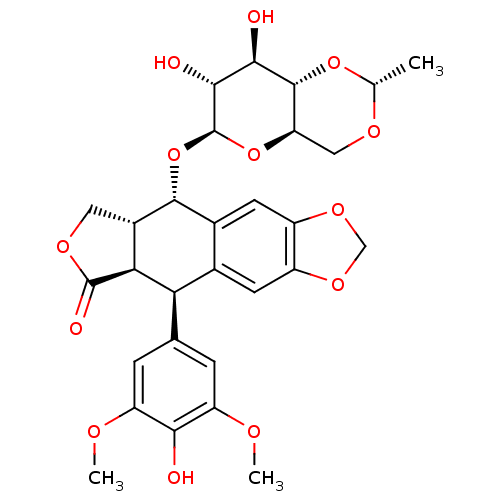
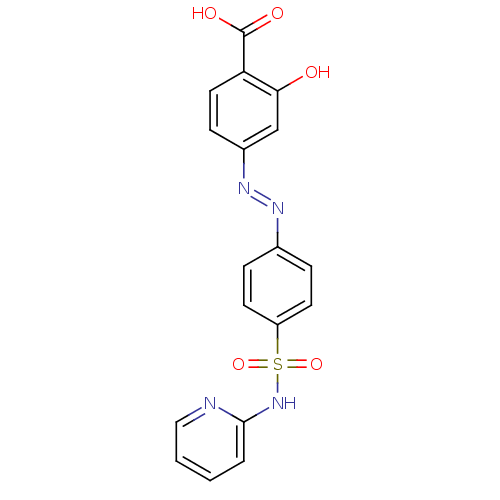
Report error Found 53 Enz. Inhib. hit(s) with Target = 'Serum paraoxonase/arylesterase 1'
Affinity DataKi: 2.39E+3nMAssay Description:Competitive inhibition of human serum PON1 using paraoxon as substrate by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6.37E+3nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 7.84E+3nMAssay Description:Inhibition of human PON1 using paraoxon as a substrate by regression analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 8.76E+3nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 9.00E+3nM ΔG°: -28.8kJ/mole IC50: 1.36E+5nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PON1 (unknown origin) expressed in baculovirus system using CMNA substrate by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PON1 (unknown origin) expressed in baculovirus system using CMNA substrate by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of PON1 (unknown origin) expressed in baculovirus system using CMNA substrate by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.65E+4nM IC50: 4.20E+4nMpH: 10.5Assay Description:Paraoxonase enzyme activity was determined at 25 °C with paraoxon (1 mM) in 50 mM glycine-NaOH (pH 10.5) containing 1 mM CaCl2. The enzyme assay was ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.91E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.37E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.14E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+4nM IC50: 1.80E+5nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+4nM ΔG°: -22.9kJ/mole IC50: 1.95E+5nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+5nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+5nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataKi: 1.31E+5nM IC50: 2.26E+5nMpH: 10.5Assay Description:Paraoxonase enzyme activity was determined at 25 °C with paraoxon (1 mM) in 50 mM glycine-NaOH (pH 10.5) containing 1 mM CaCl2. The enzyme assay was ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.74E+5nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+5nM IC50: 1.90E+5nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+5nMAssay Description:PON activities were measured in the presence of different drug concentrations. Control activity was assumed to be 100% in the absence of inhibitor. E...More data for this Ligand-Target Pair
Affinity DataIC50: 2.38E+5nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 5.70E+5nM IC50: 2.60E+5nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataKi: 2.91E+5nM IC50: 6.65E+5nMpH: 10.5Assay Description:Paraoxonase enzyme activity was determined at 25 °C with paraoxon (1 mM) in 50 mM glycine-NaOH (pH 10.5) containing 1 mM CaCl2. The enzyme assay was ...More data for this Ligand-Target Pair
Affinity DataKi: 3.06E+5nM ΔG°: -20.1kJ/mole IC50: 3.40E+5nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataIC50: 3.17E+5nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.02E+5nM IC50: 3.60E+5nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.64E+5nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6.26E+5nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6.27E+5nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 7.01E+5nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataKi: 7.43E+5nM IC50: 8.00E+5nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataKi: 8.05E+5nM ΔG°: -17.7kJ/mole IC50: 1.64E+6nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataIC50: 8.32E+5nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+6nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+6nMAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412 nm. Paraoxonase activity was determined spectrophotometrically using the same pr...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+6nM IC50: 1.24E+6nMAssay Description:The inhibitory effects of six sulfonamide derivatives was examined against the purified enzyme hPON1 and were tested in a triplicate experiment in fi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+6nMAssay Description:Inhibitory activity against human serum paraoxonase (PON1)More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+6nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.25E+6nMAssay Description:Inhibitory activity against human serum paraoxonase (PON1)More data for this Ligand-Target Pair
Affinity DataIC50: 3.27E+6nMAssay Description:Inhibitory activity against human serum paraoxonase (PON1)More data for this Ligand-Target Pair
Affinity DataIC50: 3.33E+6nMAssay Description:Inhibition of human serum PON1 using paraoxon as substrate by spectrophotometric analysisMore data for this Ligand-Target Pair
Affinity DataKi: 3.73E+6nMAssay Description:PON activities were measured in the presence of different drug concentrations. Control activity was assumed to be 100% in the absence of inhibitor. E...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+7nM ΔG°: -10.8kJ/mole IC50: 6.23E+6nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataKi: 8.97E+6nM IC50: 2.33E+7nMpH: 10.5Assay Description:Paraoxonase enzyme activity was determined at 25 °C with paraoxon (1 mM) in 50 mM glycine-NaOH (pH 10.5) containing 1 mM CaCl2. The enzyme assay was ...More data for this Ligand-Target Pair
Affinity DataKi: 1.11E+7nM ΔG°: -11.2kJ/mole IC50: 1.30E+7nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataKi: 1.83E+7nMAssay Description:PON activities were measured in the presence of different drug concentrations. Control activity was assumed to be 100% in the absence of inhibitor. E...More data for this Ligand-Target Pair