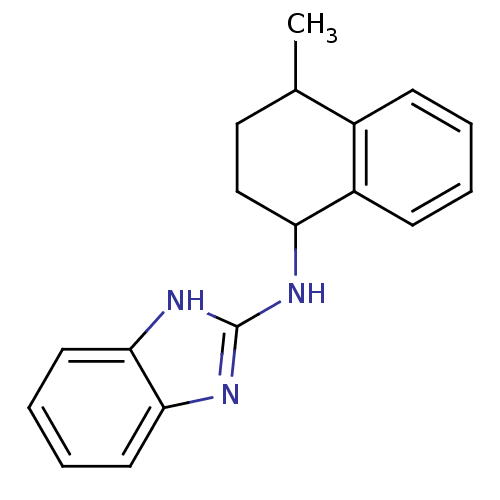
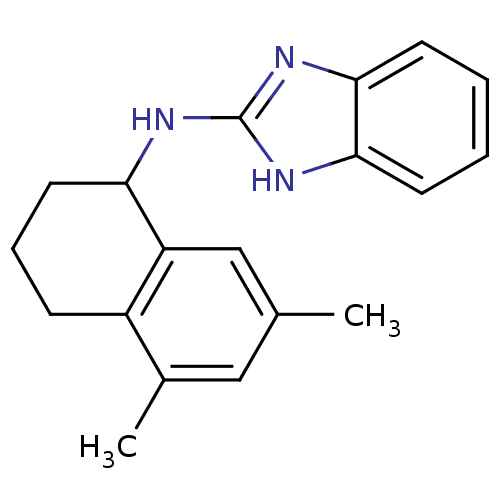
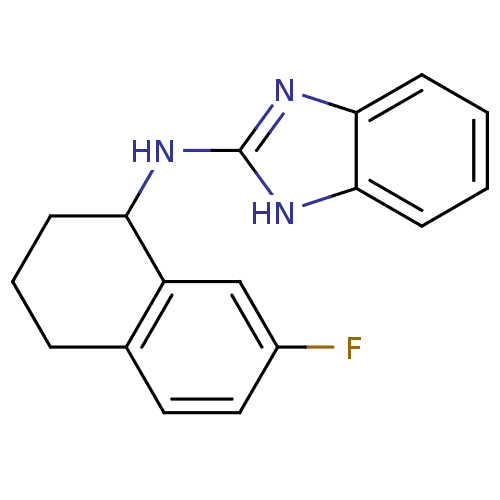
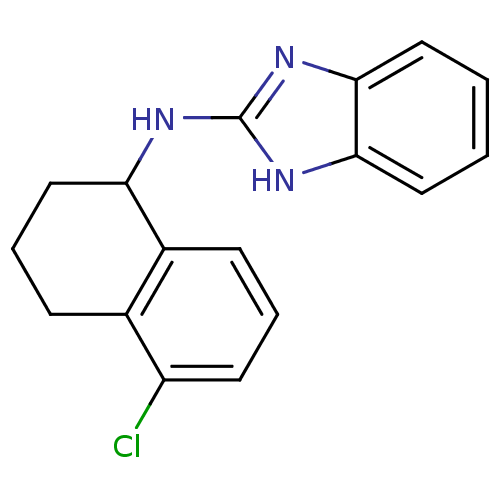
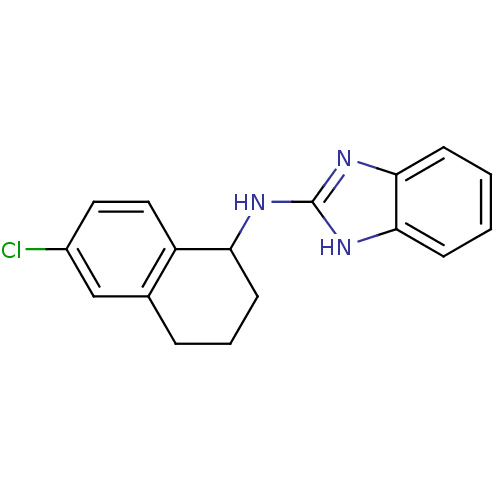
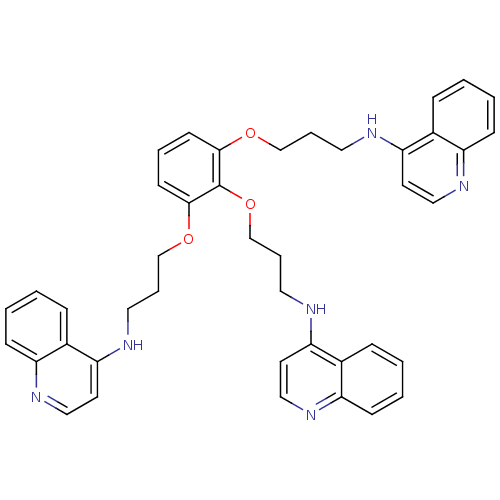
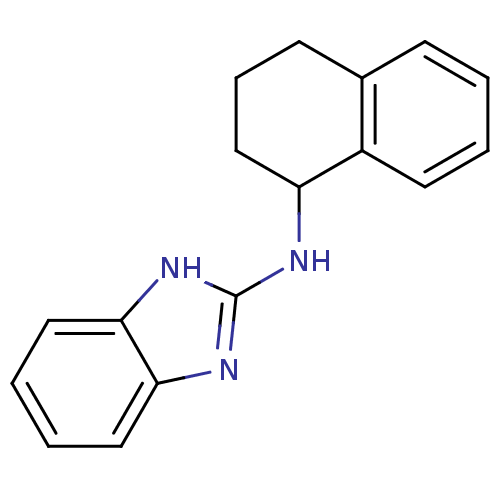
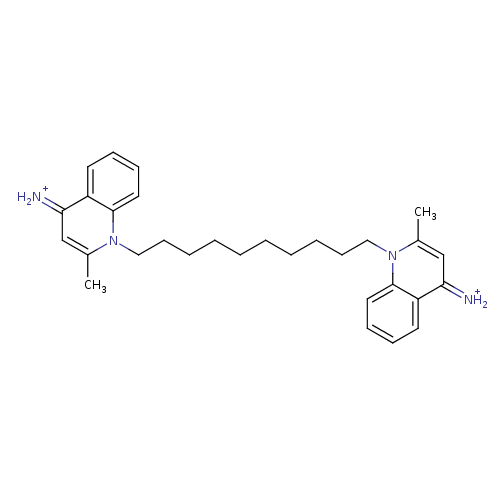
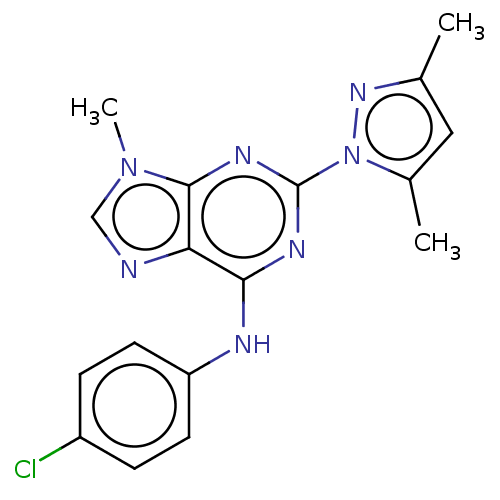
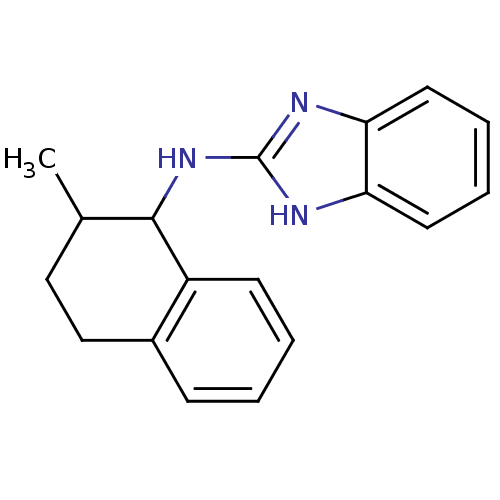
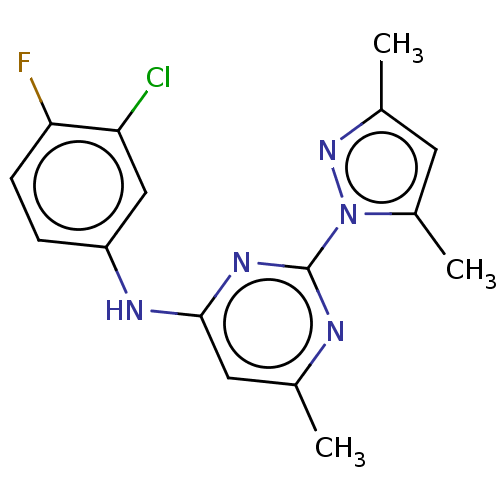
Report error Found 148 Enz. Inhib. hit(s) with Target = 'Small conductance calcium-activated potassium channel protein 3'
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.0640nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.0640nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.168nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.168nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 0.360nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as reduction in channel current with holding potential 0 mV and ramp voltage ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 0.380nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as reduction in channel current with holding potential 0 mV and ramp voltage ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 0.430nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 1.10nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as decrease in intracellular calcium level at membrane potential -120 mV for ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 1.10nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as decrease in intracellular calcium level at membrane potential -120 mV for ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Displacement of [I125]apamine from Wistar rat recombinant SK3 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 1.70nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as decrease in intracellular calcium level at membrane potential -120 mV for ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKi: 1.80nMAssay Description:Displacement of [I125]apamine from Wistar rat recombinant SK3 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 2.60nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as reduction in channel current with holding potential 0 mV and ramp voltage ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 2.70nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 3.80nMAssay Description:Inhibition of recombinant human SK3 expressed in HEK293T cells assessed as decrease in intracellular calcium level at membrane potential -120 mV for ...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Displacement of [125I]apamin from Kca2.3 channel expressed in HEK293 cells by scintillation proximity assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataKd: 7.80nMAssay Description:Inhibition of human SK3 channel expressed in HEK293 cells assessed as Ca2+ induced current after drug wash out by whole cell patch clamp technique in...More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 9.10nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 11nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 17nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 20nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 20nMAssay Description:Inhibition of human Kca 2.3 expressed in HEK293 cells by patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Displacement of [125I]apamin from Kca2.3 channel expressed in HEK293 cells by scintillation proximity assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 27nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 31nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 31nMAssay Description:Inhibition of KCa2.3 (unknown origin)More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 32nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 33nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 34nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 44nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataKi: 46nMAssay Description:Displacement of [125I]-apamin from cloned SK3 channel expressed in human HEK293 cells after 1 hr by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 56nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by electrophysiology assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 59nMAssay Description:Inhibition of Kca2.3 channel expressed in HEK293 cells by thallium flux assayMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataKd: 60nMAssay Description:Inhibition of human SK3 channel by inside-out patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 60nMAssay Description:Inhibition of KCa2.3 (unknown origin)More data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 61nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 77nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataIC50: 86nMAssay Description:Inhibition of SKca channel-mediated after-hypepolarization in rat sympathetic neuronsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataIC50: 93nMAssay Description:Inhibition of SKca channel-mediated after-hypepolarization in rat sympathetic neuronsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataKd: 100nMAssay Description:Inhibition of wild type human SK3 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 120nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of SKca channel-mediated after-hypepolarization in rat sympathetic neuronsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of SKca channel-mediated after-hypepolarization in rat sympathetic neuronsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 130nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 130nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKi: 140nMAssay Description:Displacement of [I125]apamine from Wistar rat recombinant SK3 channel expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 140nMAssay Description:Positive modulation of SK3 in HEK293 cells in presence of Ca2+ by patch-clamp methodMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 170nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Rat)
Neurosearch
Curated by ChEMBL
Neurosearch
Curated by ChEMBL
Affinity DataKd: 180nMAssay Description:Inhibition of Wistar rat recombinant SK3 channel expressed in HEK293 cells by whole cell patch clamp techniqueMore data for this Ligand-Target Pair
TargetSmall conductance calcium-activated potassium channel protein 3(Human)
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataEC50: 190nMAssay Description:Potentiation of human KCa2.3 expressed in HEK293 cells at -90 mV holding potential measured at 1 to 2 days by patch clamp electrophysiology relative ...More data for this Ligand-Target Pair