Report error Found 11043 Enz. Inhib. hit(s) with Target = 'Sodium-dependent dopamine transporter'
Affinity DataIC50: 0.0300nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]WIN 35428 from dopamine transporter in rat striatum after 120 mins by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0430nMAssay Description:Inhibition of [125I]RTI-55 binding to dopamine transport sites in rat striatal membranes.More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:In vitro binding affinity towards dopamine transporter in rat striatal membranes by [3H]GBR-12395 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 0.0770nMAssay Description:Tested for the ability to displace [125I]- RTI-55 binding in rat striatal membrane against dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0850nMAssay Description:Tested for the ability to displace [125I]- RTI-55 binding in rat striatal membrane against dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.115nMAssay Description:Inhibition of [125I]RTI-55 binding to dopamine transport sites in rat striatal membranes.More data for this Ligand-Target Pair
Affinity DataIC50: 0.120nMAssay Description:Displacement of [125I]RTI-55 from dopamine transporter of rat brain membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Displacement of [3H]WIN 35428 from dopamine transporter in rat striatum after 120 mins by liquid scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Binding affinity towards dopamine transporter using [3H]-mazindol as radioligand in rat striatal membranes.More data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.146nMAssay Description:The compound was tested in vitro for high binding affinity for the Dopamine transporter (DAT) using competitive binding assay.More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMAssay Description:Inhibition of dopamine uptake in rat synaptosomal fractionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMAssay Description:Inhibition of dopamine uptake in rat synaptosomal fractionMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]CFT from human DAT expressed in COS7 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataKi: 0.220nMAssay Description:Binding constant for the ability to displace [3H]mazindol to dopamine receptor was calculated from the Cheng-Prusoff relationshipMore data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:Binding affinity towards dopamine transporter using [3H]-mazindol as radioligand in rat striatal membranes.More data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.227nMAssay Description:The compound was tested in vitro for high binding affinity for the Dopamine transporter (DAT) using competitive binding assay.More data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.230nMAssay Description:Displacement of [3H]WIN-35428 from dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.260nMAssay Description:Inhibition of [3H]dopamine uptake at the dopamine transporter in rat striatal tissueMore data for this Ligand-Target Pair
Affinity DataIC50: 0.280nMAssay Description:Inhibition of [125I]RTI-55 binding to dopamine transport sites in rat striatal membranes.More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]CIT from dopamine transporter of rat striatal homogenateMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Displacement of [3H]WIN-35428 from DAT in rhesus monkey caudate-putamenMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Tested for radioligand [3H]BTCP displacement from dopamine transporter in rat forebrainMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of [3H]WIN-35428 binding to Dopamine transporter in rhesus (Macaca mulatta) or cynomolgus monkey (Macaca fascicularis) caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:Binding affinity towards dopamine transporter using [3H]-mazindol as radioligand in rat striatal membranes.More data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:Inhibition of [3H]dopamine binding to rat striatal synaptosome dopamine transporterMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.320nMAssay Description:Displacement of [3H]WIN-35428 from dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.350nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataKi: 0.360nMAssay Description:Competitive binding versus [N-methyl-3H]WIN-35428 in murine kidney cells transfected with human dopamine transporterMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.360nMAssay Description:Displacement of [3H]mazindol from dopamine transporterMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.360nMAssay Description:Displacement of [3H]mazindol from dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.370nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.370nMAssay Description:Inhibition of [3H]WIN-35428 binding at dopamine transporter was determinedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of [3H]WIN-35428 binding to dopamine transporter (DAT) of cynomolgus monkey caudate-putamenMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of [3H]WIN-35428 binding to dopamine transporter (DAT) of cynomolgus monkey caudate-putamenMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Tested for the ability to displace [125I]- RTI-55 binding in rat striatal membrane against dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMAssay Description:Inhibitory activity in displacing [125I]- RTI-55 binding to dopamine transporter in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.390nMAssay Description:Tested in vitro for serotonin(5-HT) neuronal uptake inhibitionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Displacement of [3H]WIN-35428 from DAT in rhesus monkey caudate-putamenMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Displacement of [125I]RTI-55 from DATMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Displacement of [125I]RTI55 from human DATMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Institut F£R Bioanorganische Und Radiopharmazeutische Chemie
Curated by ChEMBL
Affinity DataIC50: 0.400nMAssay Description:inhibition of [3H]WIN-35428 binding to the dopamine transporter (DAT) in monkey caudate-putamen.More data for this Ligand-Target Pair
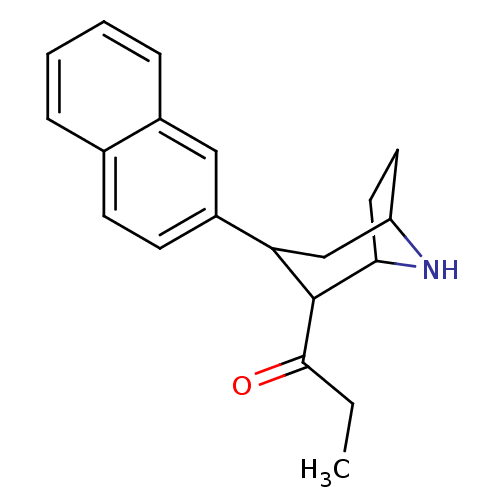

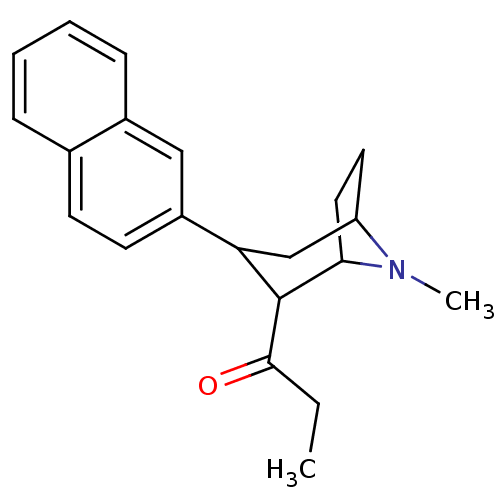
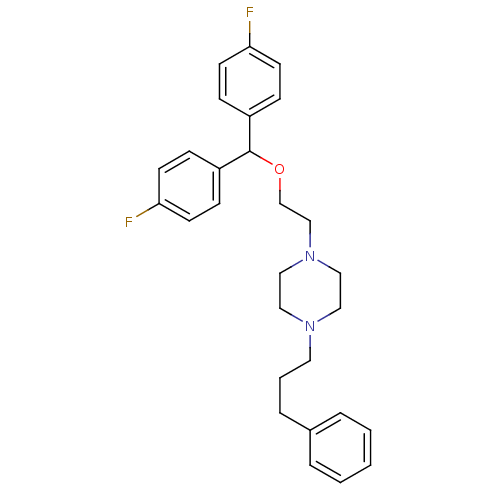
 3D Structure (crystal)
3D Structure (crystal)