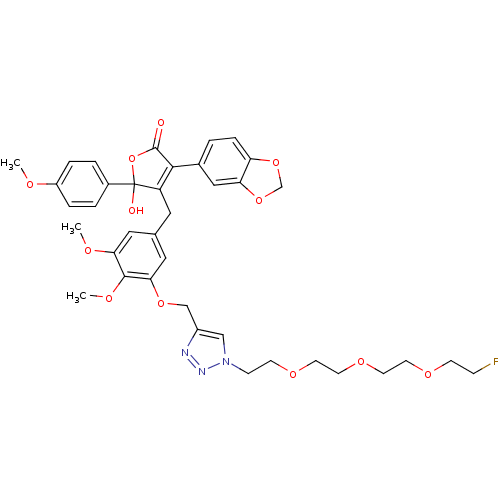
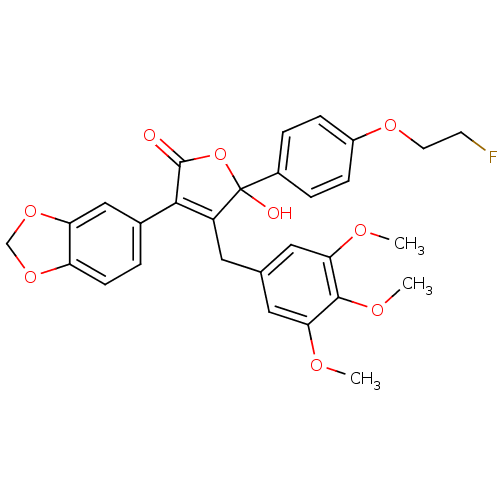
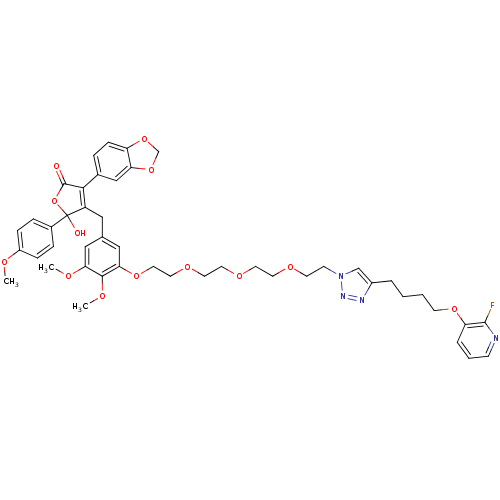
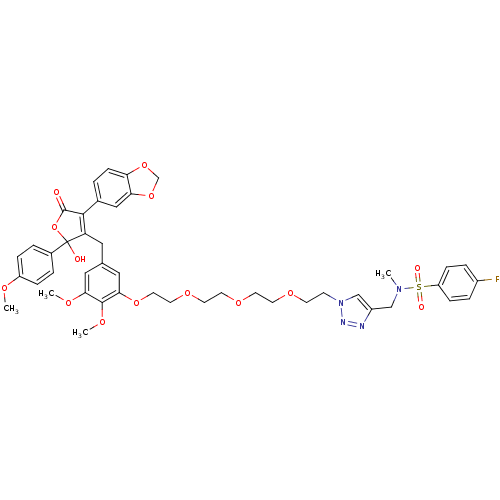
Affinity DataKi: 0.240nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 0.360nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 1.09nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.26nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 1.26nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.43nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 3.5nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.40nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 7.30nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [125I]ET1 from ETA receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Inhibition of MMP-2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Inhibition of MMP2 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of MMP-8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of MMP8 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [125I]ET1 from endothelin A receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Inhibition of MMP-9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Inhibition of MMP9 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 96nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 98nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Displacement of [125I]ET1 from endothelin B receptor in DBA mouse microsomesMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 288nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 418nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 893nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.34E+3nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 6.45E+3nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.45E+3nMAssay Description:Displacement of [125I]ET1 from ETB receptor in CD1 Nu/Nu mouse myocardial ventricles membranes after 2 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.00600nMAssay Description:Inhibition of MMP13 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incub...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of MMP13 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incub...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of MMP8 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of MMP8 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of MMP9 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of MMP13 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incub...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of MMP-13 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of MMP-13 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of MMP8 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of MMP-8 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 a...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of MMP-13 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of MMP9 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of MMP9 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of MMP-9 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 a...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of MMP-9 (unknown origin) using 7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diamino-propionyl)Ala-Arg-NH2 a...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of MMP2 using (7-methoxycoumarin-4-yl)acetyl-Pro-Leu-Gly-Leu-(3-(2,4-dinitrophenyl)-L-2,3-diaminopropionyl)Ala-Arg-NH2 as substrate incuba...More data for this Ligand-Target Pair