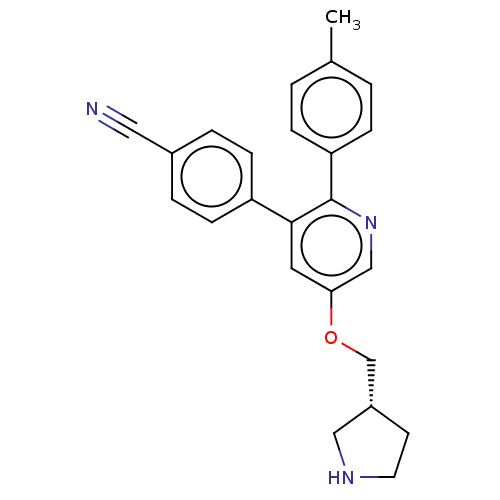
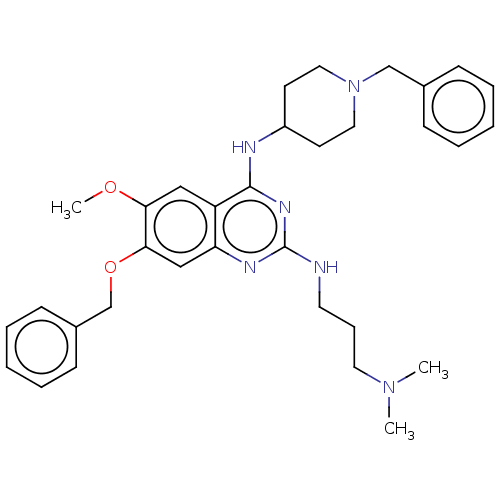
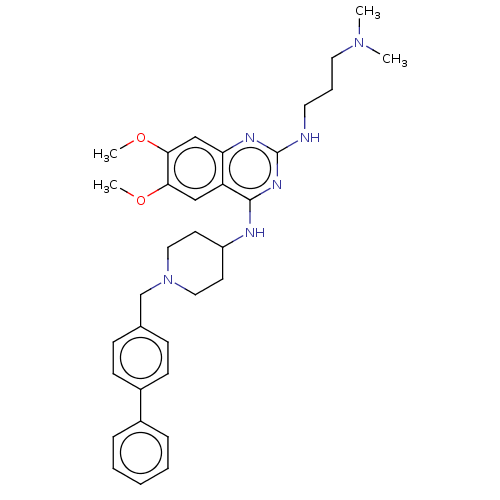
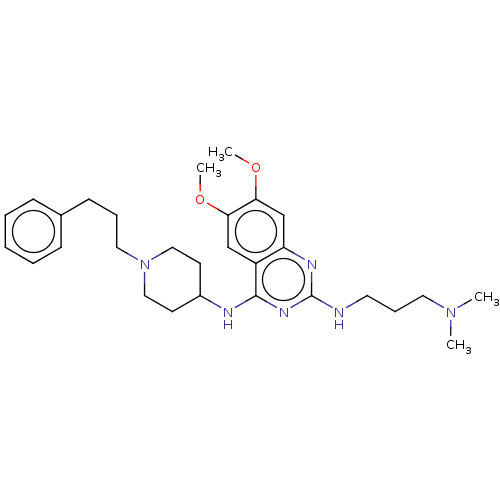
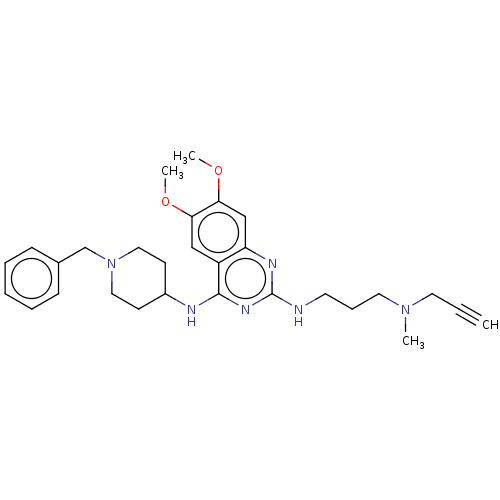
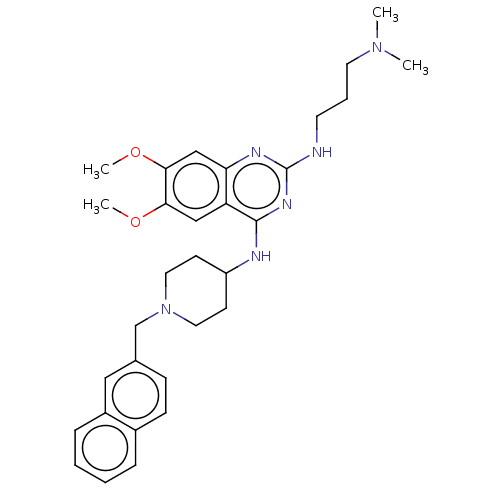
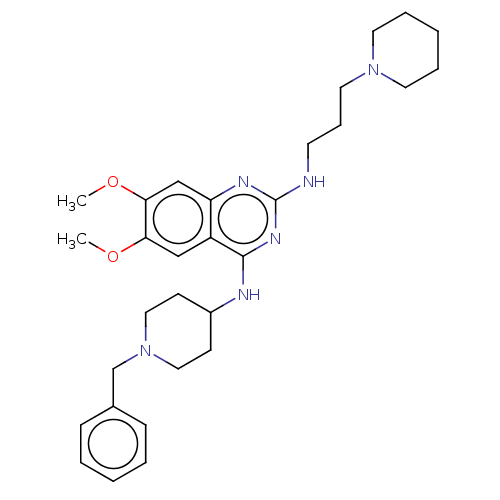
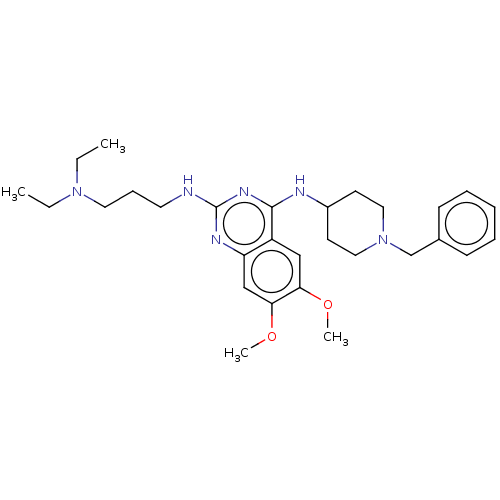
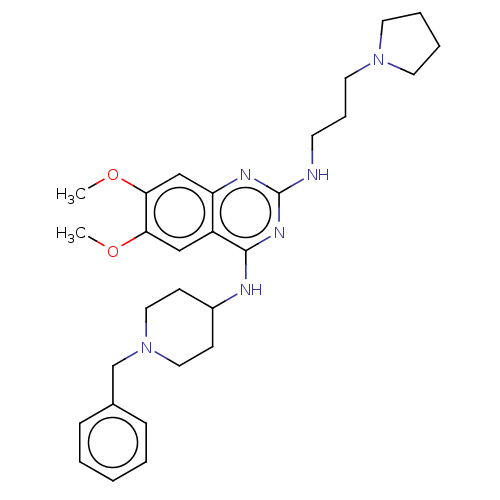
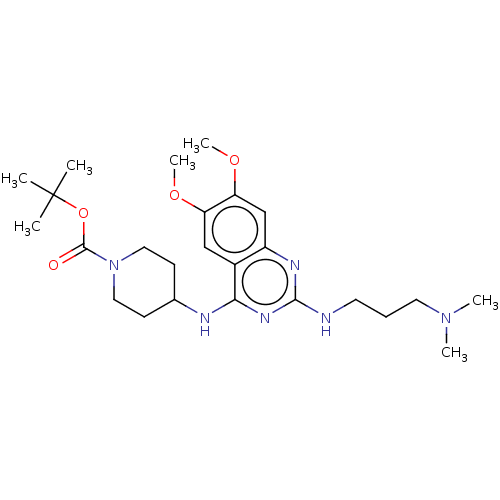
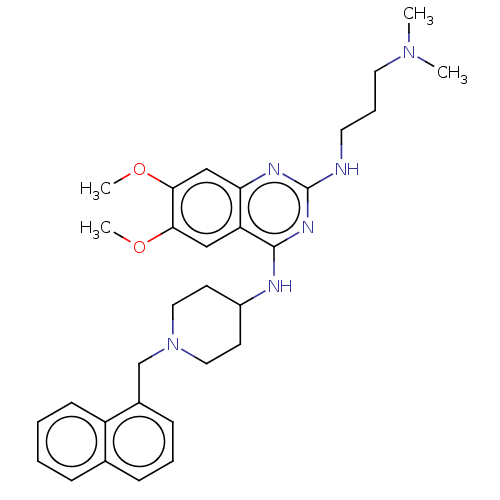
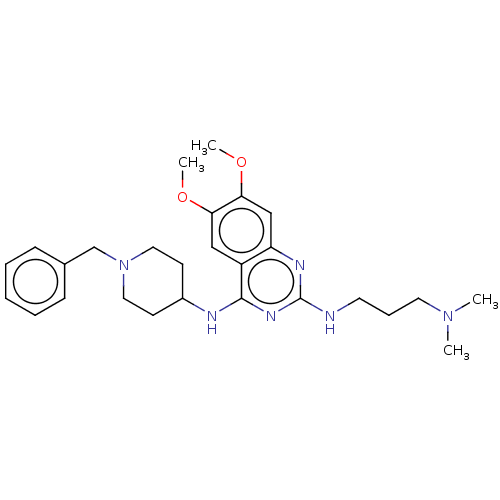
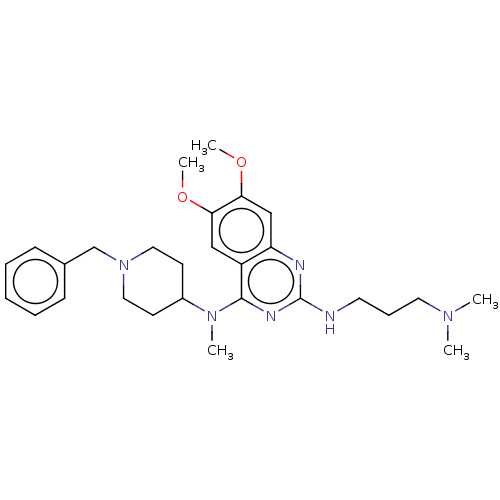
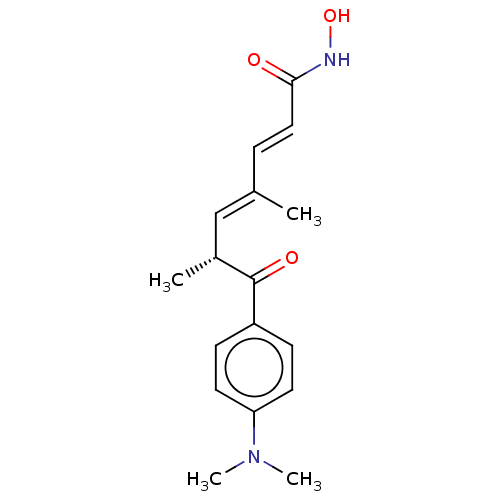
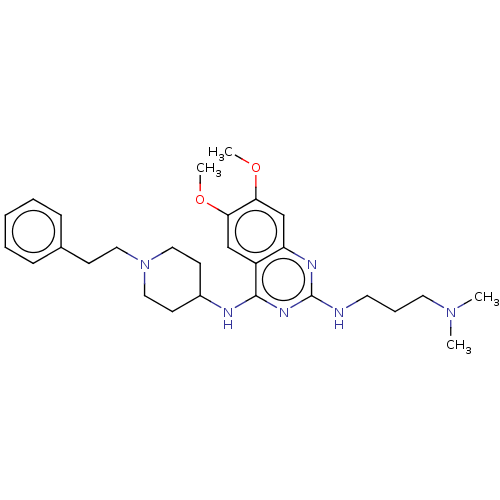
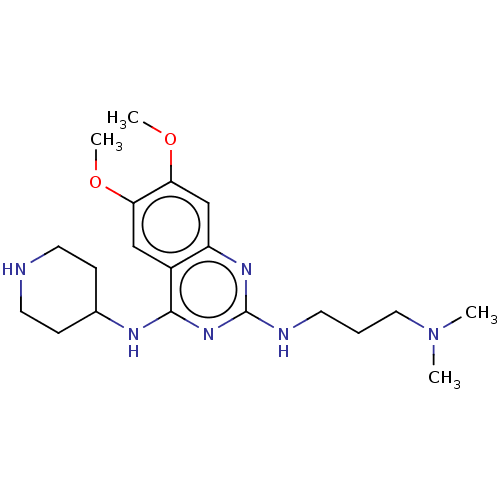
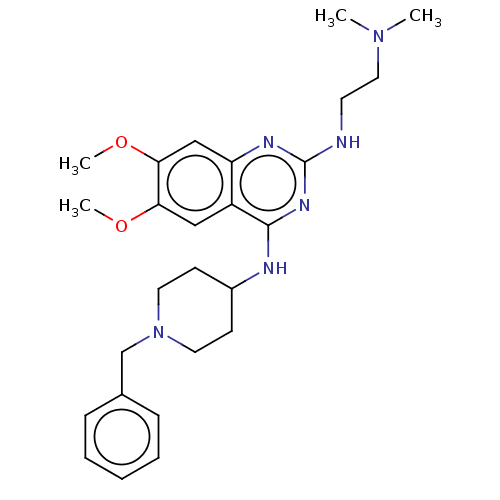
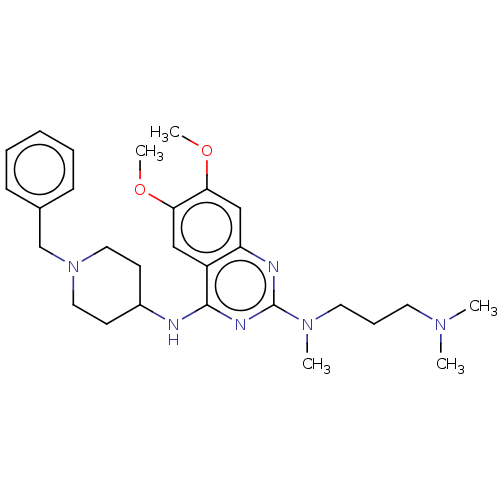
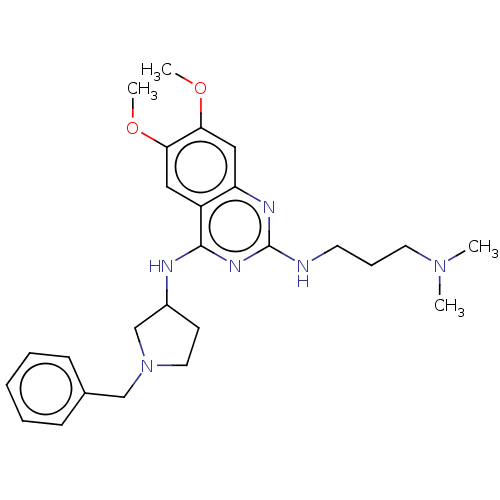
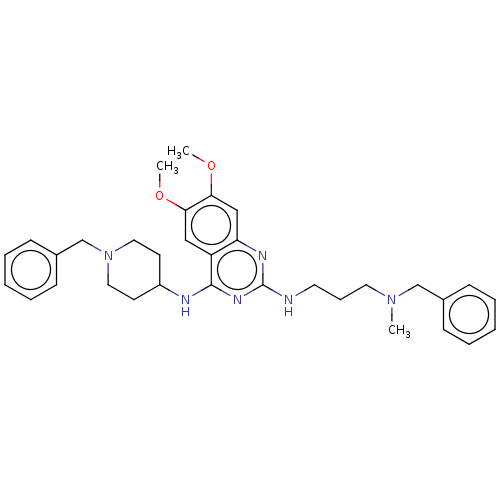
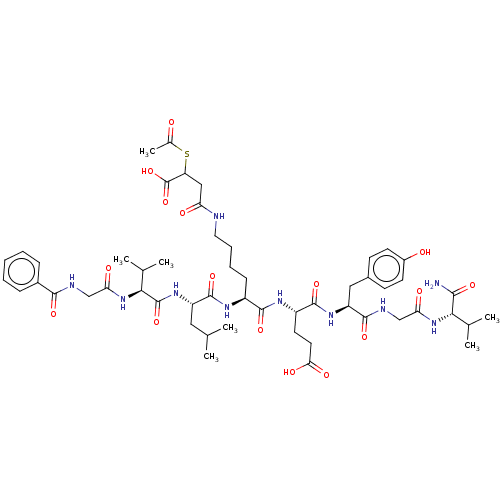
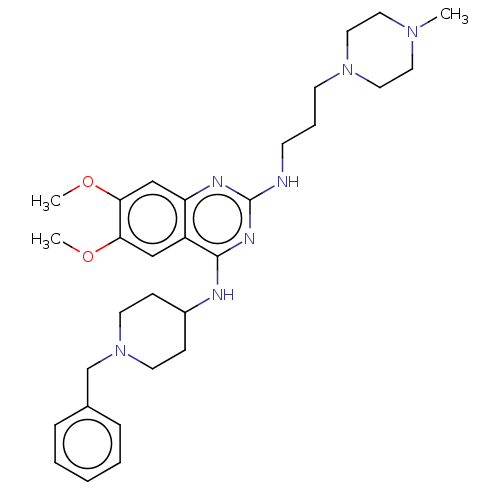
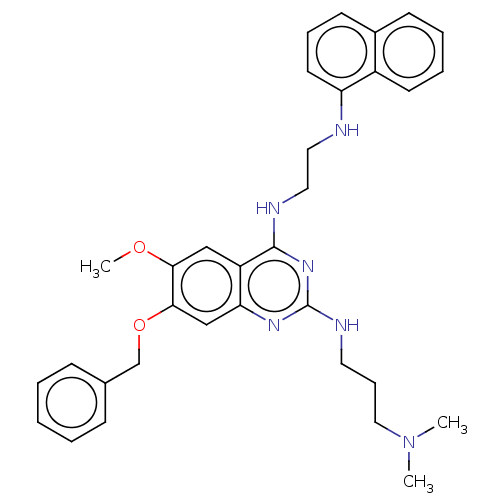
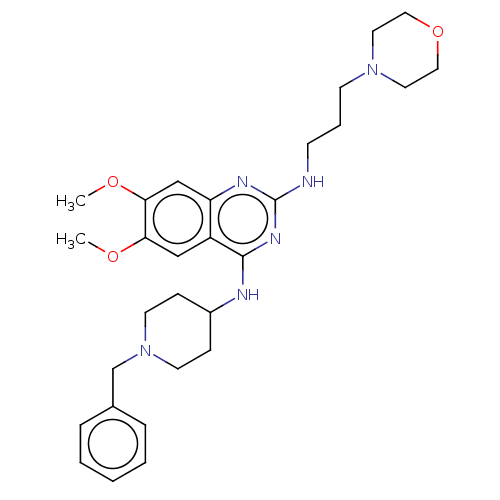
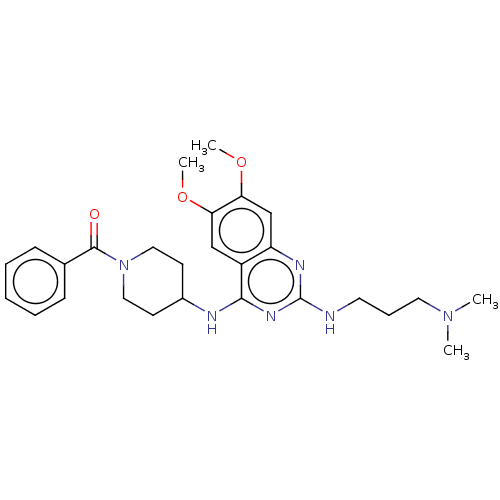
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 0.5nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as deglutarylase activity by measuring inhibition constantMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 6nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as deglutarylase activity by measuring inhibition constantMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 7nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as deglutarylase activity by measuring inhibition constantMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 22nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as deglutarylase activity by measuring inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 37nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as deglutarylase activity by measuring inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 146nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 149nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 153nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 175nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 183nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 212nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 440nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 569nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 680nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 690nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 1.18E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 2.02E+3nMAssay Description:Antagonist activity at adrenergic Alpha-1D receptor in rat aortaMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.79E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.93E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.10E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.13E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.26E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.94E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 4.01E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 4.17E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 4.24E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 4.30E+3nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as desuccinylation activity by measuring inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 4.62E+3nMAssay Description:Antagonist activity at adrenergic Alpha-1D receptor in rat aortaMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 4.83E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 5.26E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 5.82E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 6.33E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 8.03E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 9.01E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 1.06E+4nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as desuccinylation activity by measuring inhibition constantMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 1.33E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 1.34E+4nMAssay Description:Antagonist activity at adrenergic alpha1A receptor in rat tail arteryMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 1.72E+4nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as desuccinylation activity by measuring inhibition constantMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.02E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.26E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.28E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 2.61E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.29E+4nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-5, mitochondrial(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 3.81E+4nMAssay Description:Binding affinity to SIRT5 (unknown origin) assessed as desuccinylation activity by measuring inhibition constantMore data for this Ligand-Target Pair


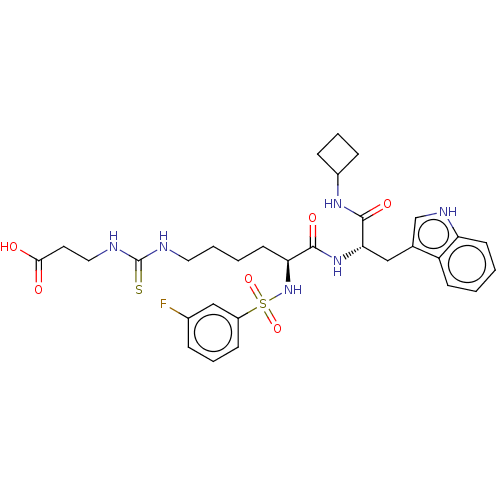
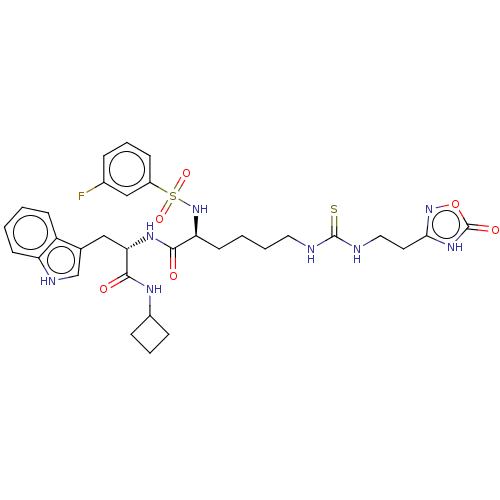
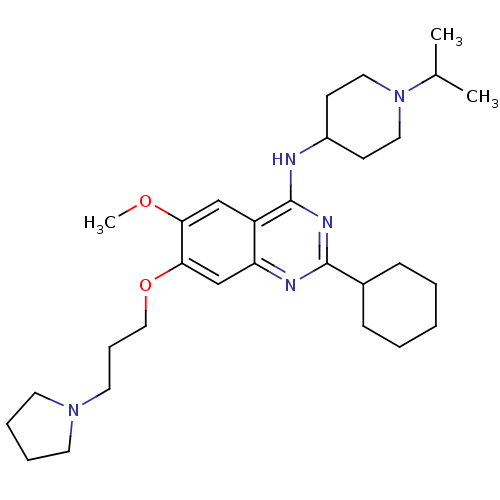
 3D Structure (crystal)
3D Structure (crystal)