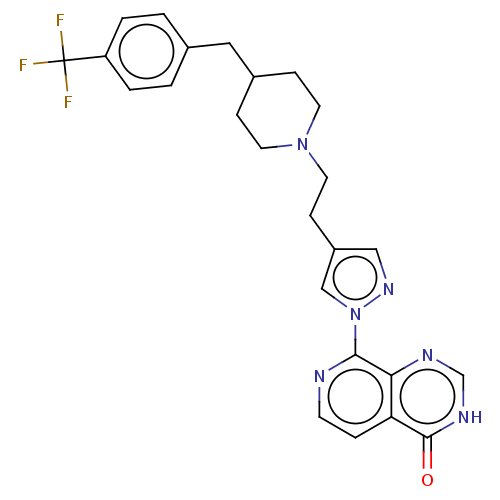
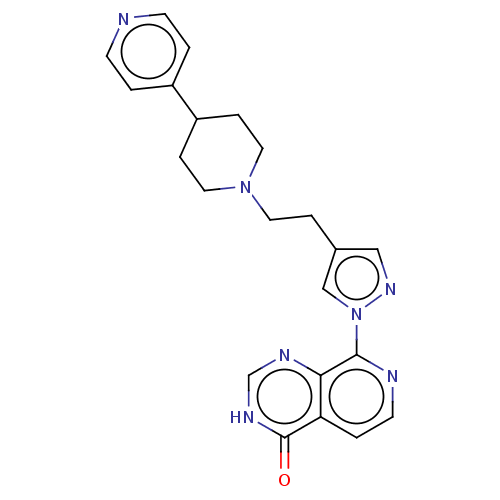
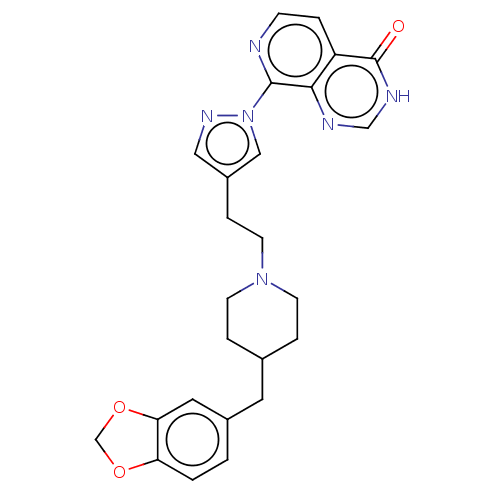
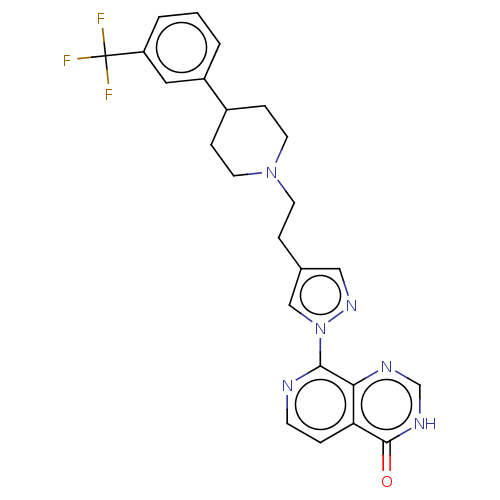
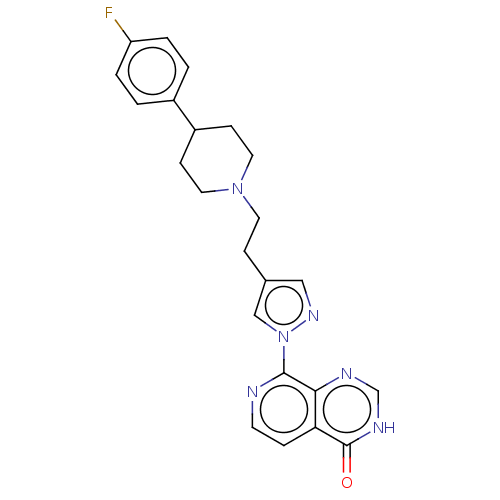
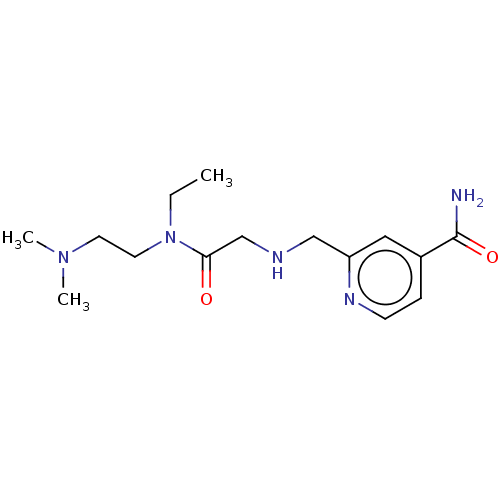
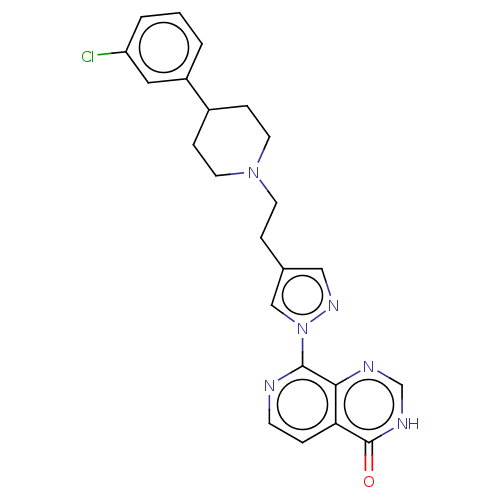
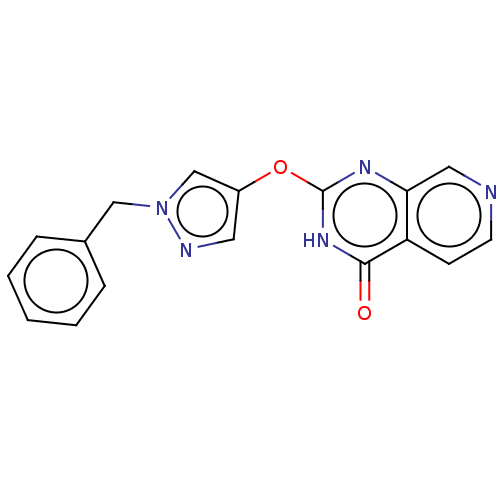
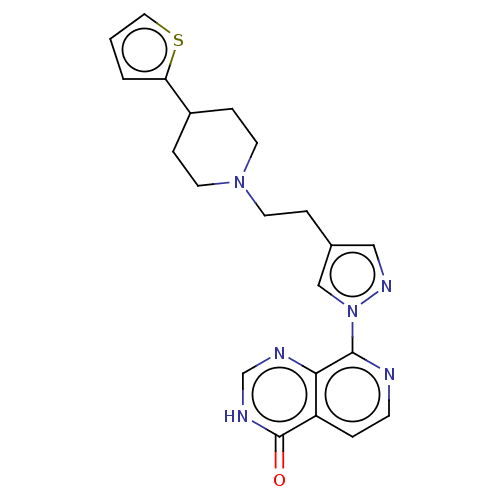
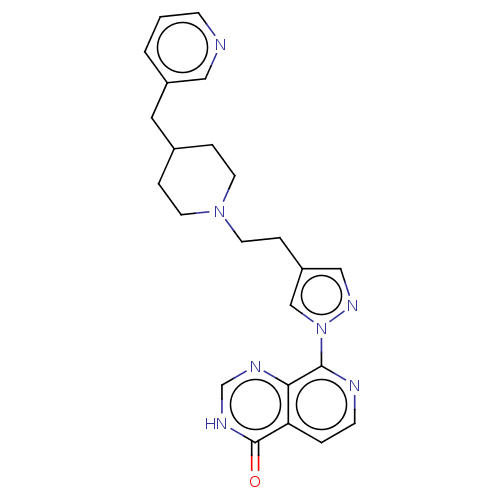
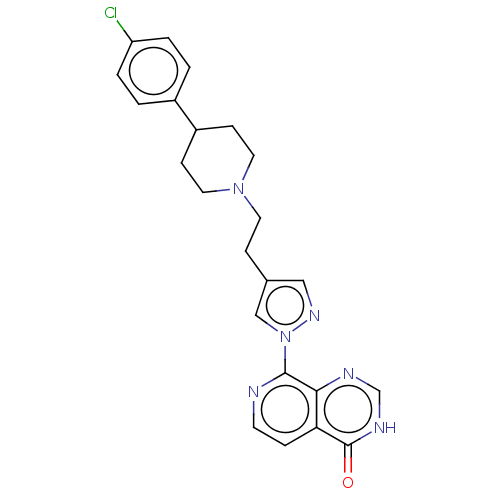
Affinity DataKi: 1.97E+3nMAssay Description:Competitive inhibition of human KDM2A expressed in Escherichia coli using 2-oxoglutarate by enzyme kinetic assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+4nMAssay Description:Mixed type inhibition of human KDM2A expressed in Escherichia coli assessed inhibition constant for compound-enzyme-substrate complex using methyl ly...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM6B (unknown origin) using biotin-H3K27me3 (21 to 44 residues) as substrate preincubated for 15 mins followed by substrate addition m...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of KDM5B (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 31nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 32nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 5C(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 34nMAssay Description:Inhibition of KDM5C (unknown origin) using biotin-H3K4me3 as substrate preincubated for 15 mins followed by substrate addition measured after 20 mins...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 36nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair
TargetLysine-specific demethylase 4B(Homo sapiens (Human))
The Institute Of Cancer Research
Curated by ChEMBL
The Institute Of Cancer Research
Curated by ChEMBL
Affinity DataIC50: 39nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotin-H3K9me3 as s...More data for this Ligand-Target Pair

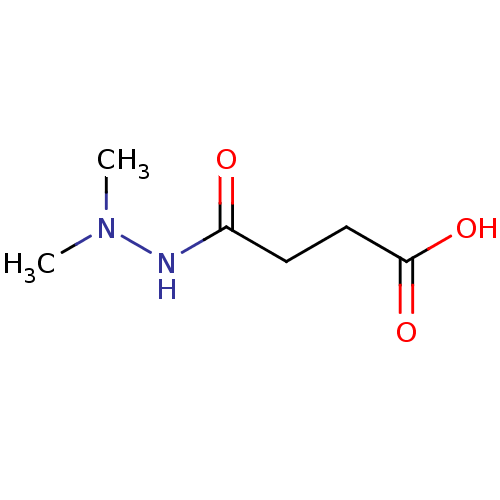
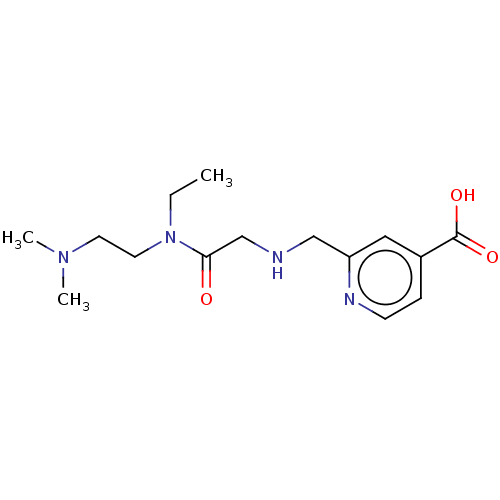

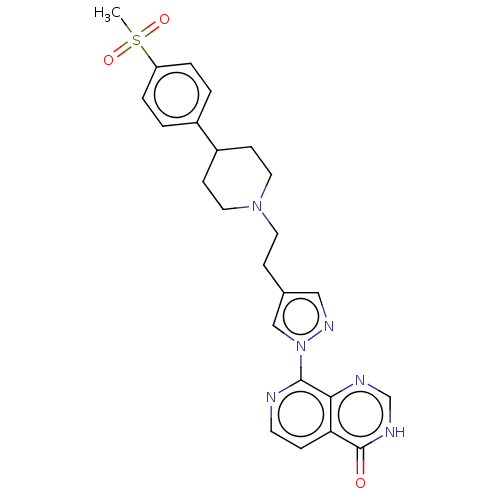
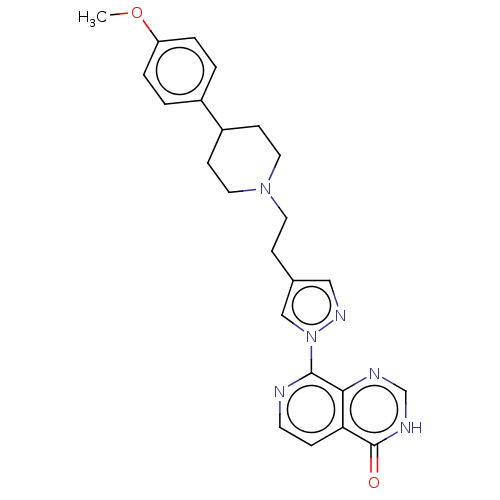
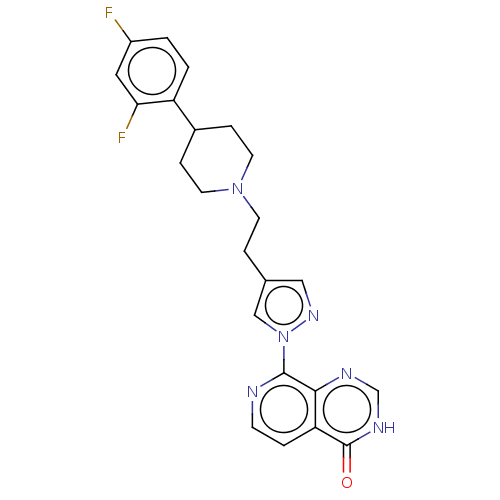

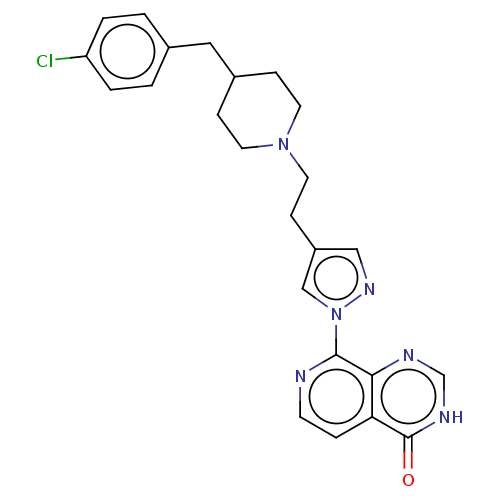
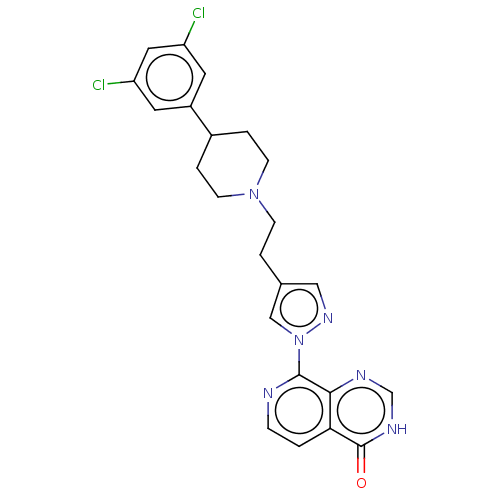
 3D Structure (crystal)
3D Structure (crystal)