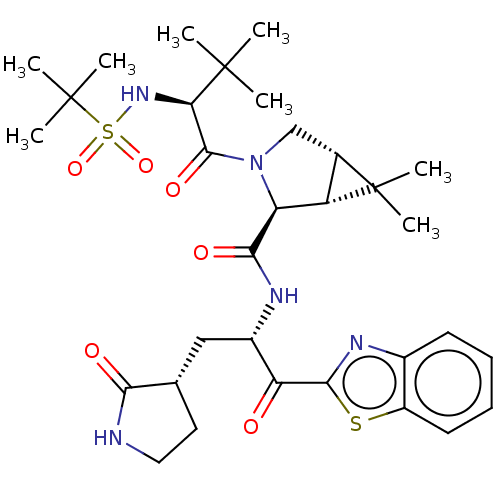
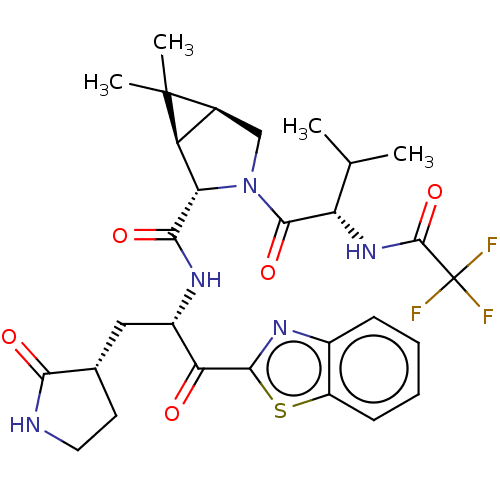
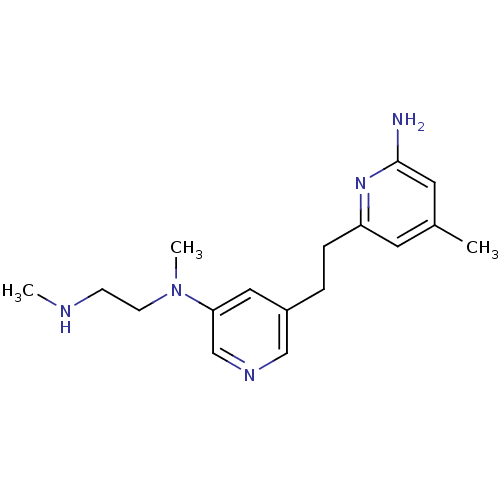
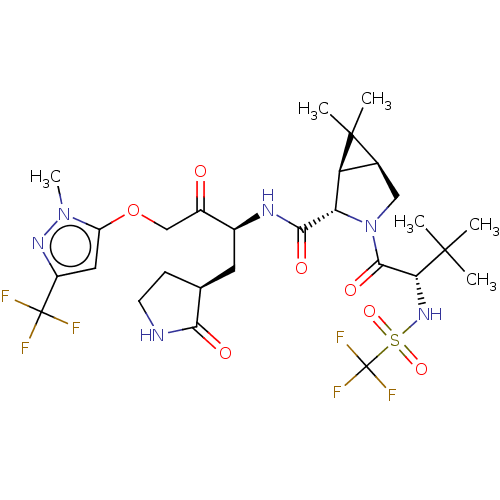
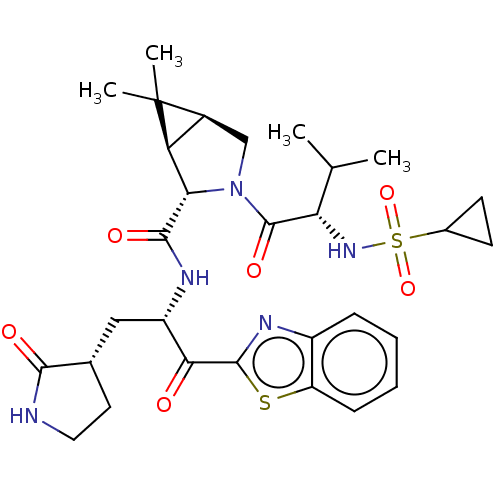
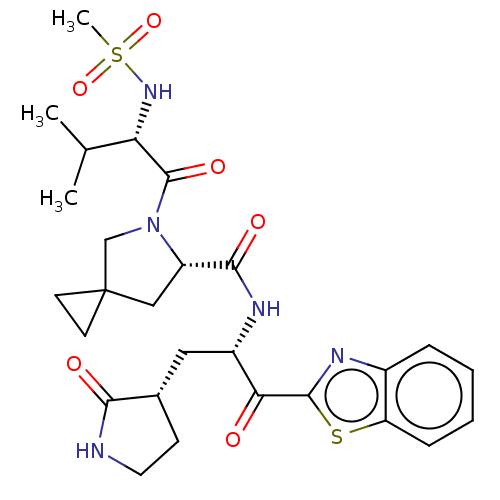
Affinity DataKi: 0.210nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: <0.440nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 1.41nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 1.55nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 1.76nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 1.78nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 1.78nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.02nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.15nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.15nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.42nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.76nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.87nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 2.88nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 3.04nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 3.5nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 3.70nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]SCH-58261 from human recombinant adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.12nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 4.22nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 5.45nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]SCH-58261 from human recombinant adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.02nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 6.14nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 6.34nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 7.05nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 7.74nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 7.87nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 8.34nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 8.84nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]SCH-58261 from human recombinant adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 9.77nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]desArg from human B1 in human WI 38 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]SCH-58261 from human recombinant adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Inhibition of human recombinant nNOS expressed in Escherichia coli using L-arginine as substrate assessed as reduction in NO production measured for ...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibition of rat recombinant nNOS expressed in Escherichia coli using L-arginine as substrate assessed as reduction in NO production measured for 6 ...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Enzyme inhibition was evaluated by measuring NO production with the hemoglobin capture assay, which was performed with purified NOSs in 96-well plate...More data for this Ligand-Target Pair
Affinity DataKi: 19.8nMAssay Description: The ability of compounds to prevent SARS-CoV-2 coronavirus-induced cell death or cytopathic effect can be assessed via cell viability, using an assa...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair

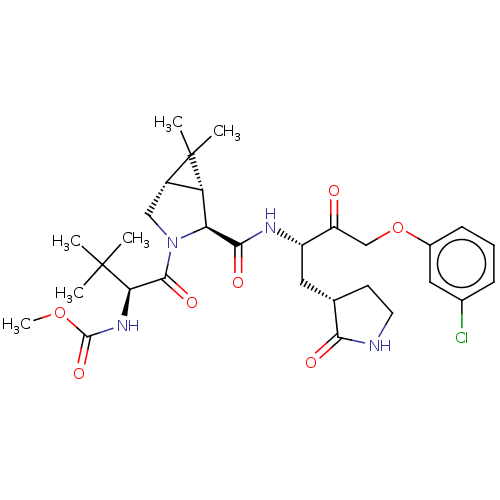
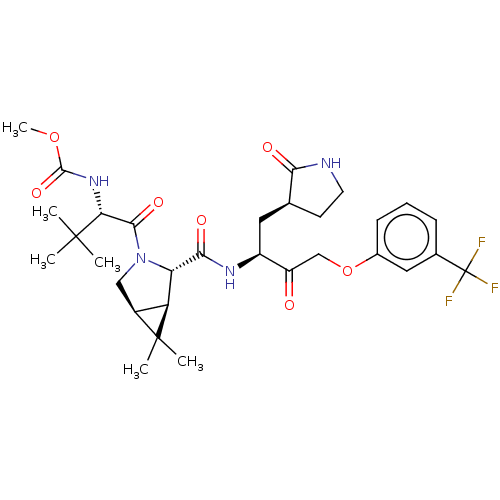
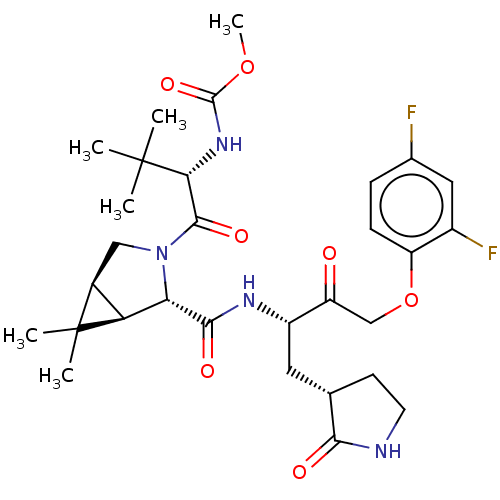
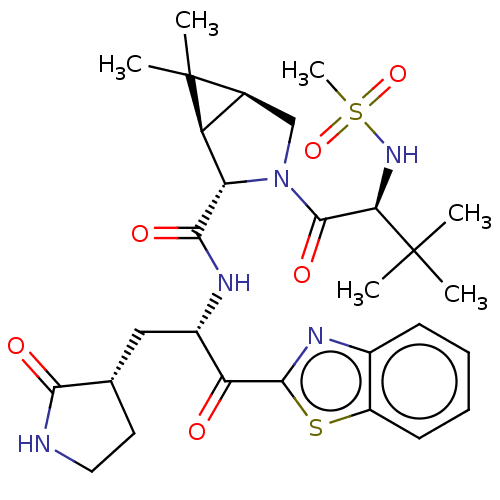
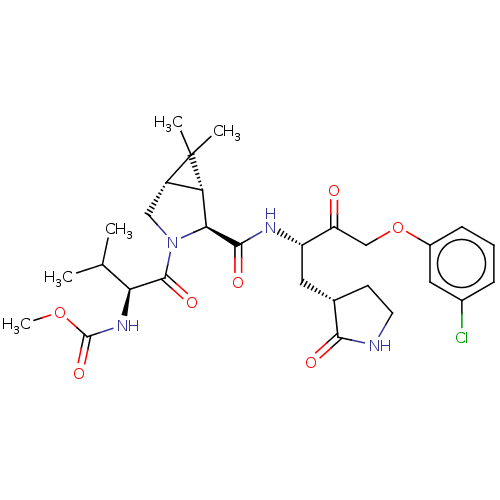
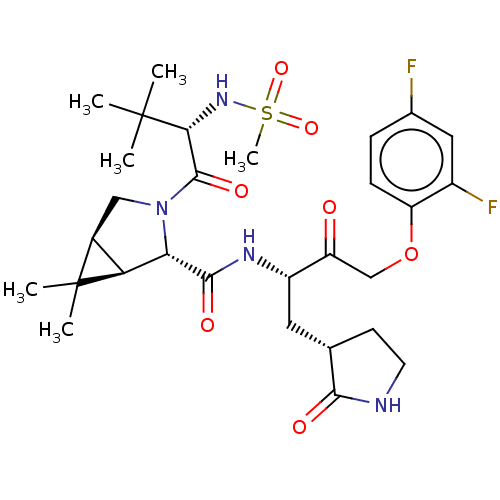

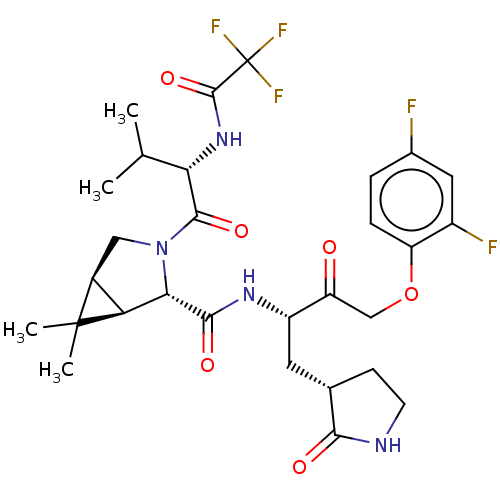
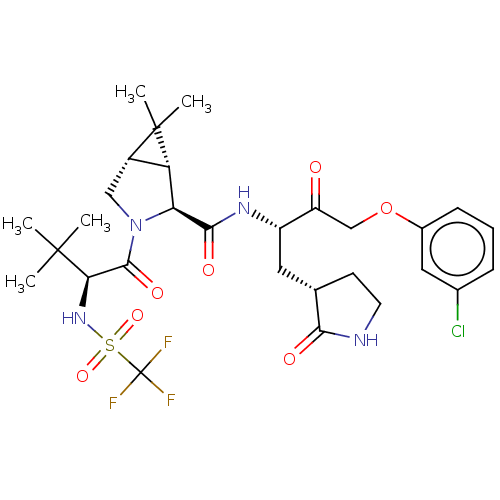
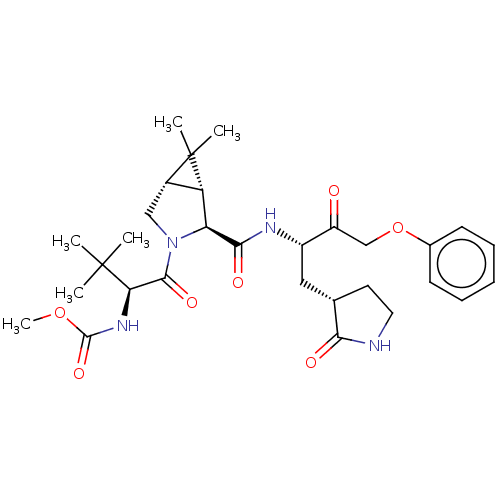
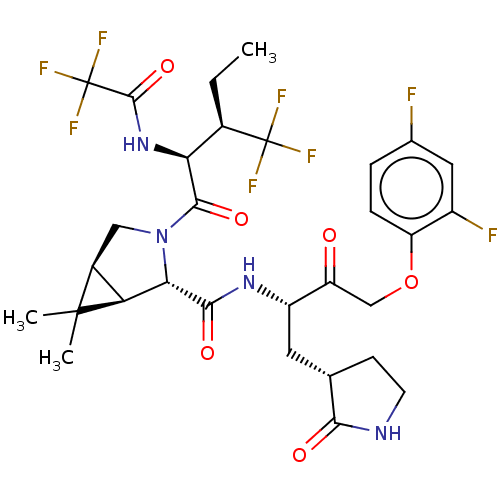


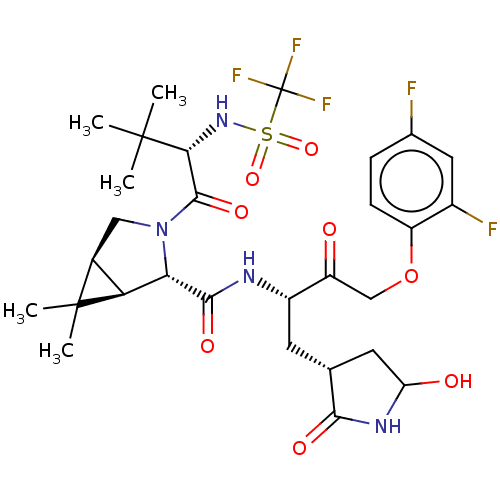
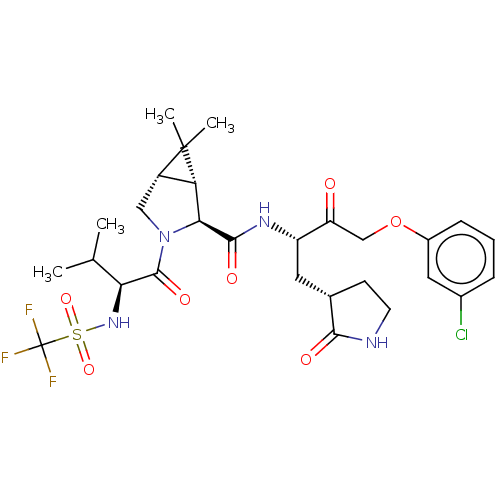
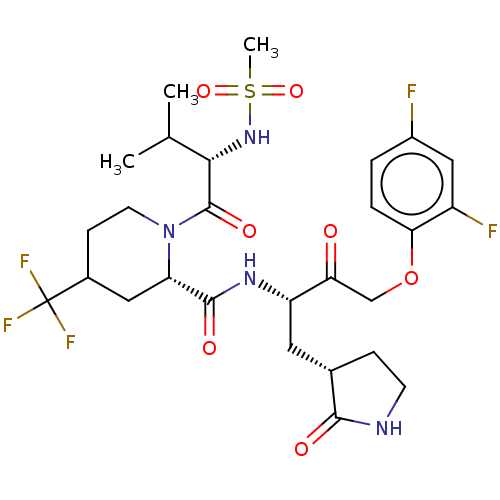
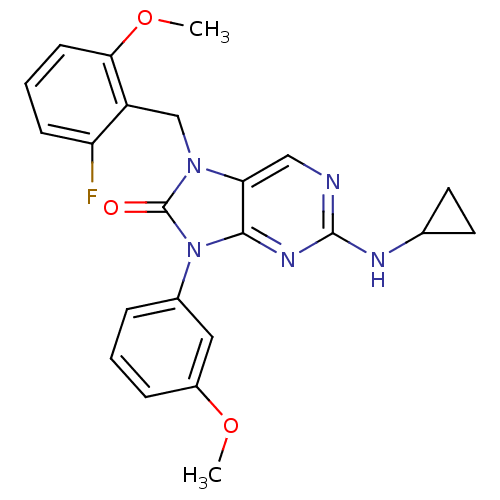
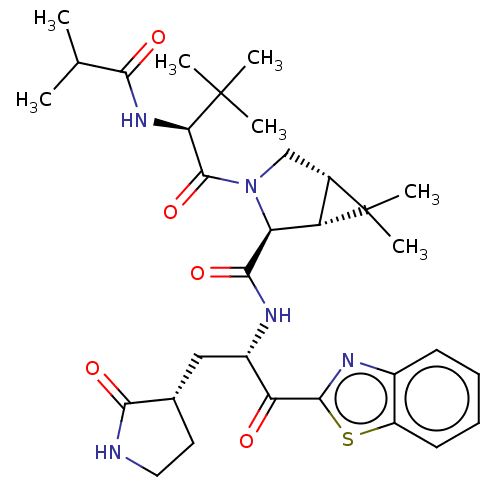
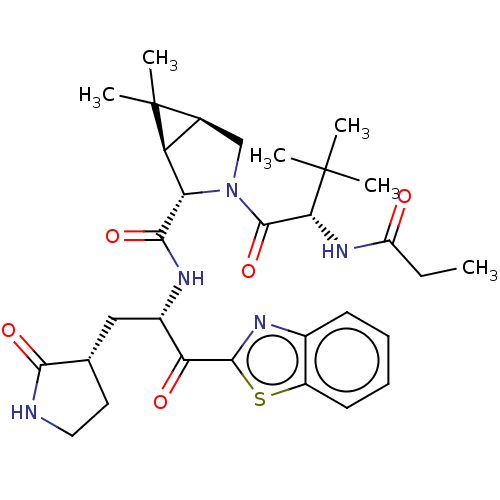
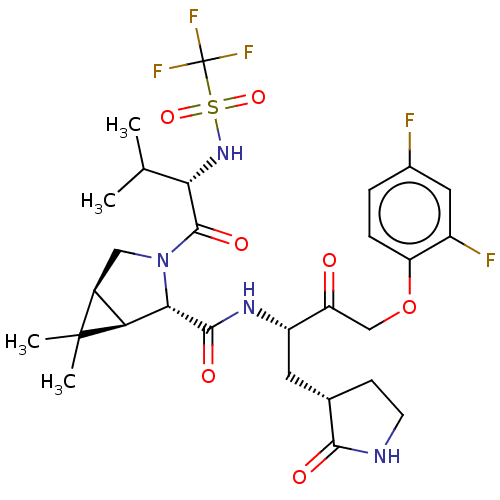
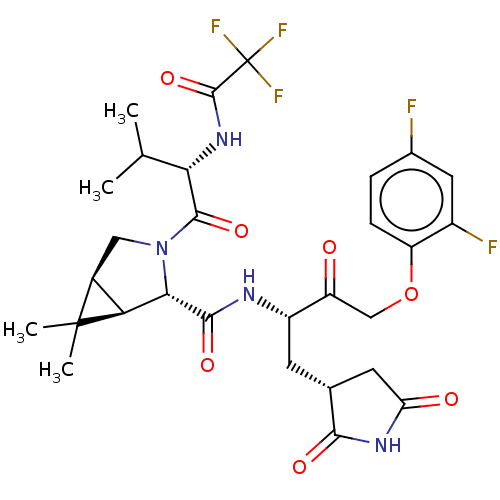
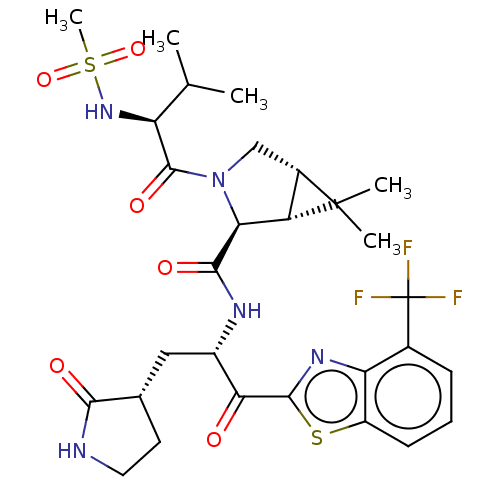

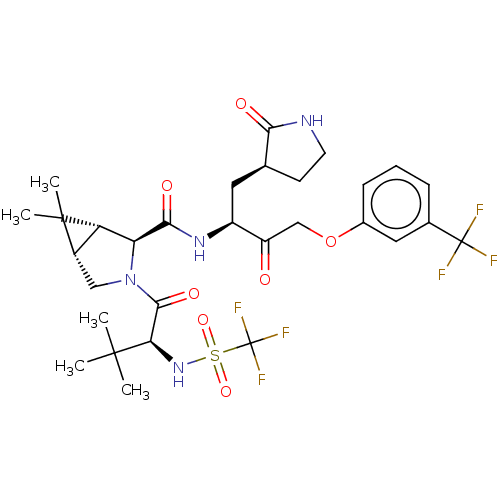

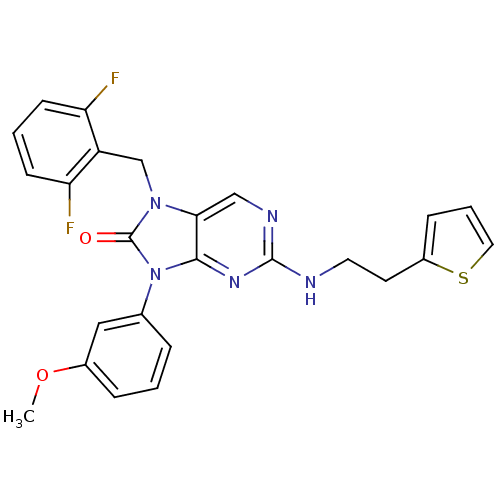
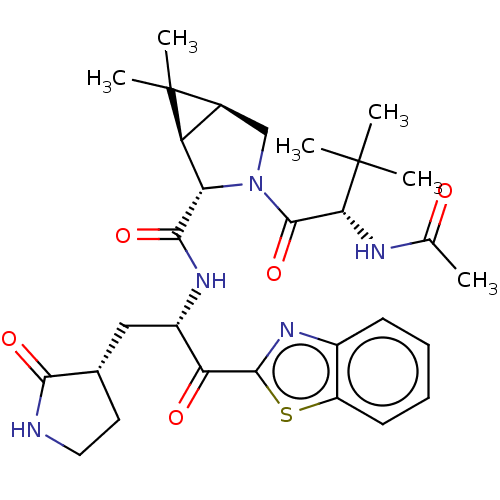



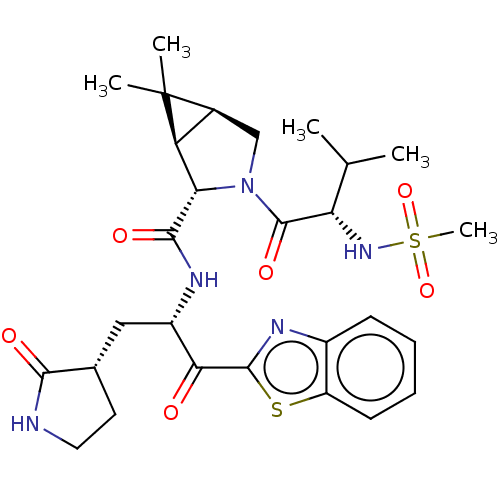
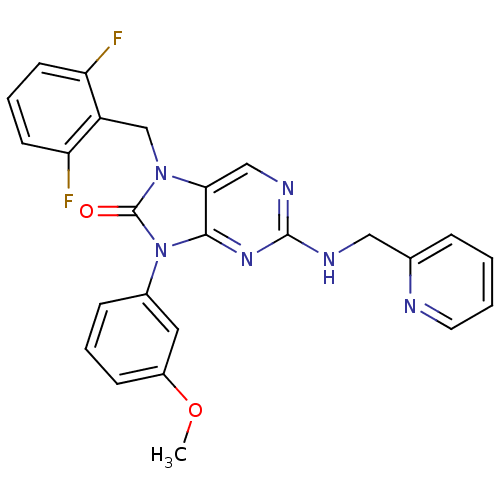
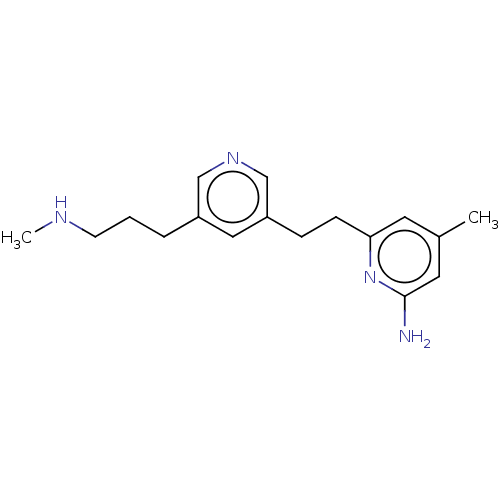
 3D Structure (crystal)
3D Structure (crystal)