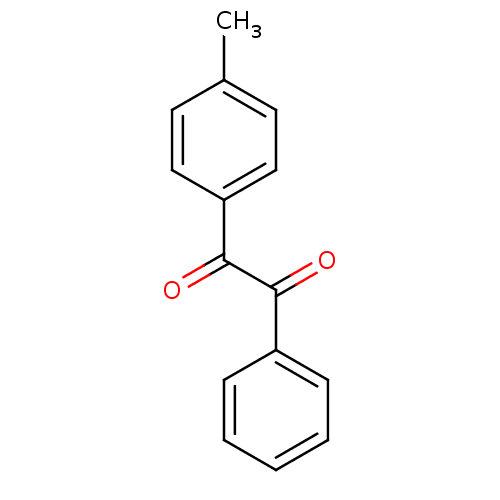
Affinity DataKi: 1.5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nM ΔG°: -47.9kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 5nM ΔG°: -47.4kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 5.60nM ΔG°: -47.1kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nM ΔG°: -46.1kJ/molepH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -46.3kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 7.90nM ΔG°: -46.2kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 8.90nM ΔG°: -46.0kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 9.5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 10.3nM ΔG°: -45.6kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 11.9nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 14.7nM ΔG°: -44.7kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 14.7nMAssay Description:Inhibition constant against human intestinal carboxylesterase (hiCE) expressed in Sf21 cells using 3 mM o-NPAMore data for this Ligand-Target Pair
Affinity DataKi: 14.7nM ΔG°: -44.7kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 15.9nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 16.2nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 17.5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 18.2nM ΔG°: -44.2kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 21.3nM ΔG°: -43.8kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 21.7nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 22.8nM ΔG°: -43.6kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 23.4nM ΔG°: -43.6kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 23.9nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Inhibition constant against human liver carboxylesterase (hCE1) expressed in Sf21 cells using 3 mM o-NPAMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Inhibition constant against human liver carboxylesterase (hCE1) expressed in Sf21 cells using 3 mM o-NPAMore data for this Ligand-Target Pair
Affinity DataKi: 30.6nM ΔG°: -42.9kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 31nM ΔG°: -42.9kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 32nM ΔG°: -42.8kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 33.3nM ΔG°: -42.7kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 36nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 45.1nMAssay Description:Inhibition constant against human liver carboxylesterase (hCE1) expressed in Sf21 cells using 3 mM o-NPAMore data for this Ligand-Target Pair
Affinity DataKi: 45.1nMAssay Description:Inhibition constant against human liver carboxylesterase (hCE1) expressed in Sf21 cells using 3 mM o-NPAMore data for this Ligand-Target Pair


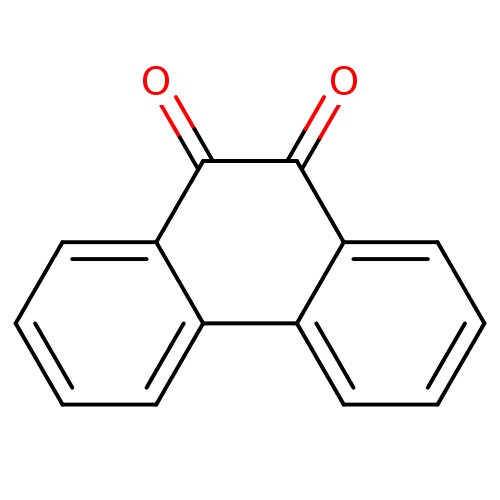
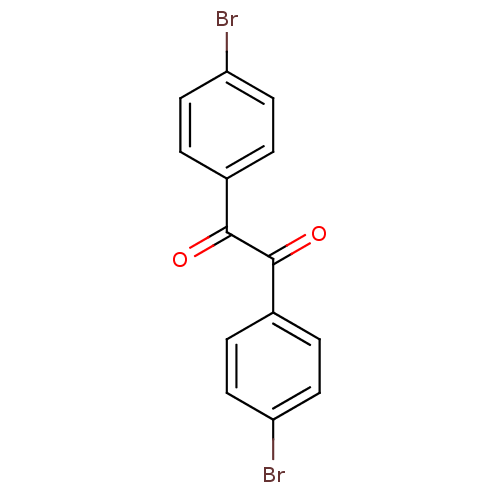
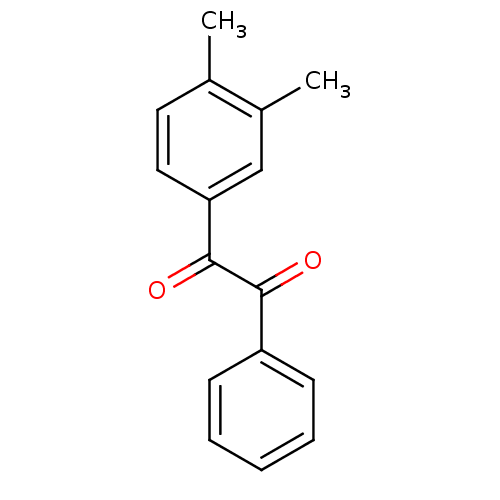
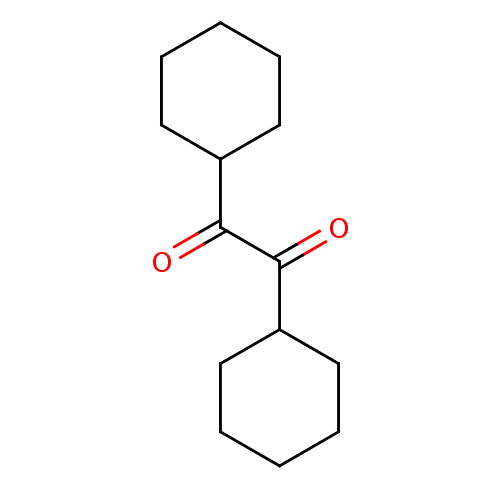

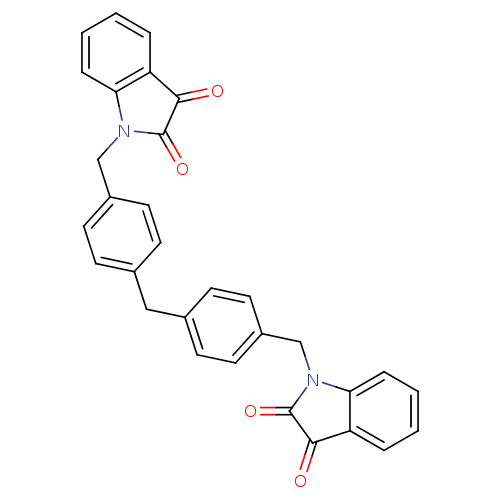
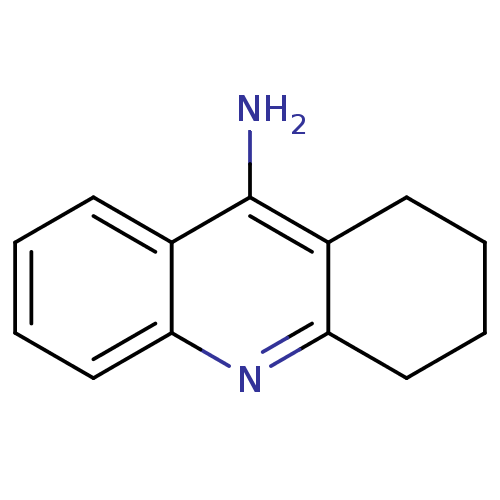
 3D Structure (crystal)
3D Structure (crystal)