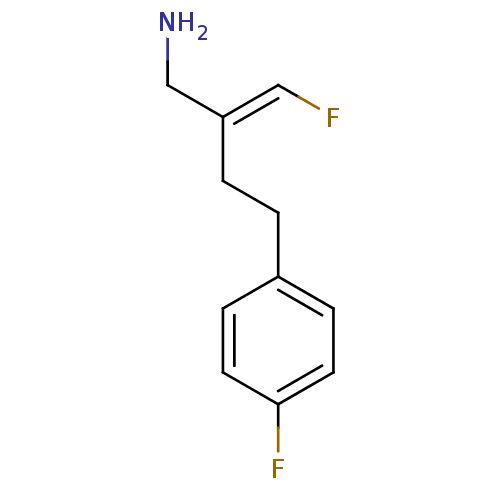
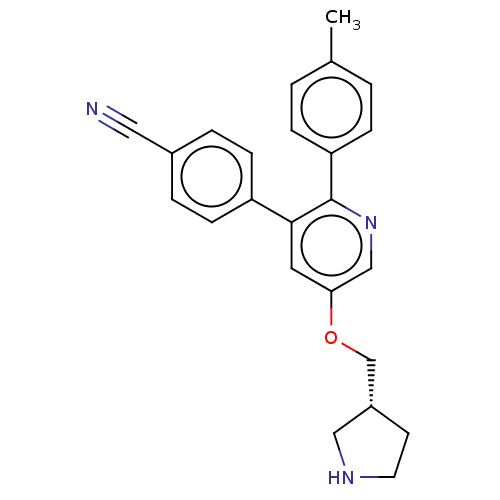
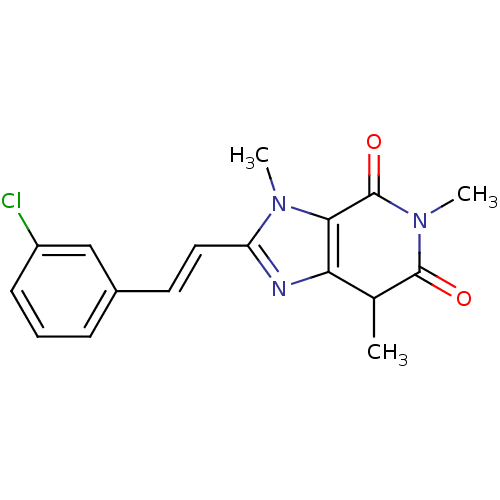
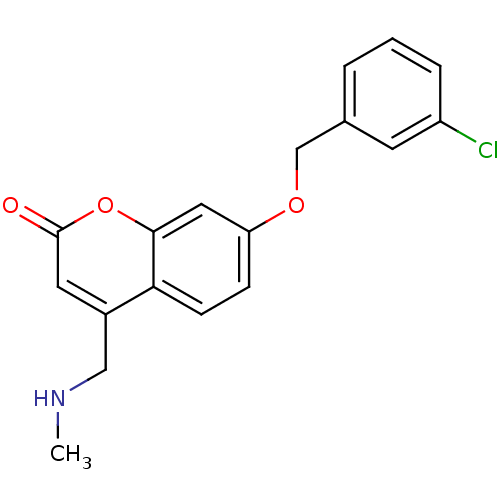
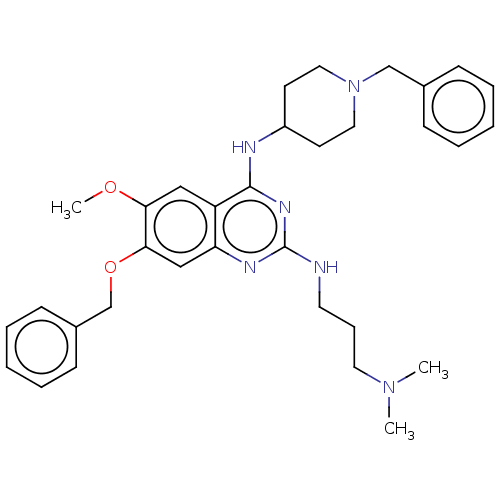
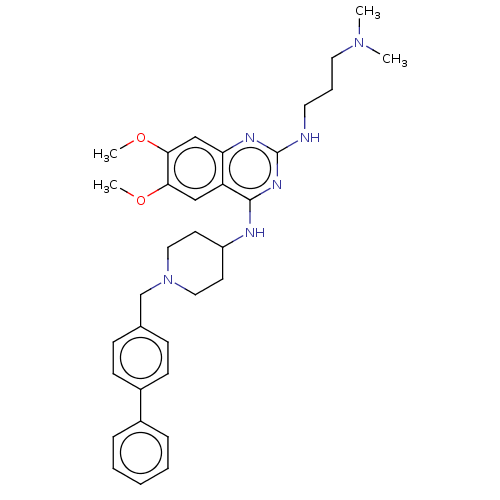
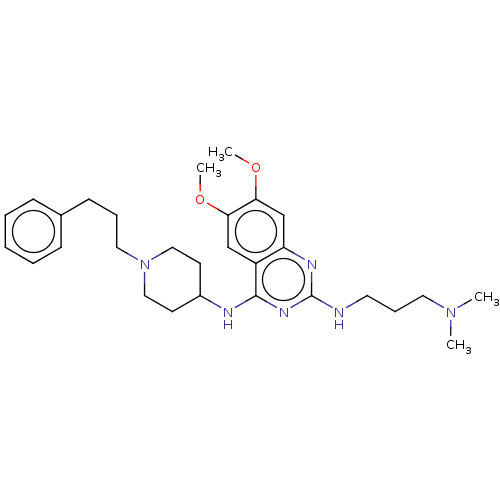
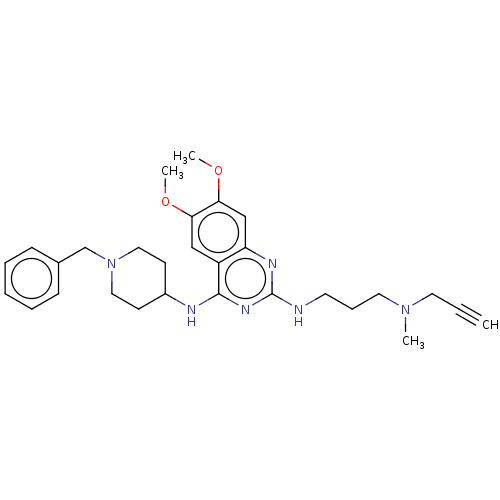
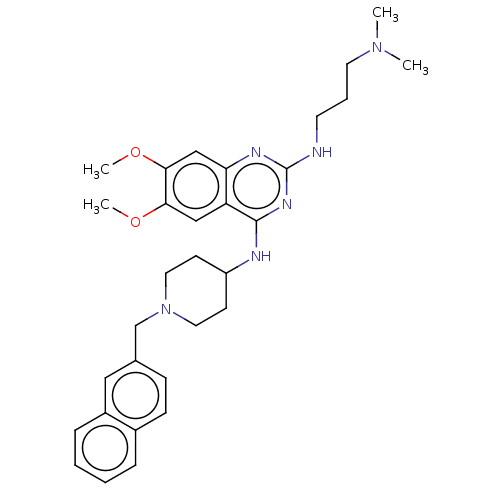
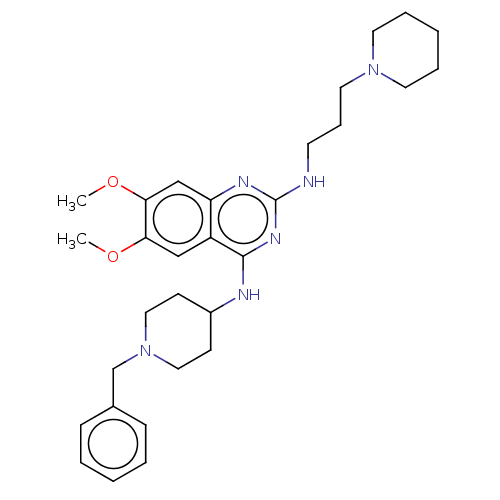
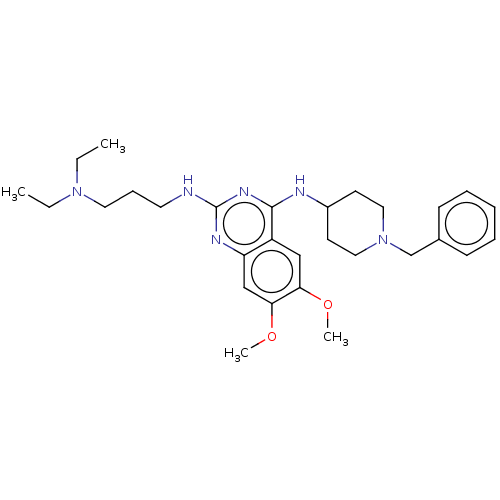
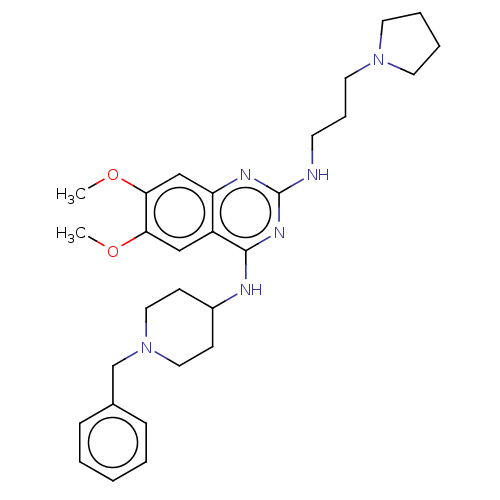
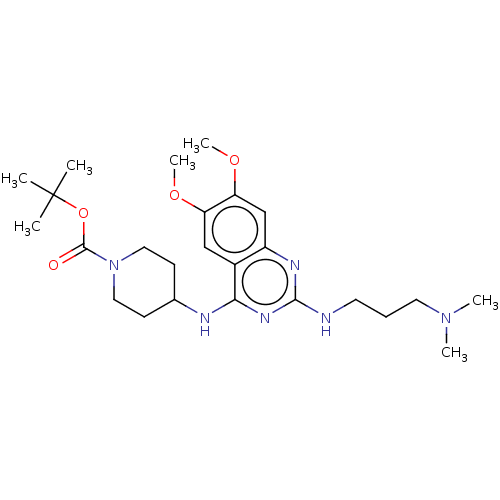
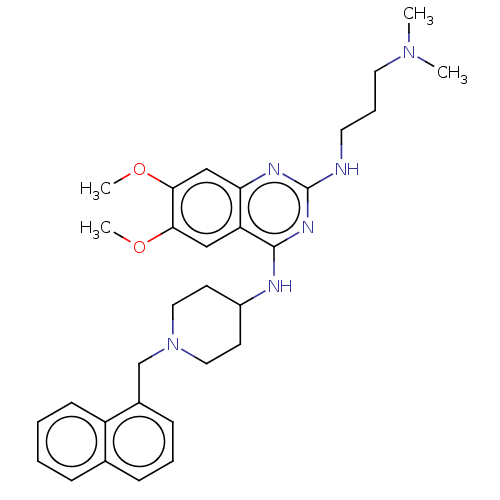
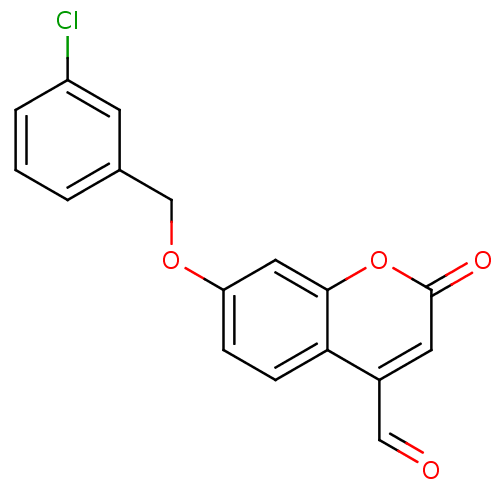
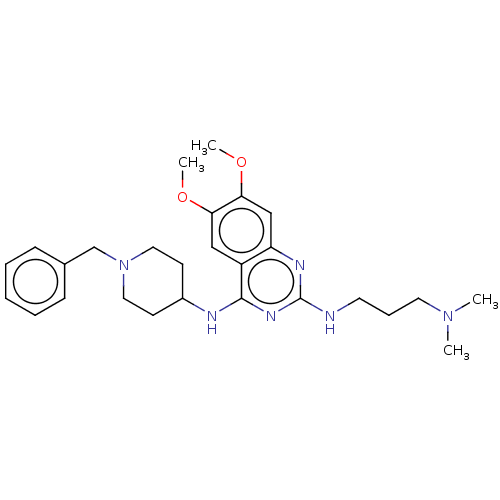
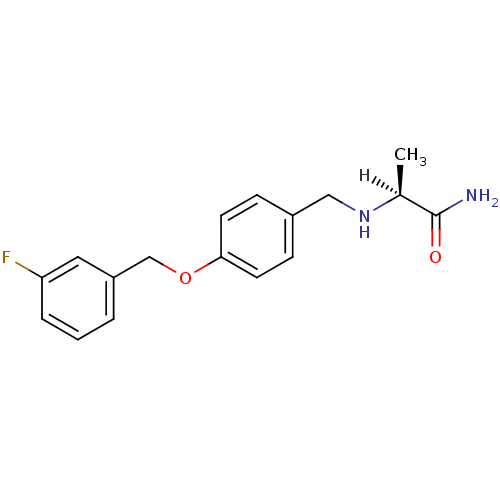
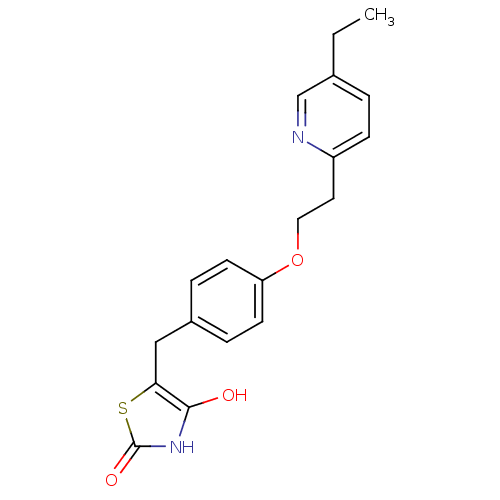
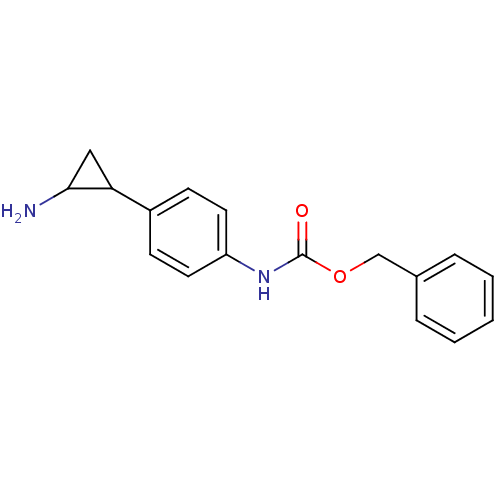
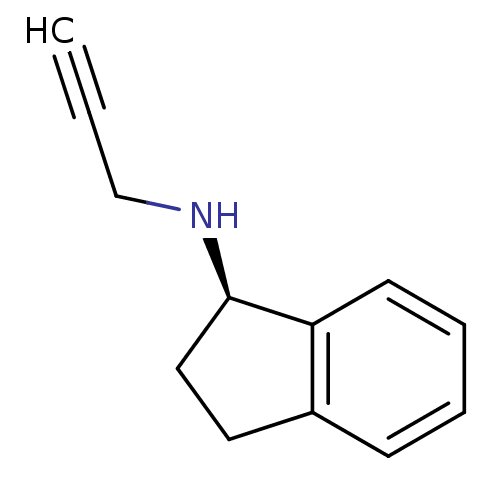
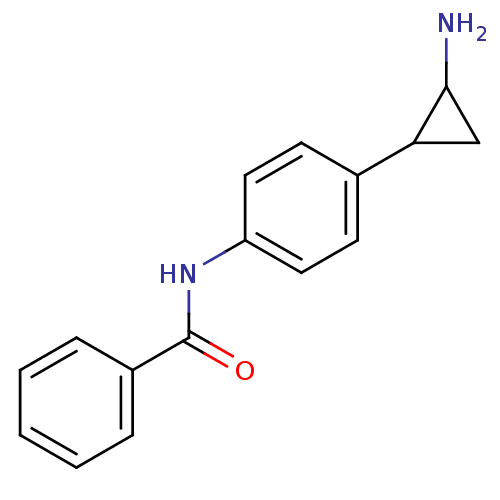
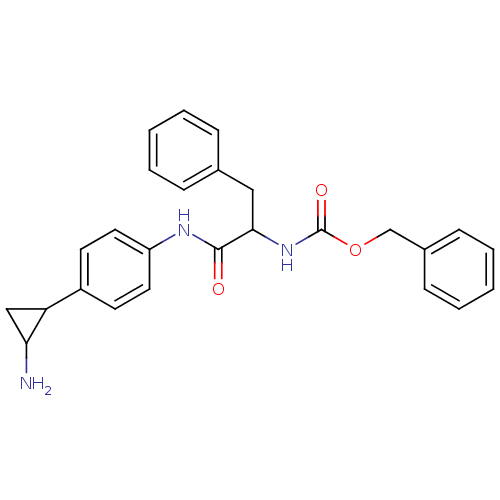
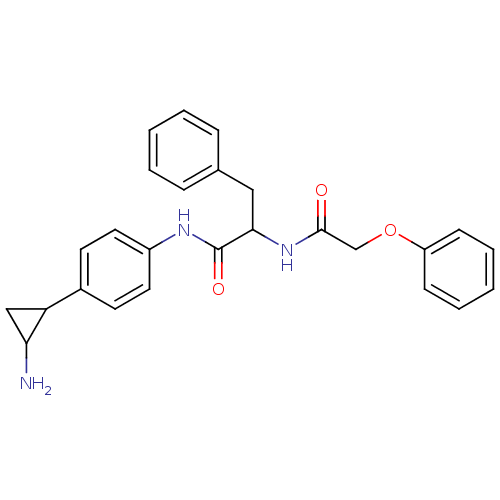
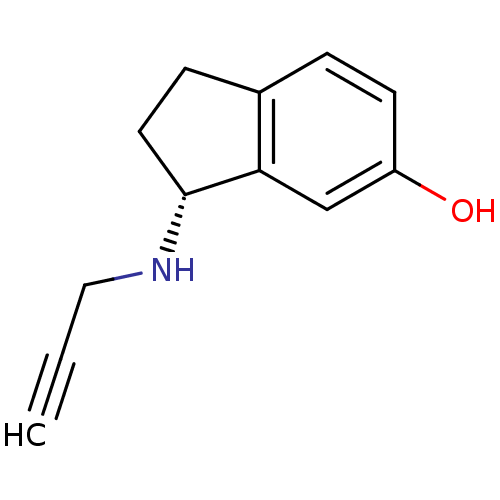
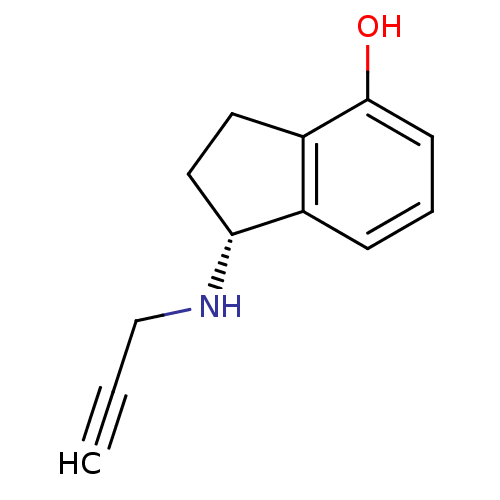
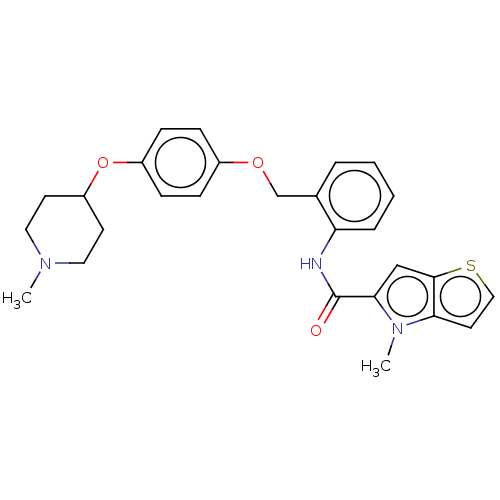
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Inhibition of human recombinant MAOB expressed in Pichia pastoris by kinetic assayMore data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nM ΔG°: -40.0kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 100nM ΔG°: -40.0kJ/molepH: 7.5 T: 2°CAssay Description:MAO B activities were determined spectrophotometrically at 250 nm using benzylamine as substrate. Competitive Ki values were determined by measuring ...More data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 146nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 149nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 153nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 175nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 183nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.2kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 212nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 400nM ΔG°: -36.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO B activities were determined spectrophotometrically at 250 nm using benzylamine as substrate. Competitive Ki values were determined by measuring ...More data for this Ligand-Target Pair
Affinity DataKi: 440nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 450nM ΔG°: -36.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO B activities were determined spectrophotometrically at 250 nm using benzylamine as substrate. Competitive Ki values were determined by measuring ...More data for this Ligand-Target Pair
Affinity DataKi: 500nMAssay Description:Competitive inhibition of human MAOB expressed in Pichia pastorisMore data for this Ligand-Target Pair
Affinity DataKi: 500nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
Affinity DataKi: 500nMAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 500nMAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 569nMAssay Description:Inhibition of human LSD1/CoREST using histone H3 peptide as substrate by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 680nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 690nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.1kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 1.10E+3nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition of LSD1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition of human recombinant MAOA expressed in Pichia pastoris by kinetic assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
Sapienza University Of Rome
Curated by ChEMBL
Sapienza University Of Rome
Curated by ChEMBL
Affinity DataKi: 1.18E+3nMAssay Description:Inhibition of G9a (unknown origin) assessed as reduction in substrate methylation using histone H3 and SAM as substrate measured after 15 to 60 mins ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 1.30E+3nMAssay Description:Inhibition of LSD1 (unknown origin)More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 1.30E+3nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nM ΔG°: -33.4kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 1.90E+3nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nM ΔG°: -32.7kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.5kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically at 316 nm and 250 nm using kynuramine and benzylamine as substrates, respectively. ...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+3nMAssay Description:Competitive inhibition of rat MAOB expressed in Pichia pastorisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Homo sapiens (Human))
Sapienza University of Rome
US Patent
Sapienza University of Rome
US Patent
Affinity DataKi: 2.10E+3nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nM ΔG°: -32.2kJ/molepH: 7.5 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A/REST corepressor 1(Homo sapiens (Human))
European Institute Of Oncology
Curated by ChEMBL
European Institute Of Oncology
Curated by ChEMBL
Affinity DataKi: 2.30E+3nMAssay Description:Inhibitory activity against beta-glucosidase of sweet almondMore data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nMpH: 7.5Assay Description:Human recombinant MAO A and MAO B were expressed in Pichia pastoris and purified as published (Binda C, et al., Proc. Natl. Acad. Sci. USA 100: 9750-...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.1kJ/molepH: 7.4 T: 2°CAssay Description:MAO A and MAO B activities were determined spectrophotometrically. Competitive Ki values for both enzymes were determined by measuring initial rates ...More data for this Ligand-Target Pair

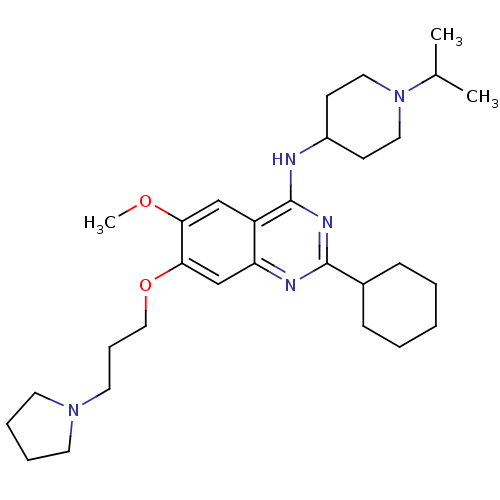
 3D Structure (crystal)
3D Structure (crystal)