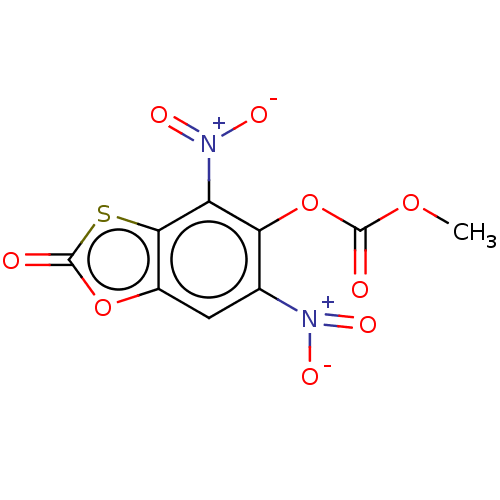
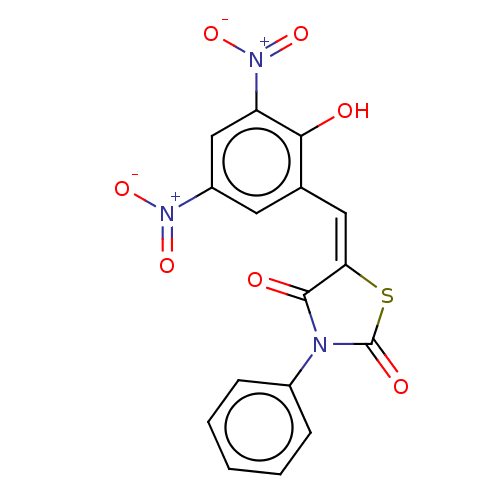
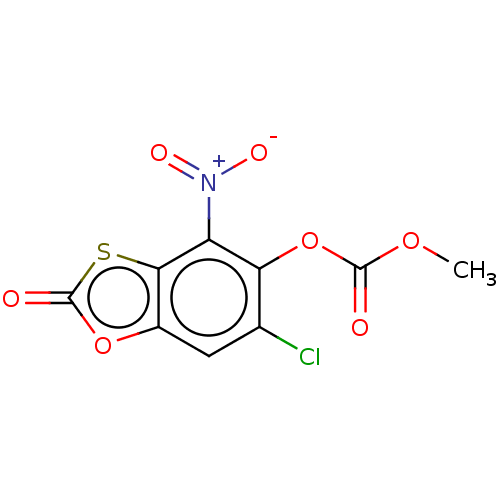
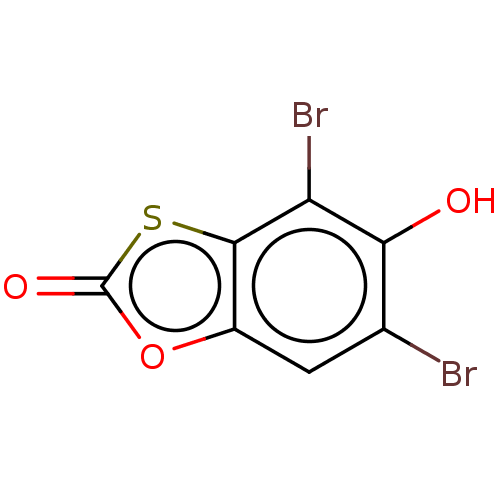
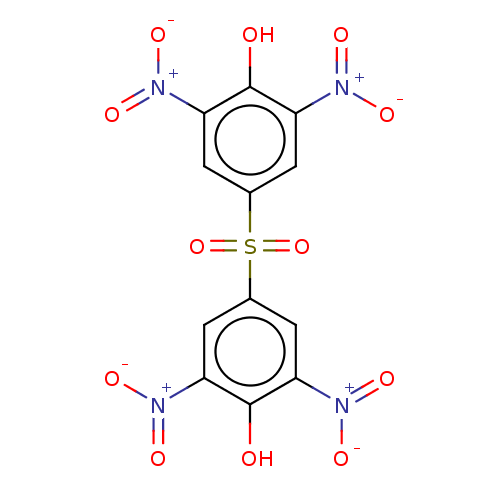
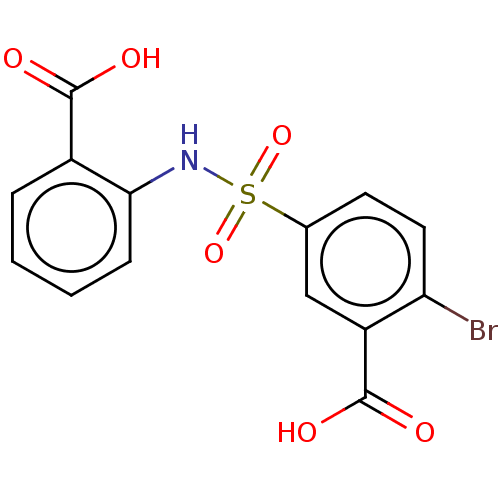
Affinity DataKi: 800nMAssay Description:Competitive inhibition of Yersinia enterocolitica Irp9 using chorismate as substrateMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataKi: 3.00E+4nMAssay Description:Competitive inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrateMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataKi: 4.00E+4nMAssay Description:Competitive inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as pyruvate production after 10 mins by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as salicylate production by fluorescence spectrometryMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+4nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as salicylate production by fluorescence spectrometryMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as salicylate production by fluorescence spectrometryMore data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+4nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as salicylate production by fluorescence spectrometryMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 8.20E+4nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 1.20E+5nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as salicylate production by fluorescence spectrometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of Yersinia enterocolitica Irp9 using chorismate as substrate assessed as pyruvate production after 10 mins by microplate reader analysisMore data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 1.70E+5nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair
TargetBifunctional chorismate mutase/prephenate dehydratase(Escherichia coli (strain K12))
University Of Kansas
Curated by ChEMBL
University Of Kansas
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Escherichia coli K-12 EcCM expressed in Escherichia coli BL21 Star (DE3) pLysS using chorismate as substrate assessed as disappearance ...More data for this Ligand-Target Pair