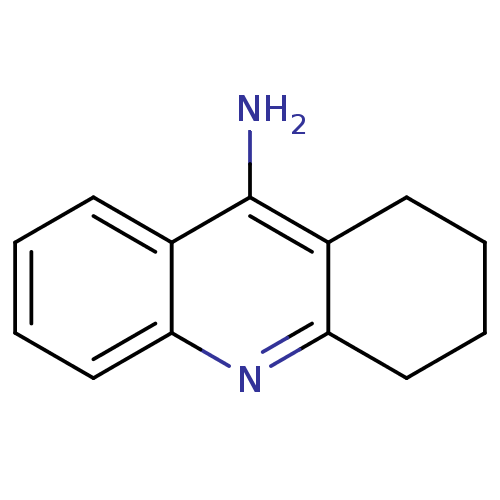
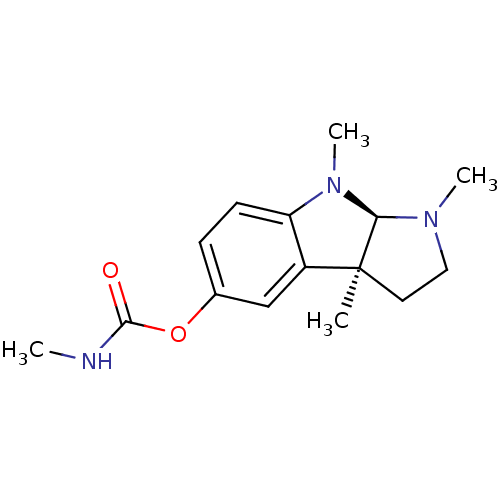
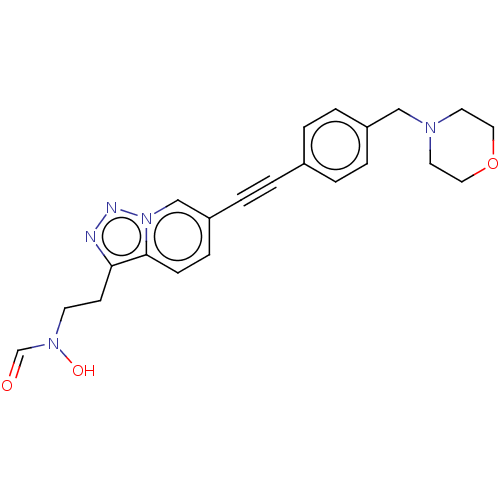
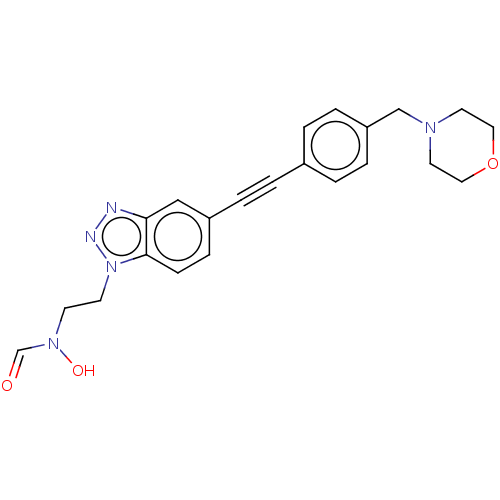
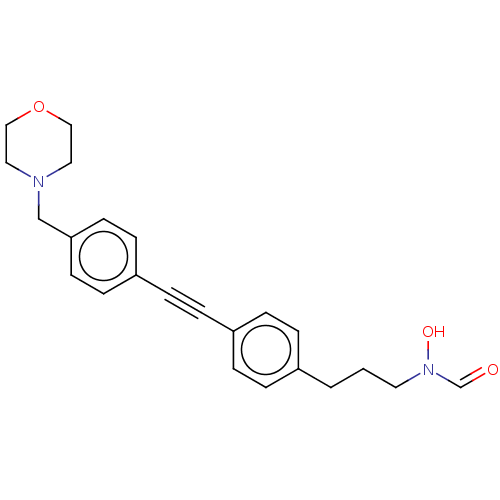
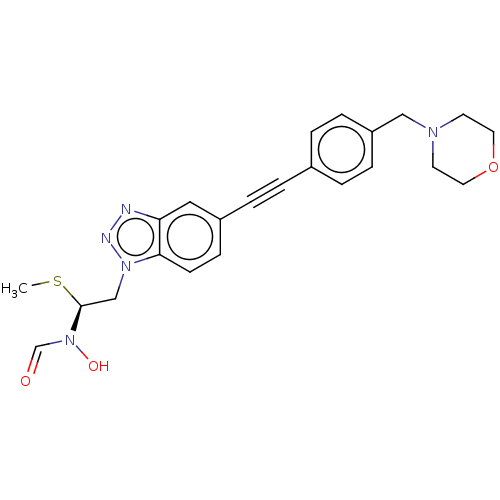
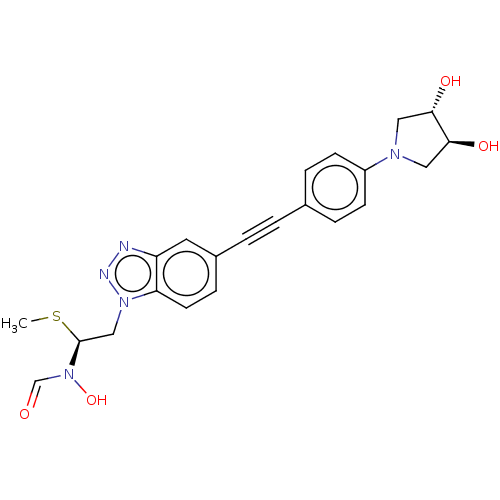
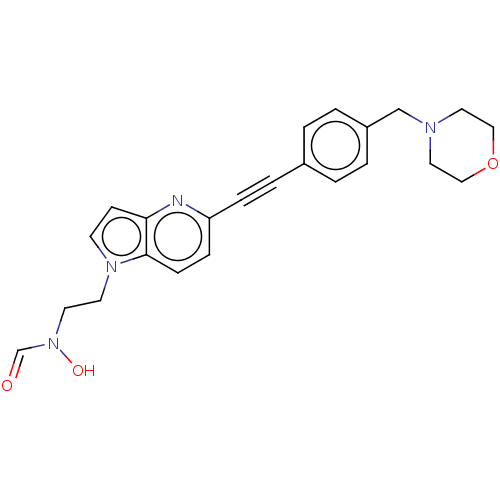
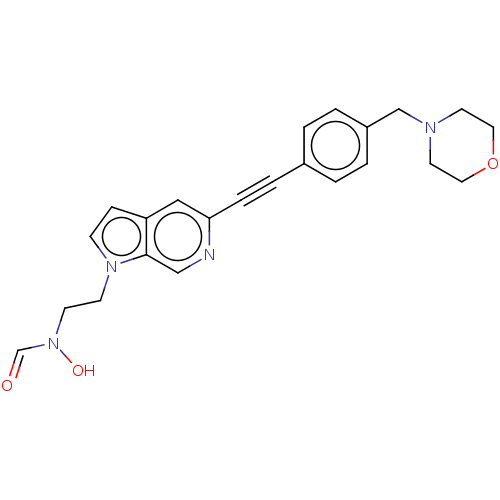
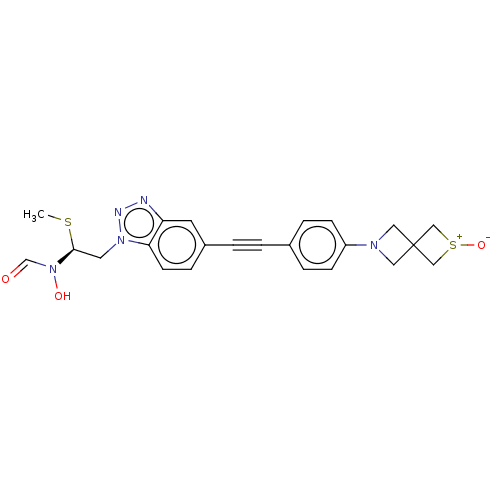
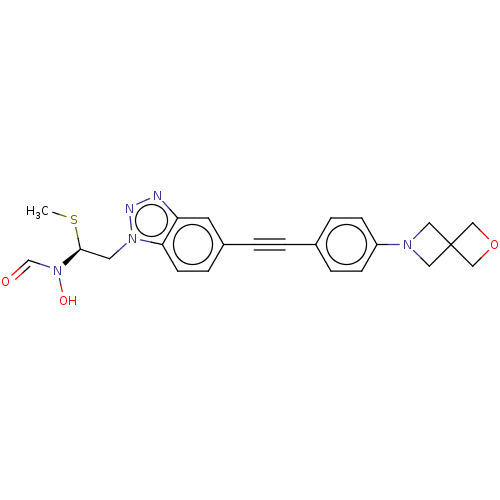
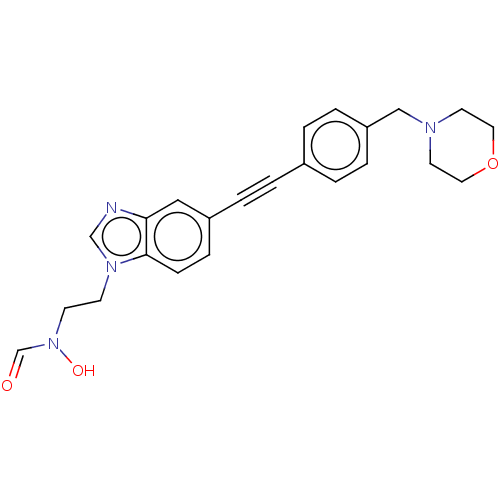
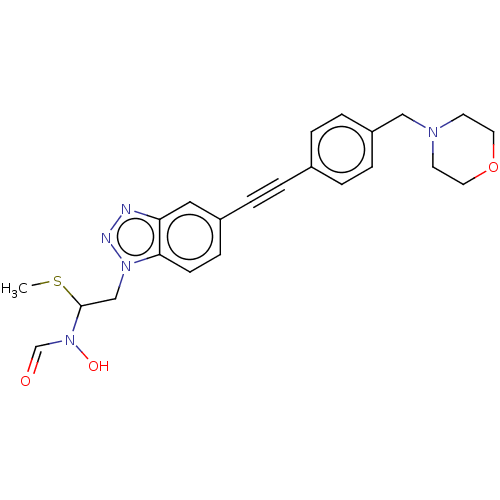
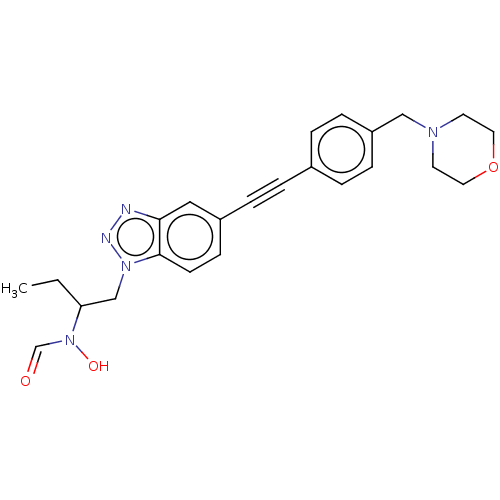
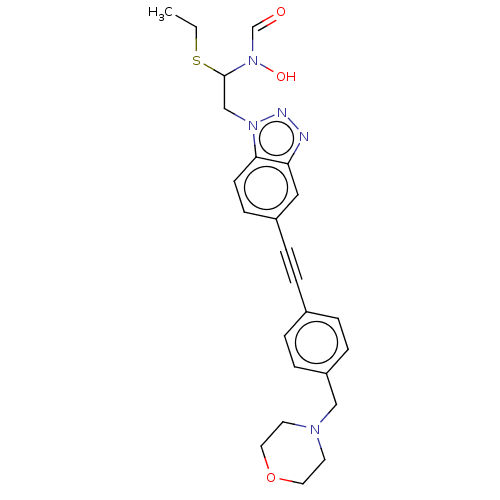
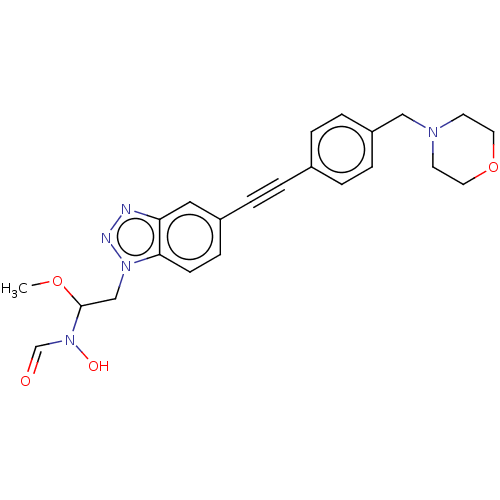
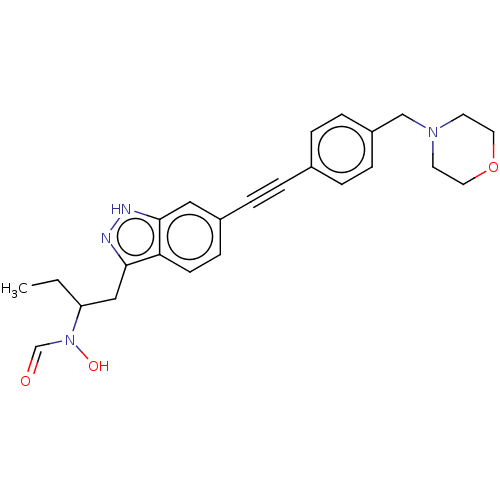
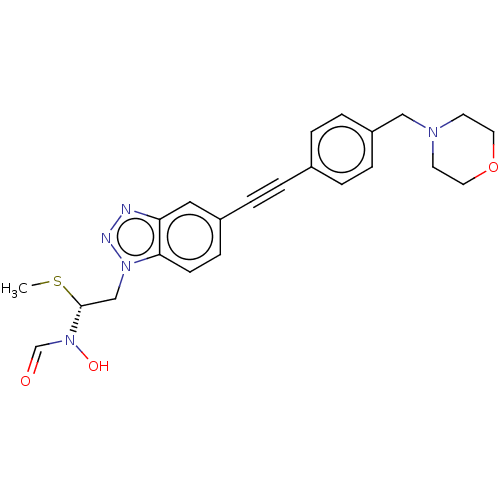
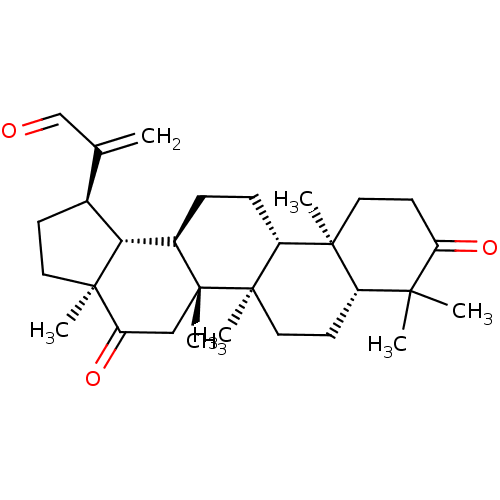
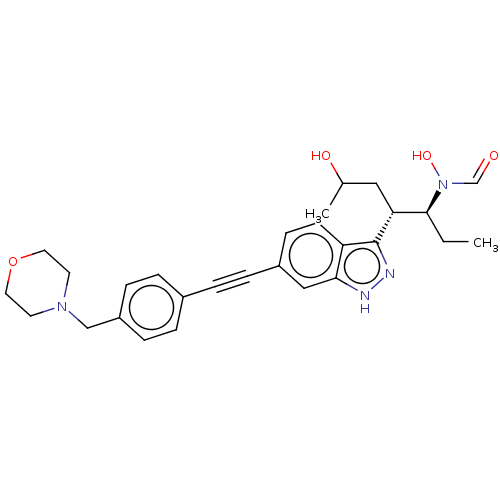
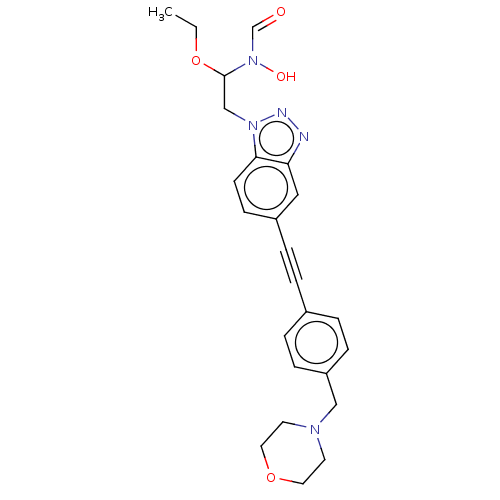
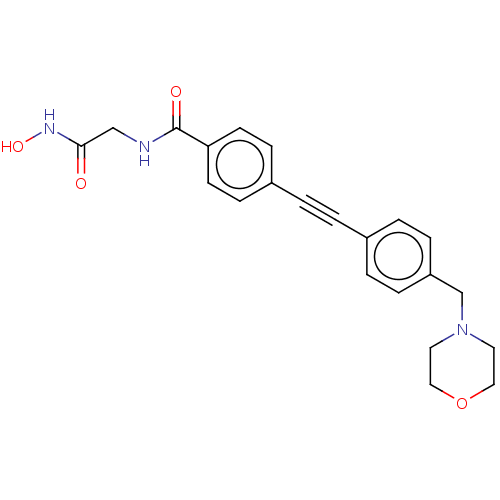
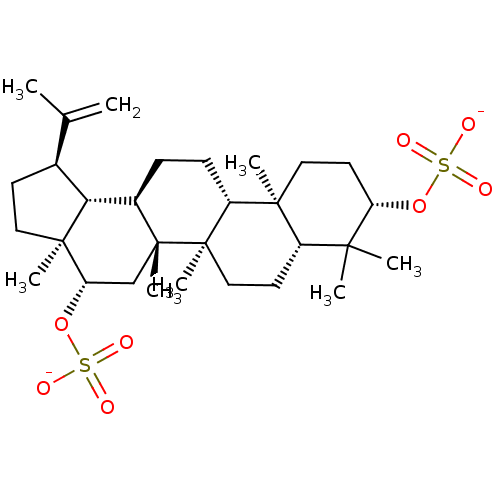
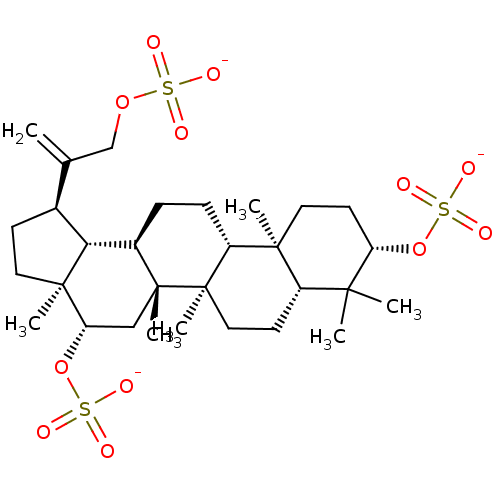
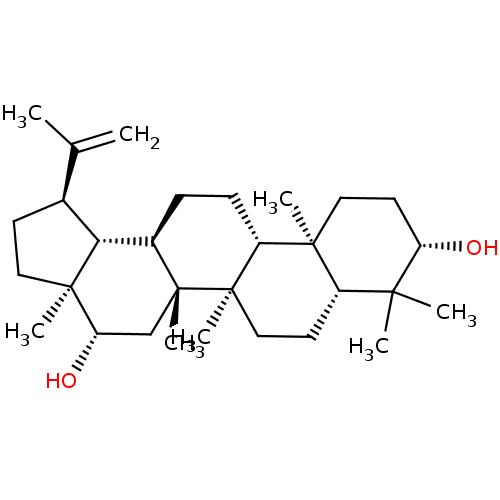
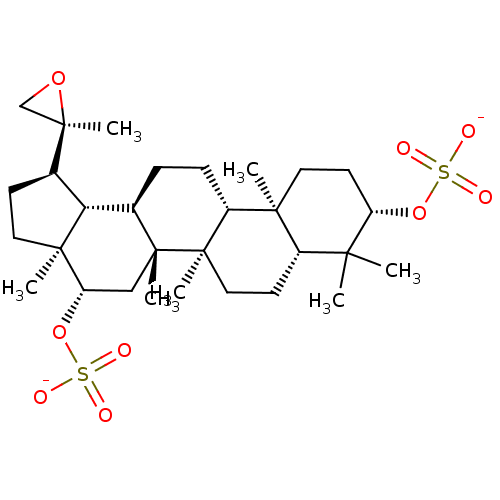
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataKi: 1.44E+5nMAssay Description:Uncompetitive inhibition of Pacific electric ray AChE using acetylthiocholine as substrate by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 64nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: <1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 5.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 7.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.15E+4nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: 5.88E+4nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 6.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 6.30E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.43E+4nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 8.06E+4nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+5nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.88E+5nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: 1.90E+5nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of BChE in horse serum using butyrylthiocholine iodide as substrate after 120 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universidad Nacional Del Sur
Curated by ChEMBL
Universidad Nacional Del Sur
Curated by ChEMBL
Affinity DataIC50: >2.00E+5nMAssay Description:Inhibition of Pacific electric ray AChE using acetylthiocholine iodide as substrate after 60 mins by Ellman's methodMore data for this Ligand-Target Pair

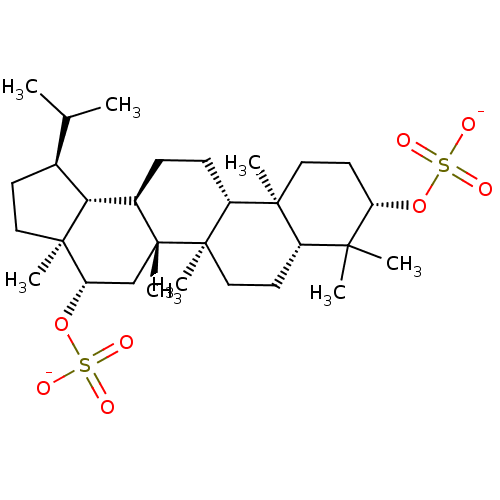
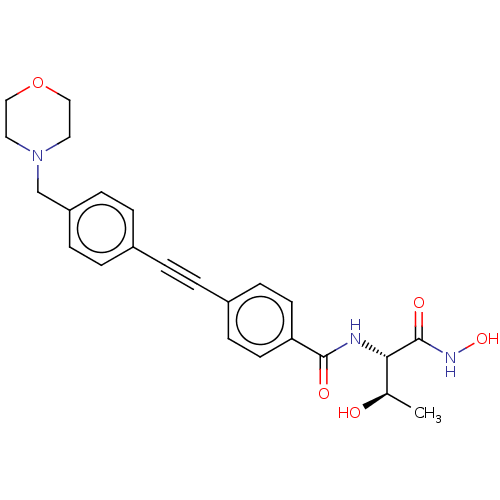
 3D Structure (crystal)
3D Structure (crystal)