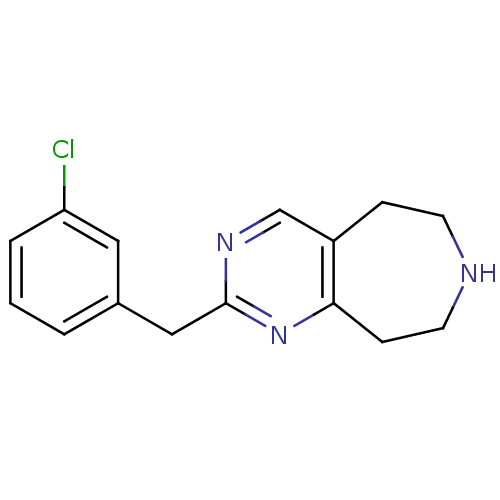
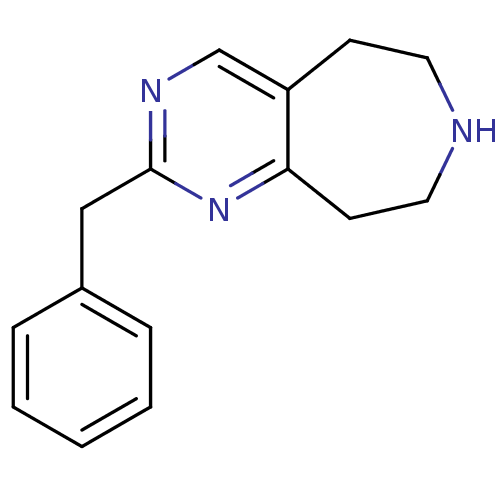
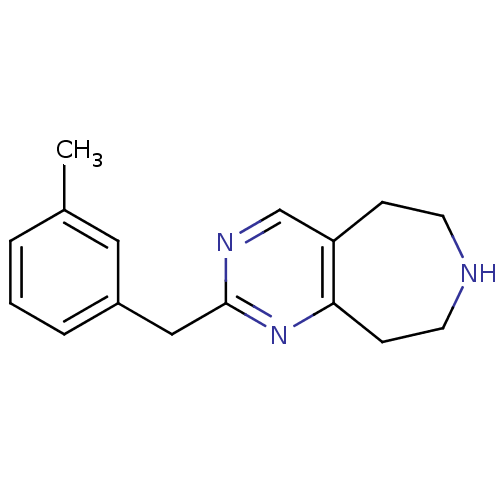
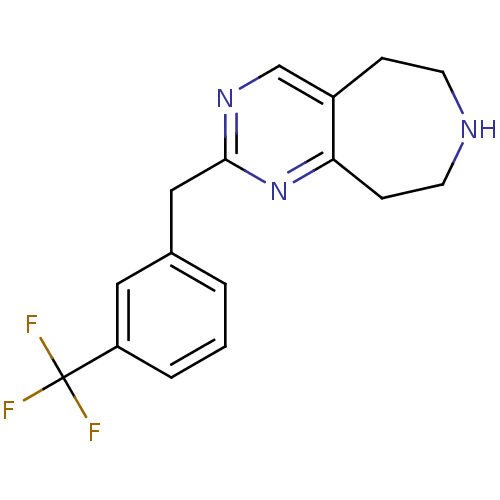
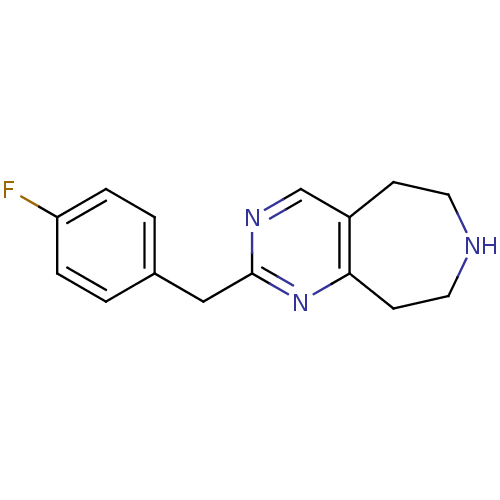
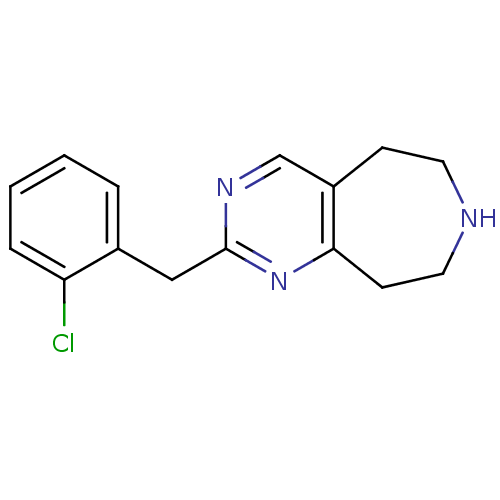
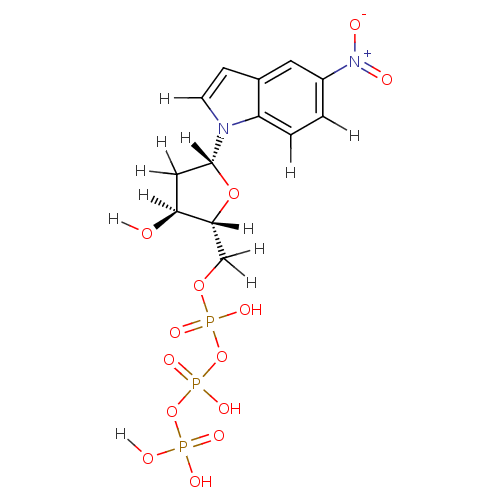
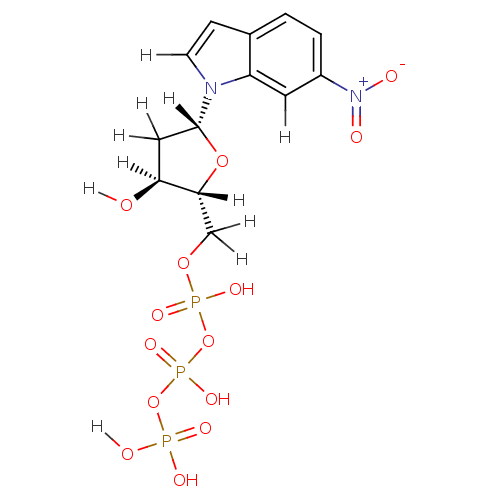
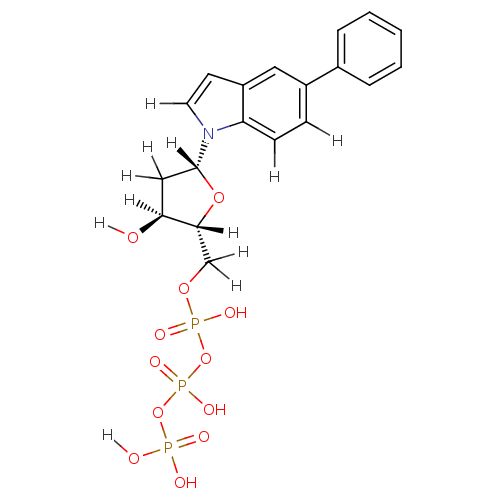
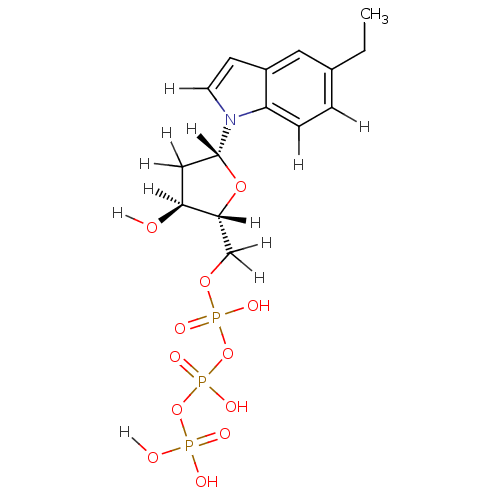
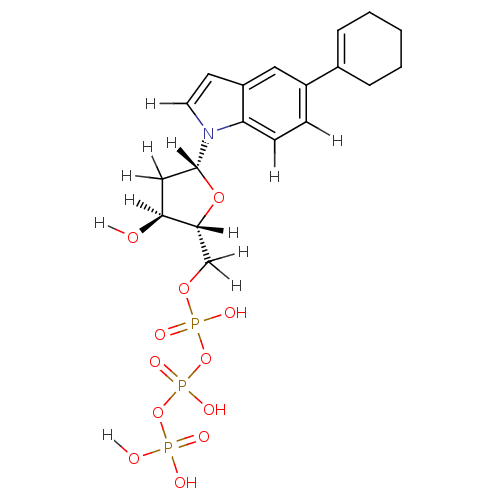
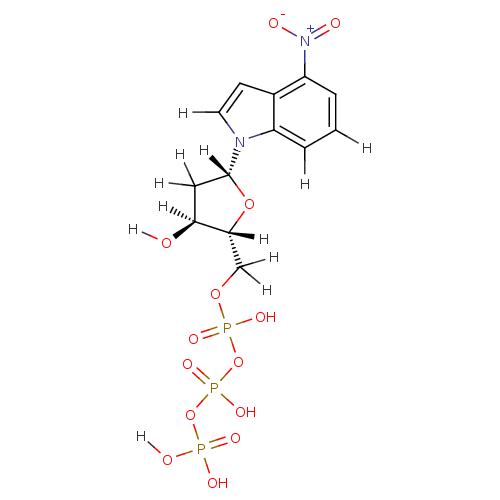
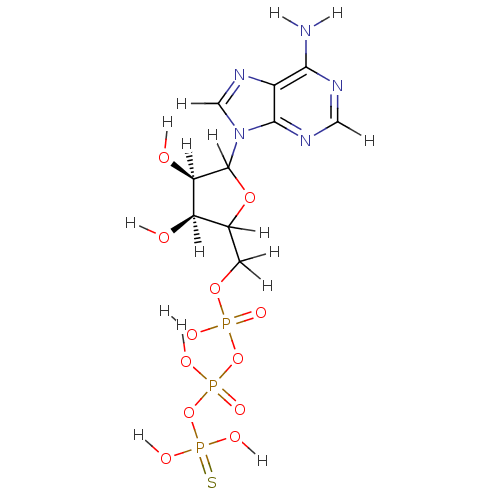
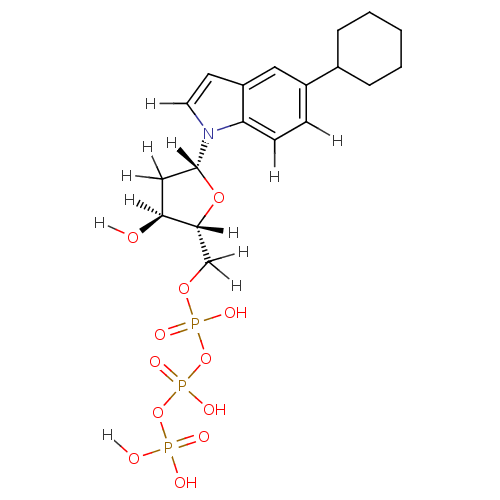
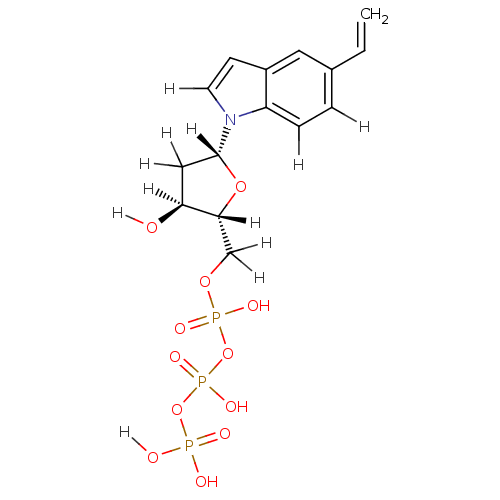
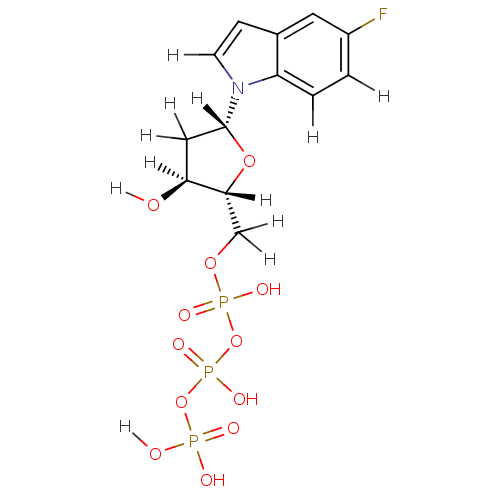
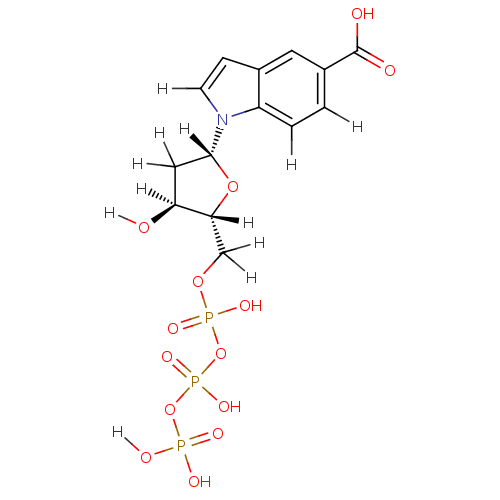
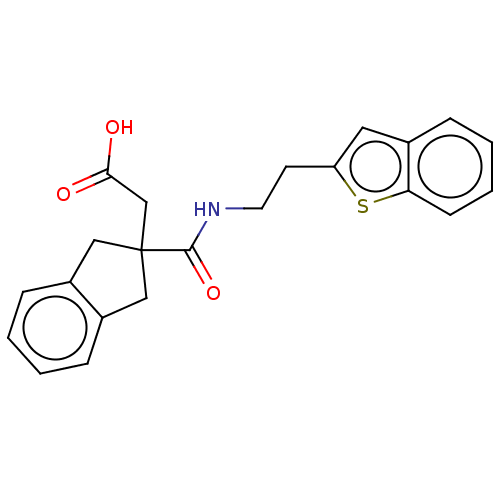
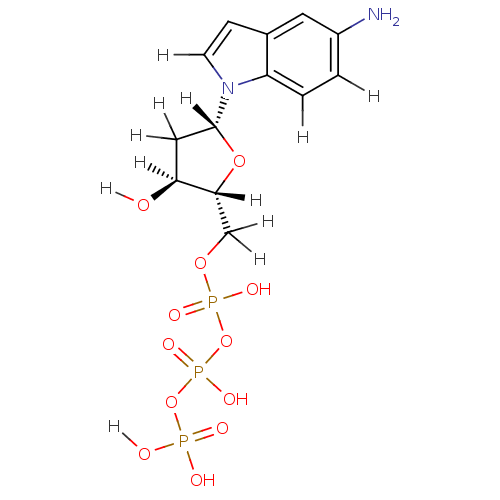
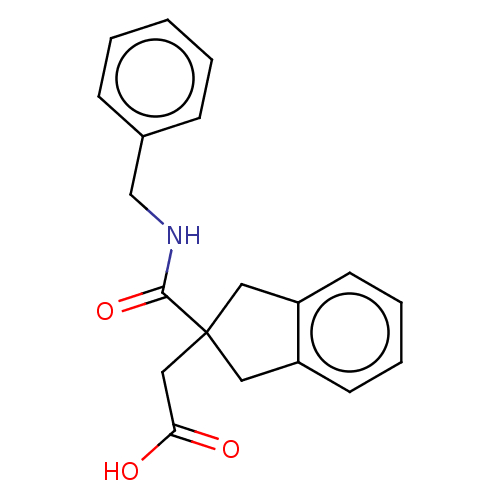
Affinity DataKi: 15nMAssay Description:Displacement of [125I]DOI from human recombinant 5HT2C receptor expressed in HEK293 cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 84nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 167nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 244nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 305nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 308nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 470nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.85E+3nMAssay Description:Displacement of [3H]meselurgine from human 5HT2C receptor expressed in Swiss mouse 3T3 cells by scintillation proximity assayMore data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 4.80E+3nM ΔG°: -30.4kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 5.00E+3nM ΔG°: -30.3kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 5.10E+3nM ΔG°: -30.2kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nM ΔG°: -30.4kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 8.10E+3nM ΔG°: -30.2kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 9.00E+3nM ΔG°: -30.0kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 9.10E+3nM ΔG°: -28.8kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 1.00E+4nM ΔG°: -28.5kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 1.10E+4nM ΔG°: -28.3kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 1.11E+4nM ΔG°: -29.4kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+4nM ΔG°: -29.3kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+4nM ΔG°: -29.0kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 1.38E+4nM ΔG°: -27.7kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 1.40E+4nM ΔG°: -27.7kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 1.64E+4nM ΔG°: -27.3kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.17E+4nM ΔG°: -27.7kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+4nM ΔG°: -27.4kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 2.52E+4nM ΔG°: -26.2kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 2.62E+4nM ΔG°: -26.2kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 2.89E+4nM ΔG°: -25.9kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nM ΔG°: -26.9kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 3.02E+4nM ΔG°: -25.8kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 3.13E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 3.32E+4nM ΔG°: -25.6kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 3.40E+4nM ΔG°: -25.5kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+4nM ΔG°: -26.5kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 3.42E+4nM ΔG°: -25.5kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 3.70E+4nM ΔG°: -25.3kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+4nM ΔG°: -26.1kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175K] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 4.06E+4nM ΔG°: -25.1kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 4.06E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
TargetComplex of Sliding-clamp-loader large subunit [R175L] and Sliding-clamp-loader small subunit(bacteriophage T4)
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 4.16E+4nM ΔG°: -25.0kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 4.20E+4nM ΔG°: -25.0kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: 4.50E+4nM ΔG°: -25.8kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
TargetSliding-clamp-loader large/small subunit(Enterobacteria phage T4 (Bacteriophage T4))
Case Western Reserve University
Case Western Reserve University
Affinity DataKi: 4.75E+4nM ΔG°: -24.7kJ/molepH: 7.5 T: 2°CAssay Description:ATPase assay contained 10 mM Mg2+, 1uM 13/20 DNA, 500uM (r)NTP or (d)NTP, 500 nM gp45, and 500 nM gp44/62 in T4 buffer. ATPase activity was measured ...More data for this Ligand-Target Pair
Affinity DataKi: >5.70E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: >5.70E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: >5.70E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: >5.70E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataKi: >5.70E+4nMAssay Description:Inhibition of rabbit ACE using Abz-FRK(DNP)-P as substrate measured for every 30 sec for 5 mins by UV-fluorescence based assayMore data for this Ligand-Target Pair

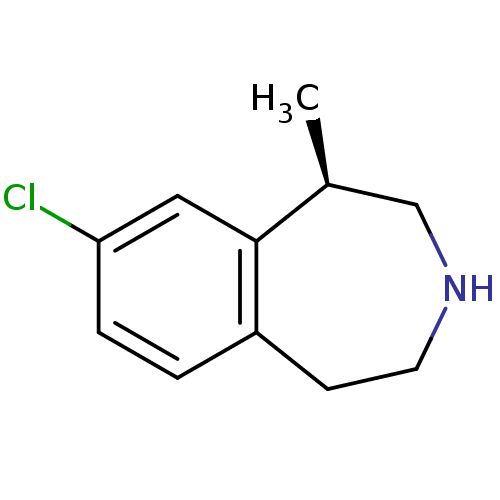
 3D Structure (crystal)
3D Structure (crystal)