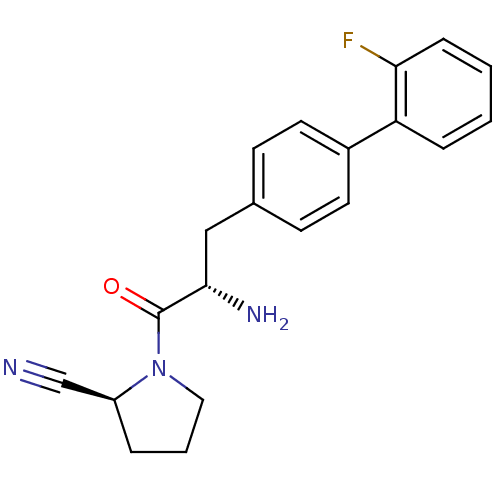
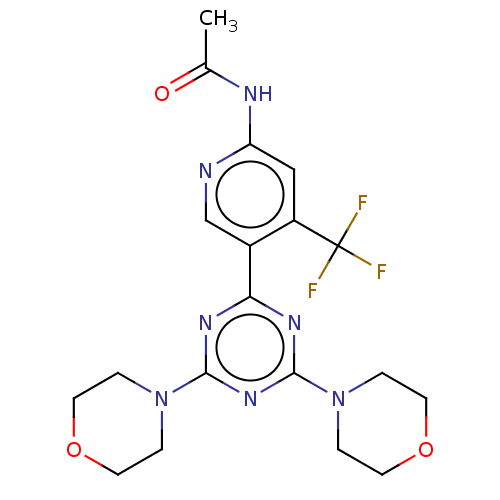
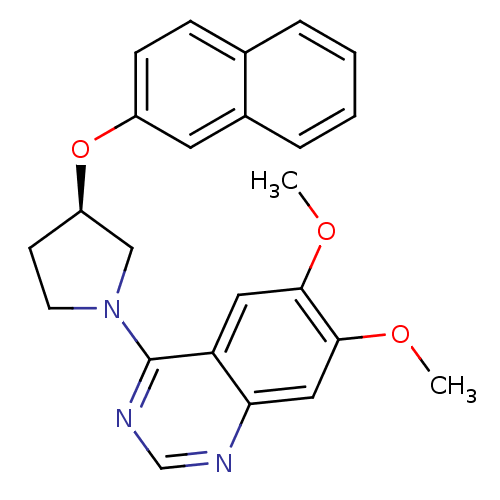
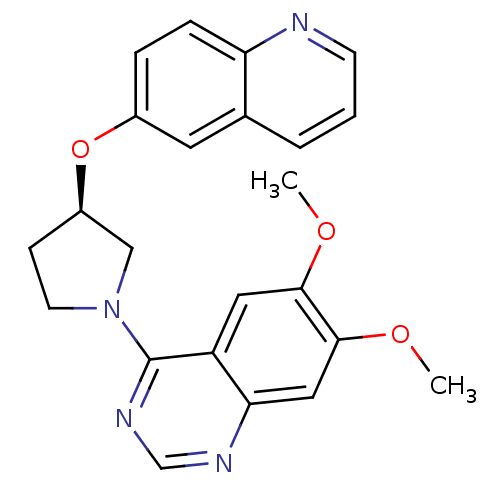
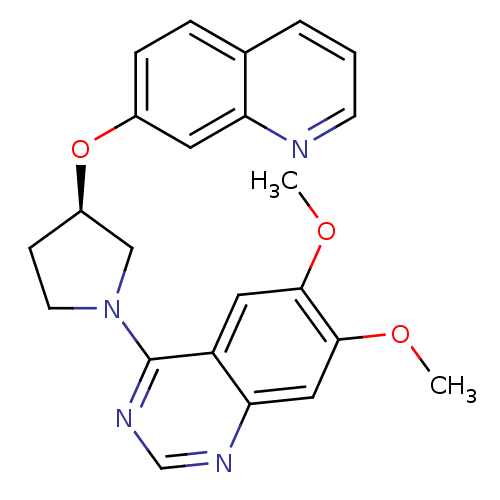
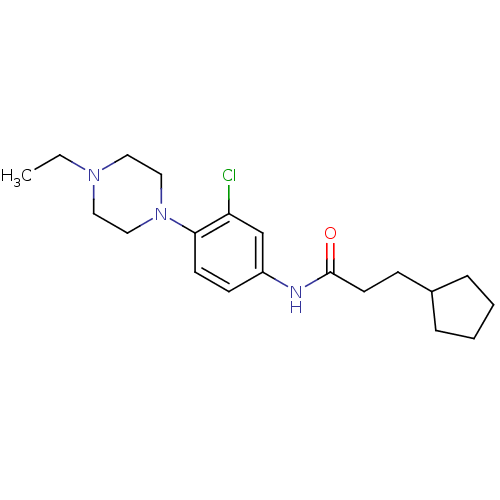
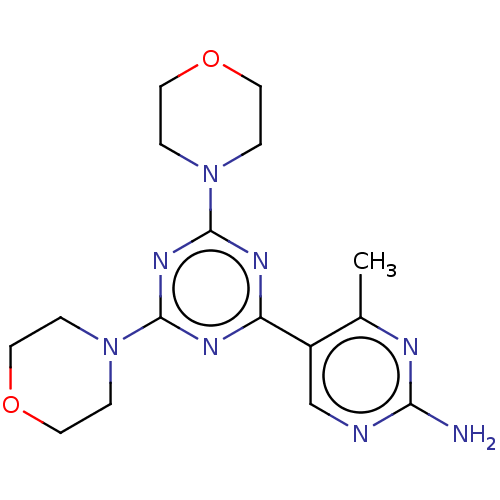
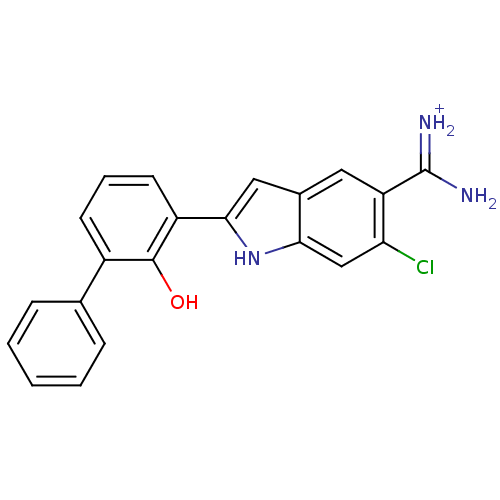
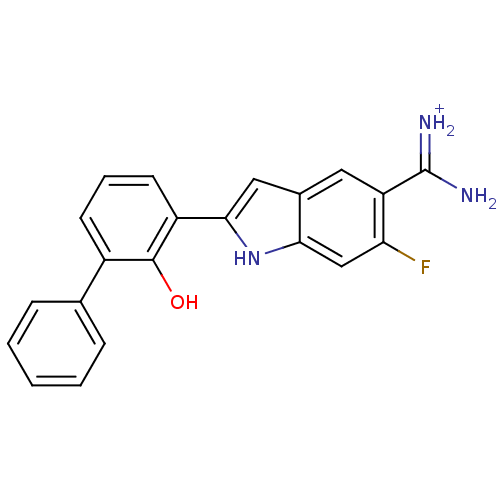
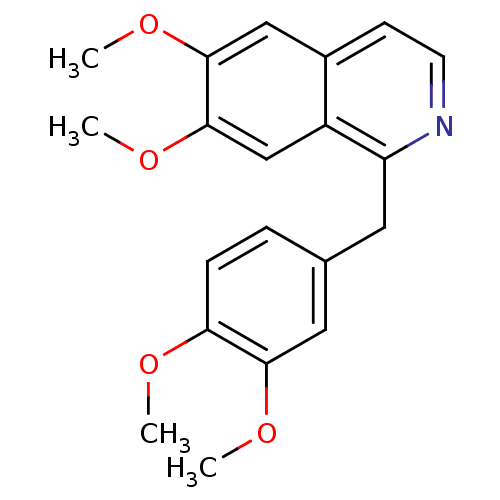
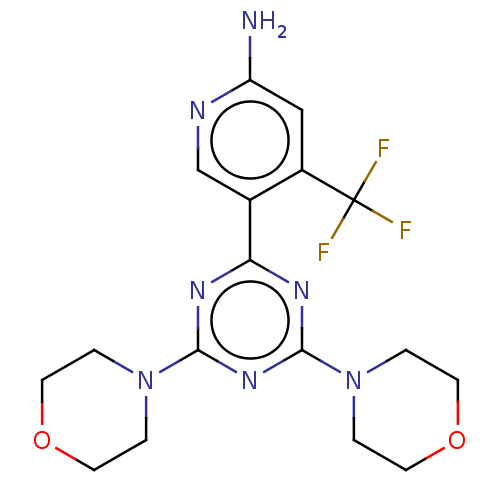
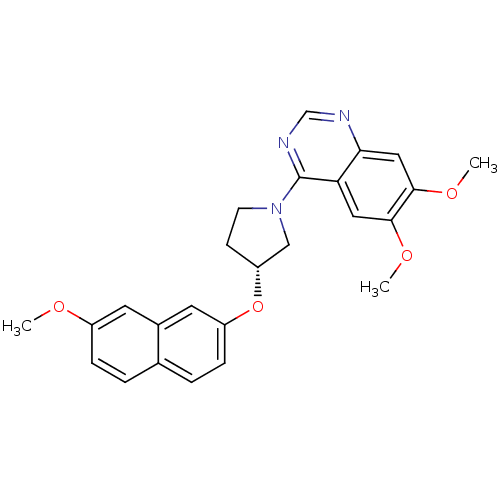
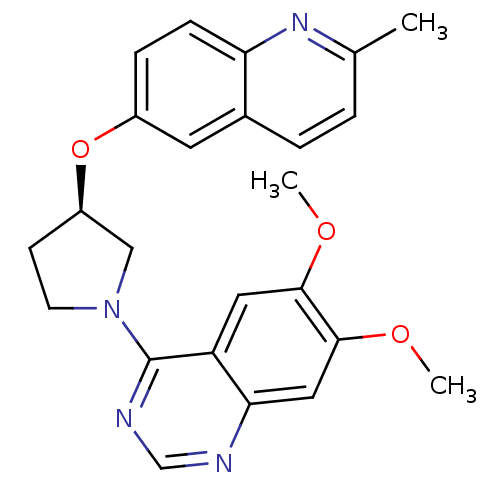
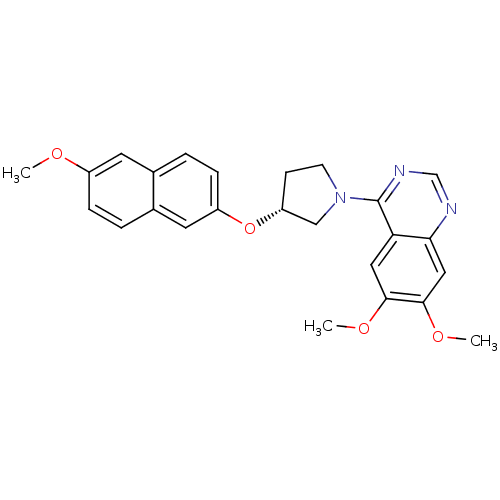
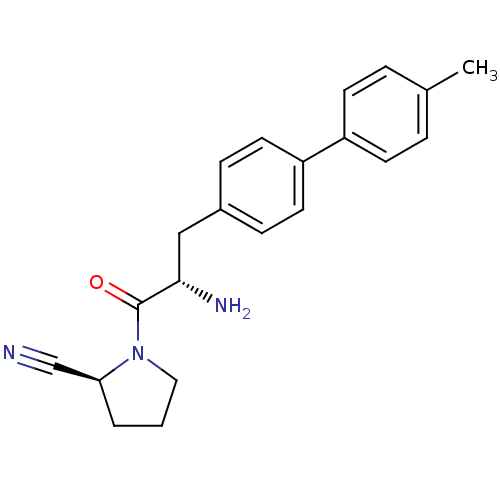
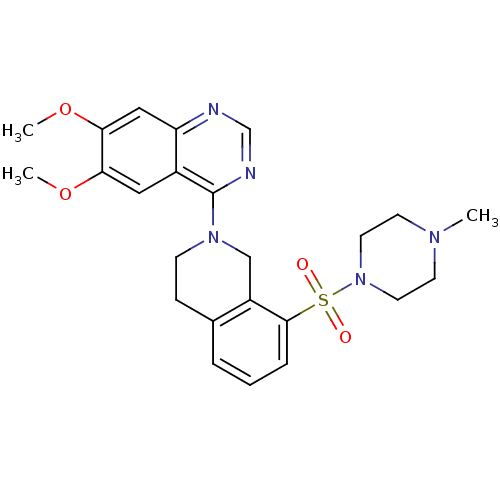
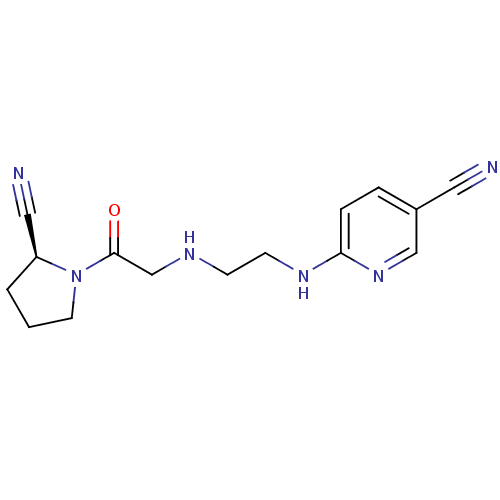
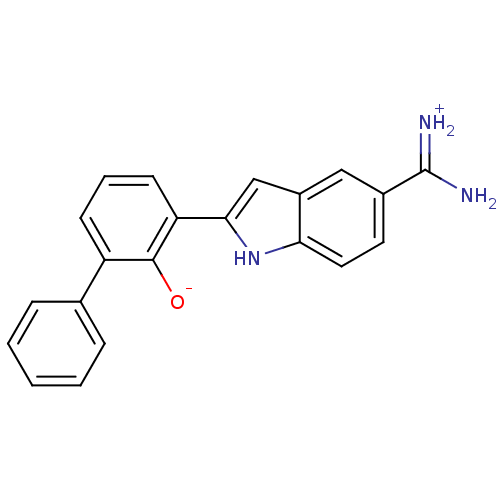
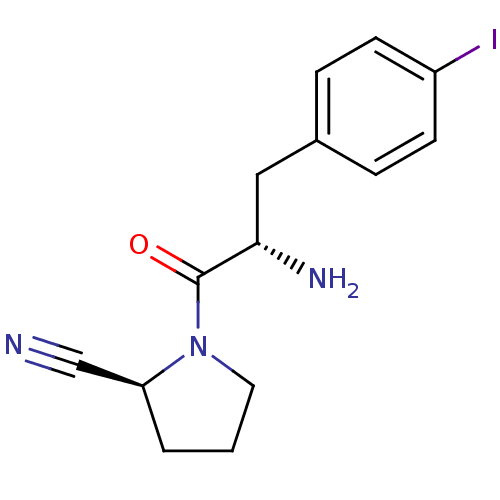
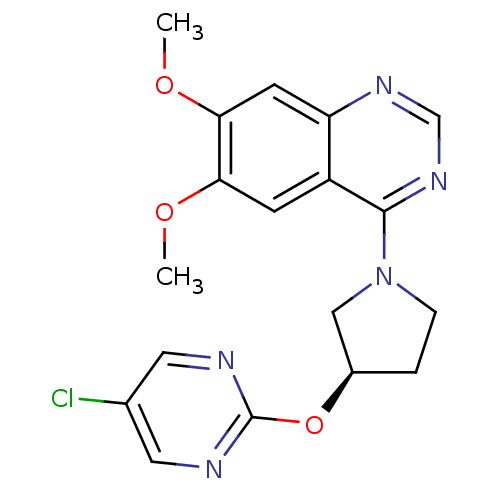
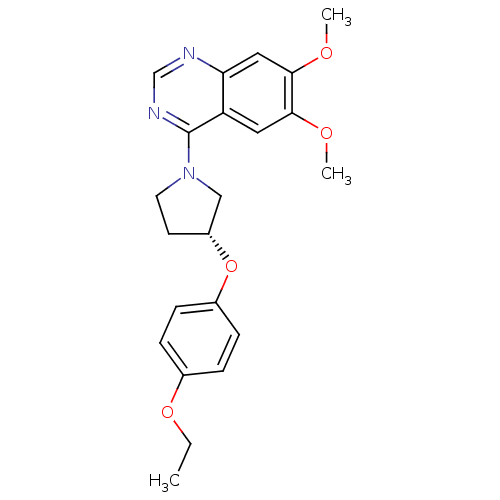
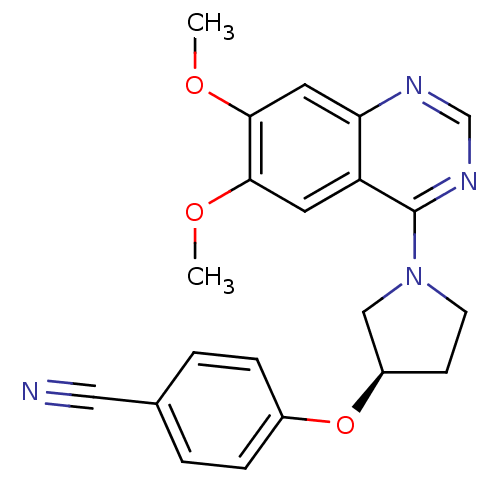
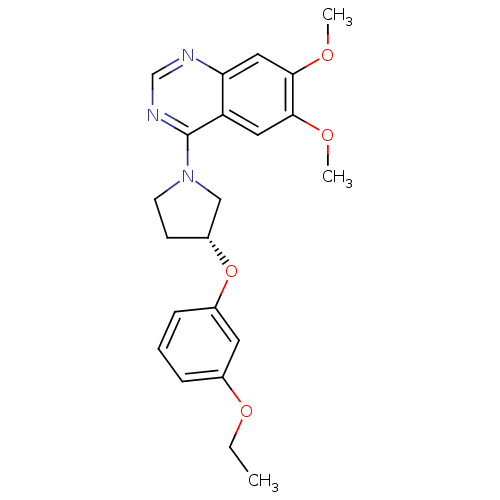
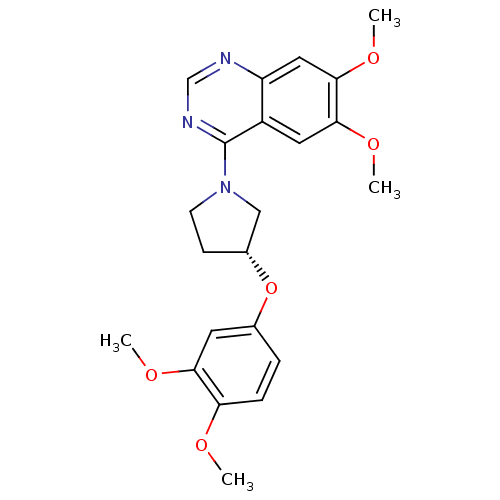
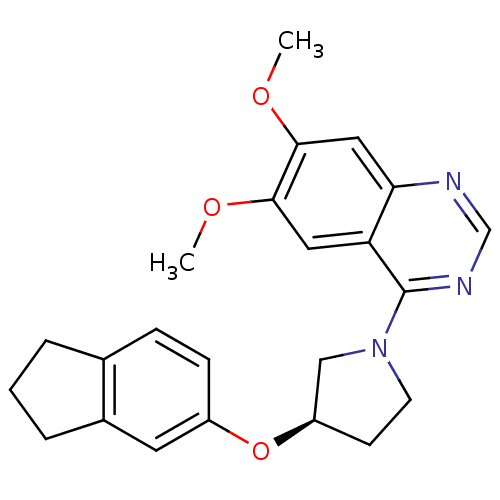
TargetMuscarinic acetylcholine receptor M4(RAT)
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Affinity DataKi: 0.520nMAssay Description:Displacement of [3H]NMS from rat recombinant muscarinic M4 receptor expressed in CHO cells after 2 hrs by microplate scintillation countingMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(RAT)
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Affinity DataKi: 0.520nMAssay Description:Displacement of [3H]NMS from rat recombinant muscarinic M4 receptor expressed in CHO cells after 2 hrs by microplate scintillation countingMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M4(RAT)
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Affinity DataKi: 0.560nMAssay Description:Displacement of [3H]NMS from rat muscarinic M4 receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M5(RAT)
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Affinity DataKi: 0.860nMAssay Description:Displacement of [3H]NMS from rat recombinant muscarinic M5 receptor expressed in CHO cells after 2 hrs by microplate scintillation countingMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M5(RAT)
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Dvanderbilt Program In Drug Discovery
Curated by ChEMBL
Affinity DataKi: 1.80nMAssay Description:Displacement of [3H]NMS from rat muscarinic M5 receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -49.4kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 3.10nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 4nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 5.30nM ΔG°: -47.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 8.10nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 9.70nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 12nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 12nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 12nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(RAT)
Vanderbilt Institute Of Chemical Biology
Curated by ChEMBL
Vanderbilt Institute Of Chemical Biology
Curated by ChEMBL
Affinity DataKi: 12.7nMAssay Description:Displacement of [3H]NMS from rat muscarinic M1 receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -45.0kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 17nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 18nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 18nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 19nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 25nM ΔG°: -43.0kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -43.3kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -43.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 36nM ΔG°: -42.5kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 44nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 46nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 54nM ΔG°: -41.1kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 56nM ΔG°: -41.0kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 61nMAssay Description:Inhibition of human N-terminal His6-tagged PI3K p110alpha/p85alpha expressed in baculovirus expression system after 1 hr using AlexaFluor647-labeled ...More data for this Ligand-Target Pair
TargetSerine/threonine-protein kinase mTOR(Homo sapiens (Human))
University Of Basel
Curated by ChEMBL
University Of Basel
Curated by ChEMBL
Affinity DataKi: 62nMAssay Description:Inhibition of human C-terminal GST-tagged mTOR (1360 to 2549 residues) expressed in baculovirus expression system using after 1 hr AlexaFluor647-labe...More data for this Ligand-Target Pair
Affinity DataKi: 63nM ΔG°: -41.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 67nM ΔG°: -40.5kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 68nM ΔG°: -40.5kJ/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M3(RAT)
Vanderbilt Institute Of Chemical Biology
Curated by ChEMBL
Vanderbilt Institute Of Chemical Biology
Curated by ChEMBL
Affinity DataKi: 74.8nMAssay Description:Displacement of [3H]NMS from rat muscarinic M3 receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: 85nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair

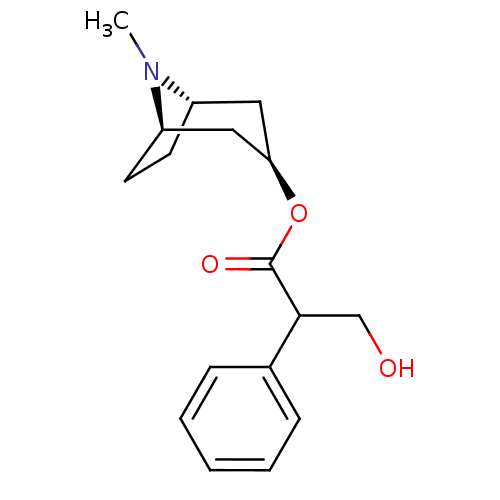
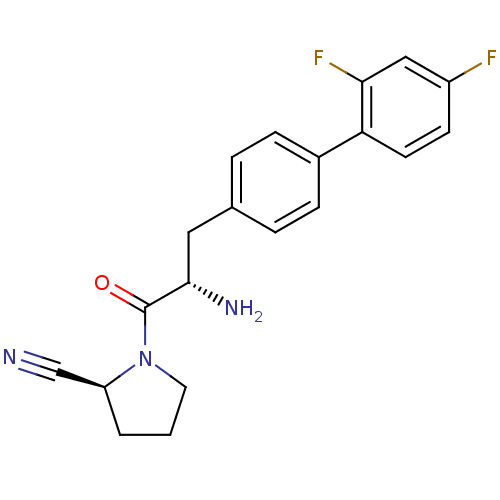

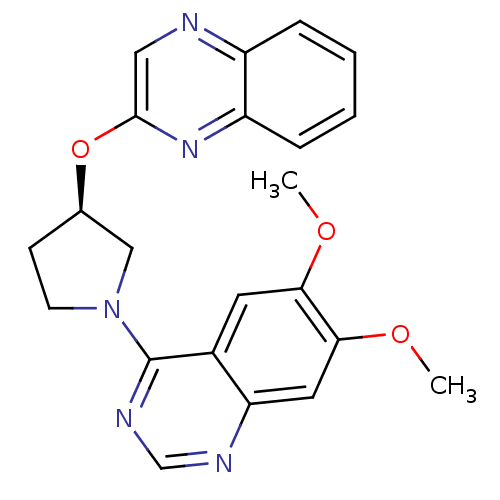
 3D Structure (crystal)
3D Structure (crystal)