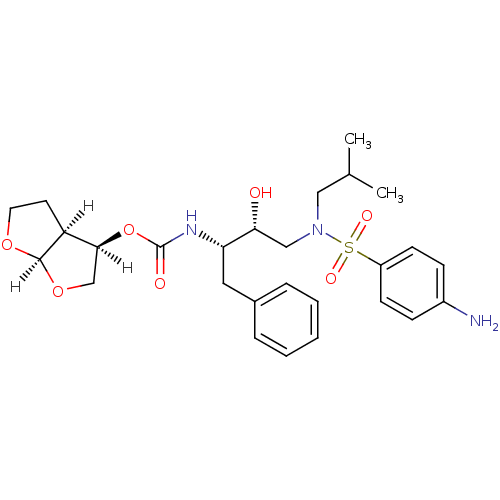
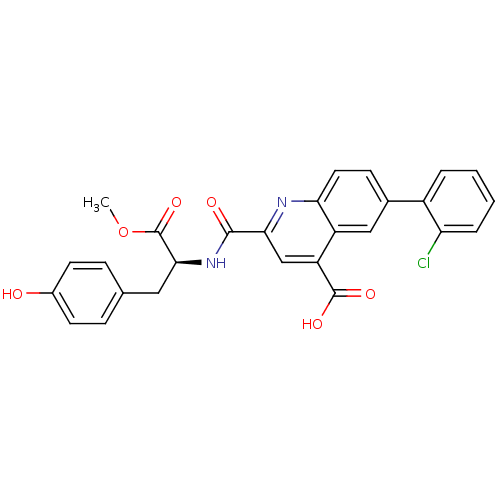
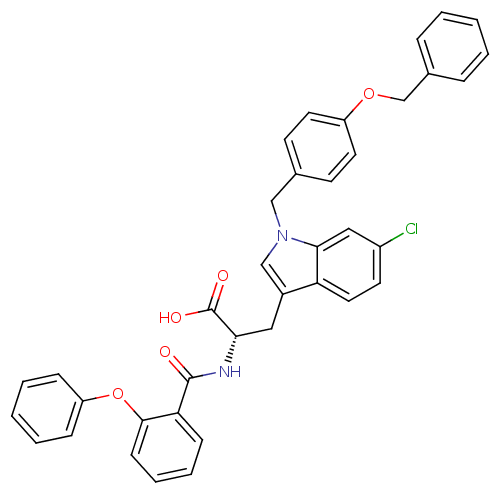
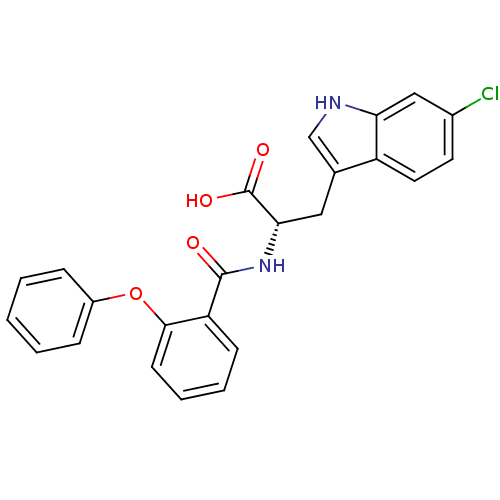
Affinity DataKi: 0.5nMAssay Description:In vitro inhibitory effect of histamine H3 antagonist on the electrically evoked contractile response of isolated guinea pig jejunum segments.More data for this Ligand-Target Pair
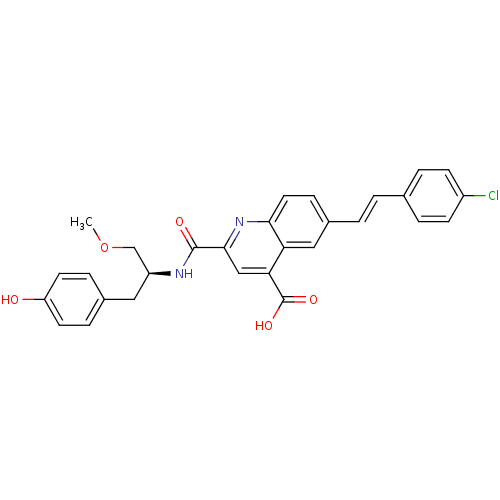
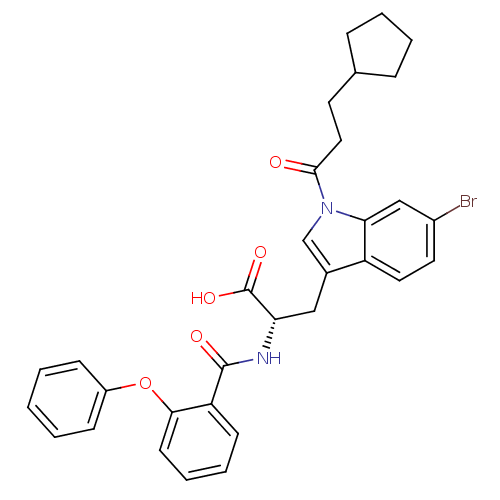
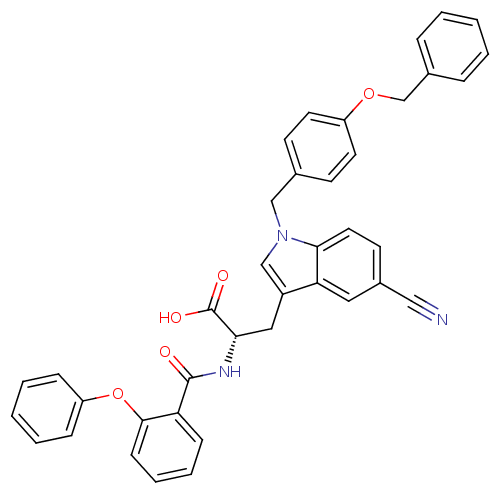
Affinity DataKi: 10nMAssay Description:In vitro inhibitory effect of histamine H3 antagonist on the electrically evoked contractile response of isolated guinea pig jejunum segments.More data for this Ligand-Target Pair
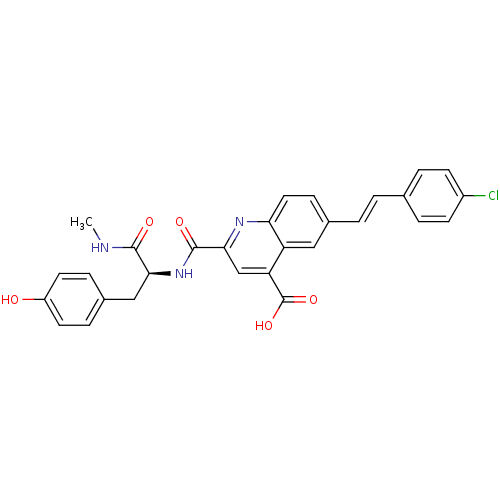
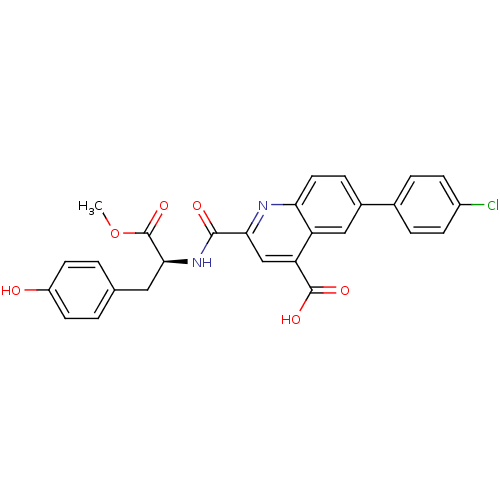
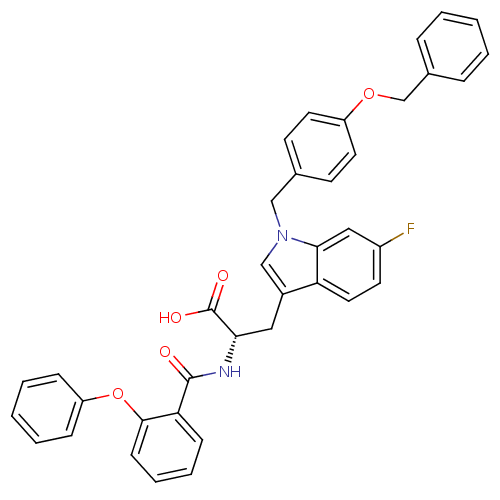
Affinity DataKi: 10nMAssay Description:In vitro inhibitory effect of histamine H3 antagonist on the electrically evoked contractile response of isolated guinea pig jejunum segments.More data for this Ligand-Target Pair
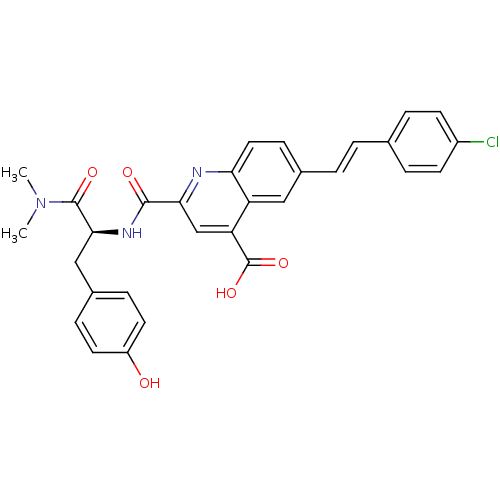
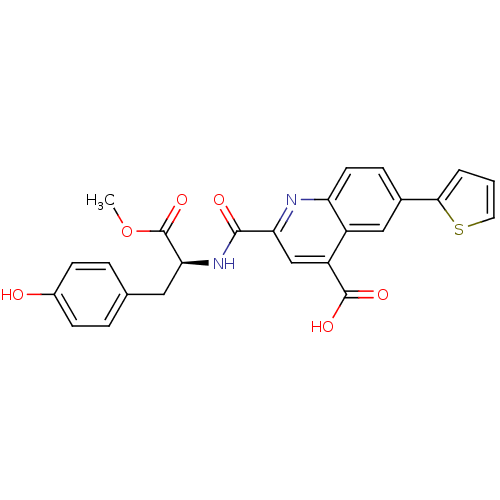
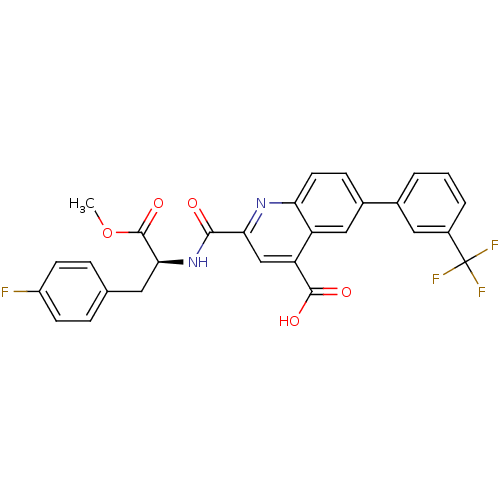
Affinity DataKi: 100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nM ΔG°: -39.7kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 400nM ΔG°: -36.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -35.7kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 900nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 900nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 900nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+3nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nM ΔG°: -31.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Compound was evaluated for its binding affinity against NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nM ΔG°: -29.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 9.00E+3nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nMAssay Description:Binding affinity against Hepatitis C virus NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nMAssay Description:Displacement of B-Alexa-Fluor647 from CDK2 (unknown origin) by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair



 3D Structure (crystal)
3D Structure (crystal)