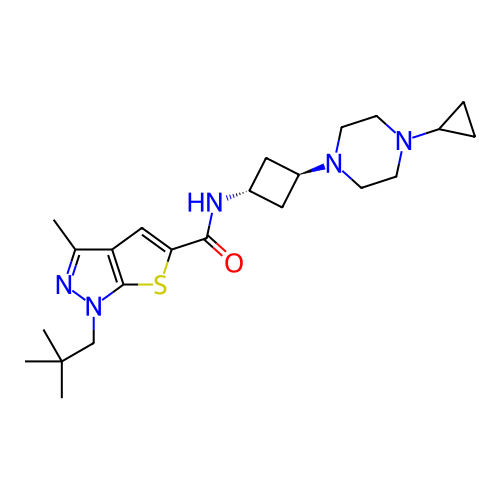
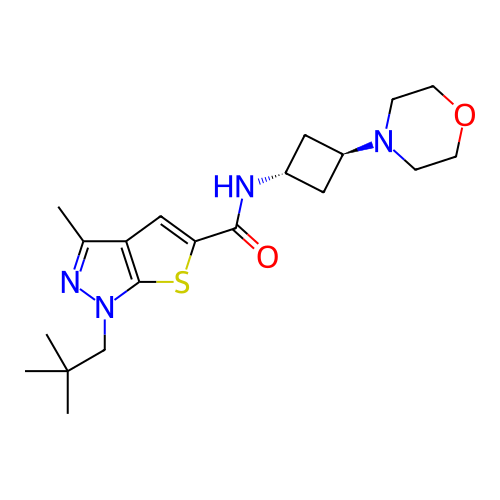
Report error Found 106 Enz. Inhib. hit(s) with all data for entry = 12743
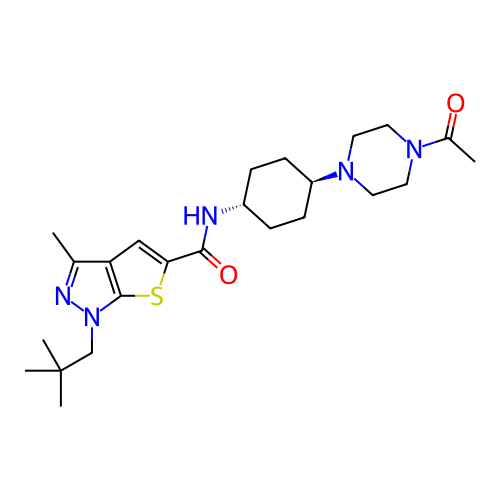
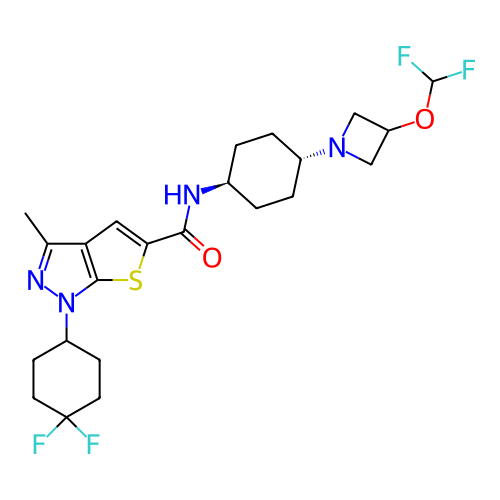
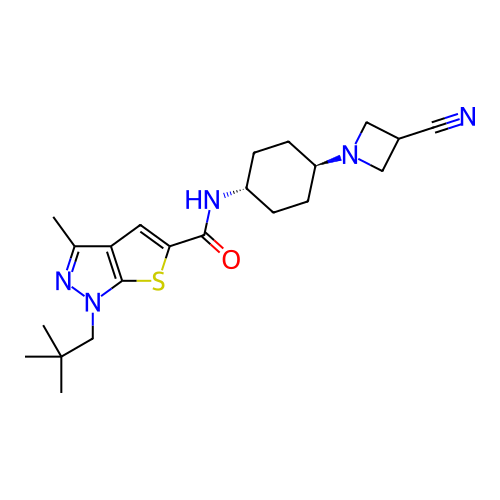
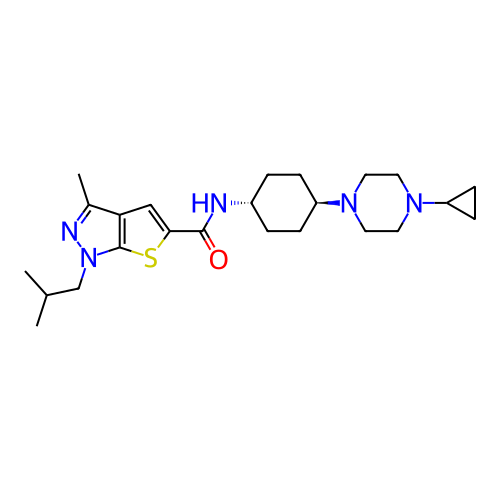
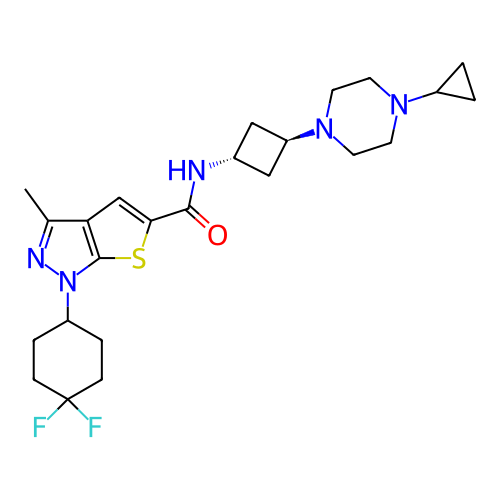
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
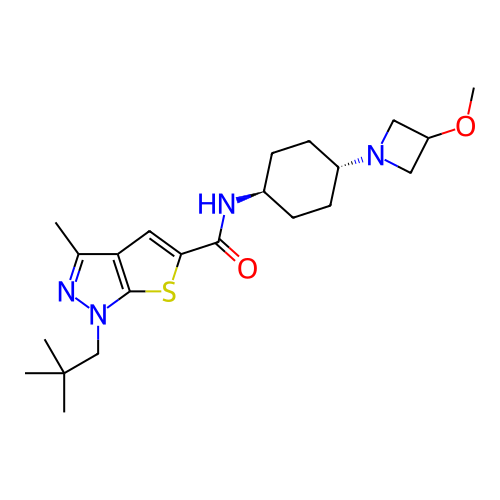
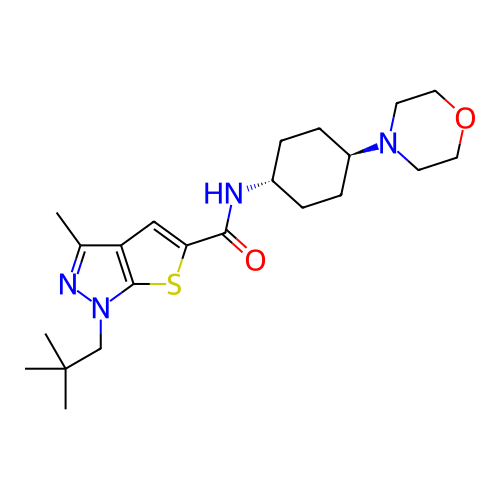
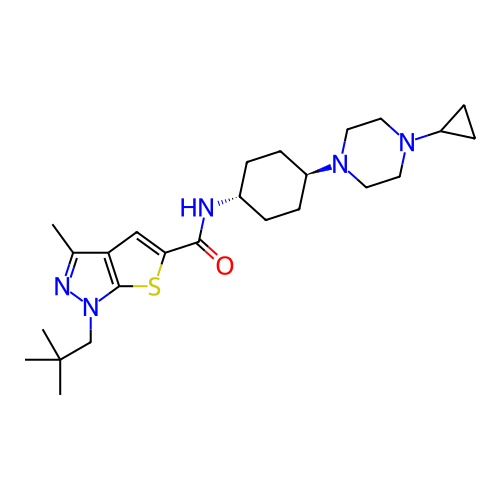
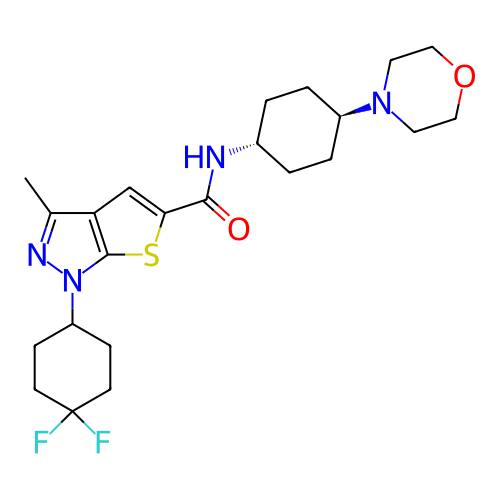
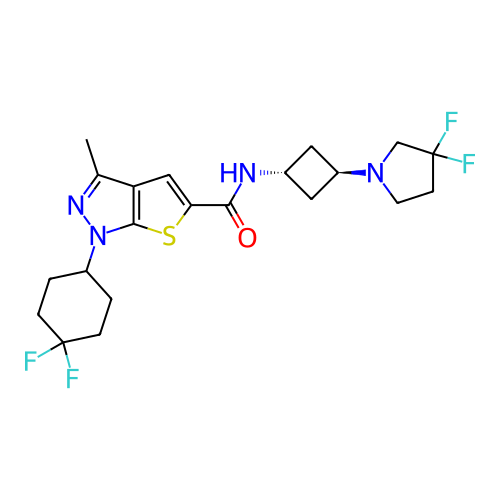
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
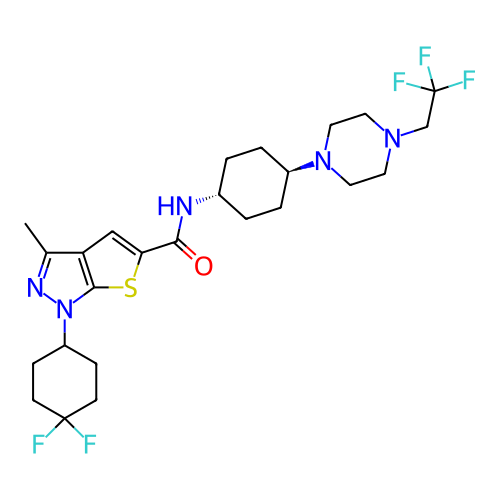
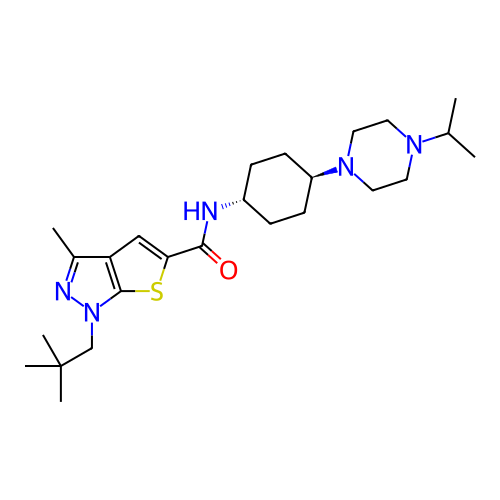
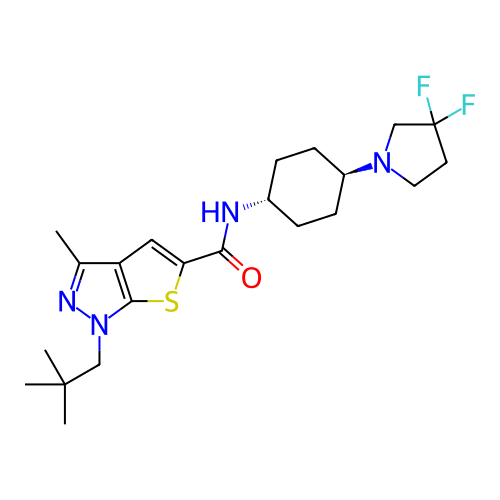
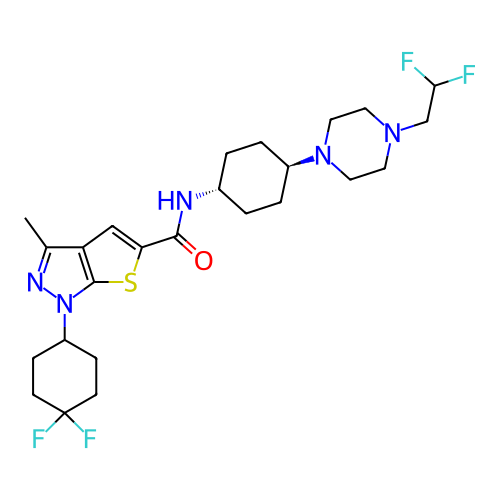
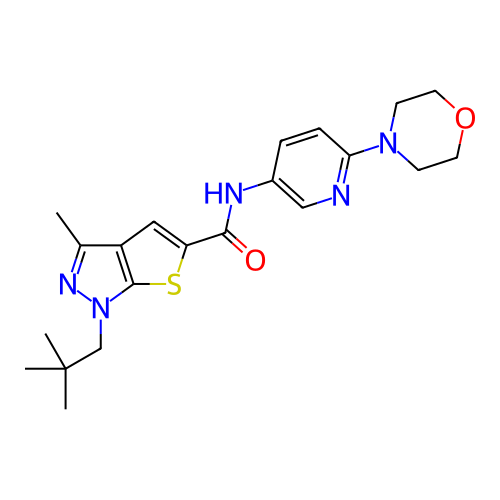
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
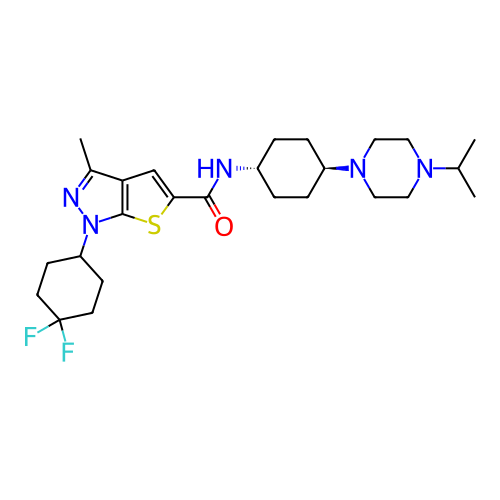
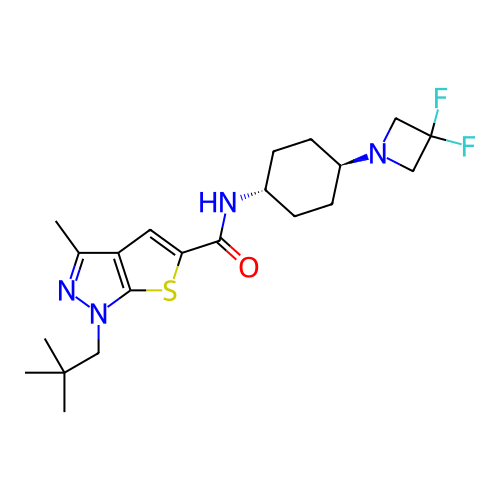
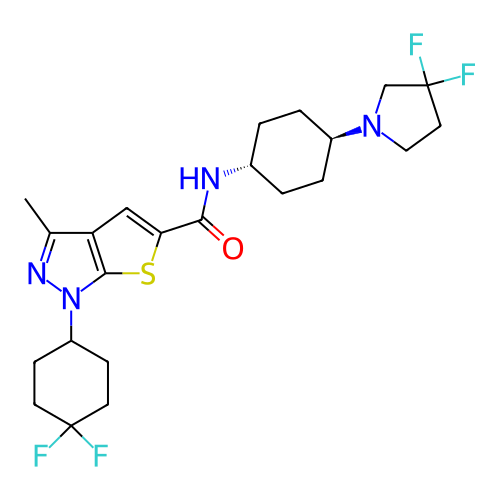
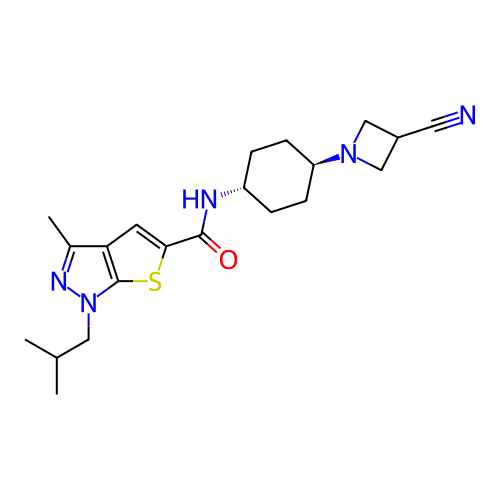
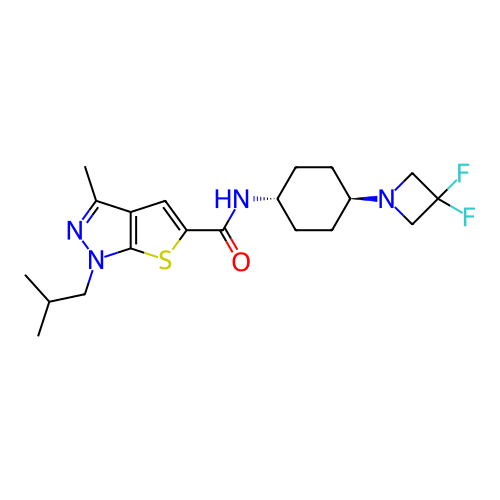
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair